IRAS16293 Band9 - Imaging for CASA 3.4: Difference between revisions
| (49 intermediate revisions by 3 users not shown) | |||
| Line 1: | Line 1: | ||
*'''This portion of the guide covers imaging of the fully calibrated data see: [[IRAS16293_Band9_-_Calibration_for_CASA_3.4]]'''. | |||
<pre style="background-color: #ffa07a;"> | |||
WARNING: On June 15, 2012 the calibration guide and the final data products (calibrated science | |||
data: IRAS16293_Band9_CalibratedMS_FIXED.tgz and reference images: IRAS16293_Band9_ReferenceImages_FIXED.tgz)) | |||
were changed to correct for a 1.2" position error in the phase calibrator (1625-254). Without | |||
correction, the science images will suffer from a similar offset. | |||
</pre> | |||
== Overview == | == Overview == | ||
This section of the casa guide continues from | This section of the casa guide continues from [http://casaguides.nrao.edu/index.php?title=IRAS16293_Band9_-_Calibration_for_CASA_3.4 IRAS16293_Band9_-_Calibration_for_CASA_3.4] . If you completed the calibration section, then you already have the final calibrated dataset. If you just started to work on this IRAS16293 B9 data, you can download the fully calibrated dataset '''IRAS16293_Band9_CalibratedMS''' from | ||
[https://almascience.nrao.edu/almadata/sciver/IRAS16293B9 North America] | [https://almascience.nrao.edu/almadata/sciver/IRAS16293B9 North America] | ||
| Line 8: | Line 17: | ||
[http://almascience.nao.ac.jp/almadata/sciver/IRAS16293B9 East Asia] | [http://almascience.nao.ac.jp/almadata/sciver/IRAS16293B9 East Asia] | ||
This casa guide covers imaging of both continuum and | This casa guide covers imaging of both continuum and spectral lines of IRAS16293 Band 9. '''This guide requires CASA 3.4'''. Among other things, CASA 3.4.0 contains an improved antenna primary beam model for ALMA. In this casa guide you will also need the [http://casaguides.nrao.edu/index.php?title=Analysis_Utilities Analysis Utils] package, so make sure you have it imported into CASA. | ||
==Confirm your version of CASA== | |||
This guide has been written for CASA release 3.4.0. Please confirm your version before proceeding. | |||
<source lang="python"> | |||
# In CASA | |||
version = casalog.version() | |||
print "You are using " + version | |||
if (int(version.split()[4][1:-1]) < 19874): | |||
print "\033[91m YOUR VERSION OF CASA IS TOO OLD FOR THIS GUIDE." | |||
print "\033[91m PLEASE UPDATE IT BEFORE PROCEEDING." | |||
else: | |||
print "Your version of CASA is appropriate for this guide." | |||
</source> | |||
==Install Analysis Utilities== | |||
Analysis Utilities (or analysisUtils for short) is a small set of Python scripts that provide a number of analysis and plotting utilities for ALMA data reduction. This guide uses a few of these utilities. They are very easy to install (just download and untar). See | |||
http://casaguides.nrao.edu/index.php?title=Analysis_Utilities | |||
for a full description and download instructions. If you do not wish to do this, see the CASA 3.3 version of the guide linked at the top of this page for alternative (but slow) plotting options. Analysis Utilities are updated frequently so if its been a while since you installed it, its probably worth doing it again. If you are at an ALMA site or ARC, the analysis utilities are probably already installed and up to date. | |||
== List the contents of the Data== | |||
Before we start with the imaging itself, do a {{listobs}} to the calibrated dataset, so you can check that everything is ok. | Before we start with the imaging itself, do a {{listobs}} to the calibrated dataset, so you can check that everything is ok. | ||
| Line 14: | Line 47: | ||
<source lang="python"> | <source lang="python"> | ||
# listobs to check data | # listobs to check data | ||
listobs(vis='IRAS16293_Band9.rebin.ms', | listobs(vis='IRAS16293_Band9.fixed.rebin.ms', | ||
listfile='IRAS16293_Band9.rebin.ms.listobs',verbose=F) | listfile='IRAS16293_Band9.fixed.rebin.ms.listobs',verbose=F) | ||
</source> | </source> | ||
| Line 41: | Line 74: | ||
ID= 9-13: 'DV12'='A011', 'DV13'='A072', 'DV14'='A025', 'DV17'='A138', | ID= 9-13: 'DV12'='A011', 'DV13'='A072', 'DV14'='A025', 'DV17'='A138', | ||
</pre> | </pre> | ||
'''If you are using the calibrated data from the science portal, it is essential to do the following.''' | |||
<source lang="python"> | |||
# In CASA | |||
tb.open('IRAS16293_Band9.fixed.rebin.ms/POINTING', nomodify = False) | |||
a = tb.rownumbers() | |||
tb.removerows(a) | |||
tb.close() | |||
</source> | |||
== Continuum Emission Imaging == | == Continuum Emission Imaging == | ||
As | As shown in Figure 2 of [[IRAS16293Band9]], these observations have a 7 pointing mosaic, which you can visualize with the next command (see also Figure 1). | ||
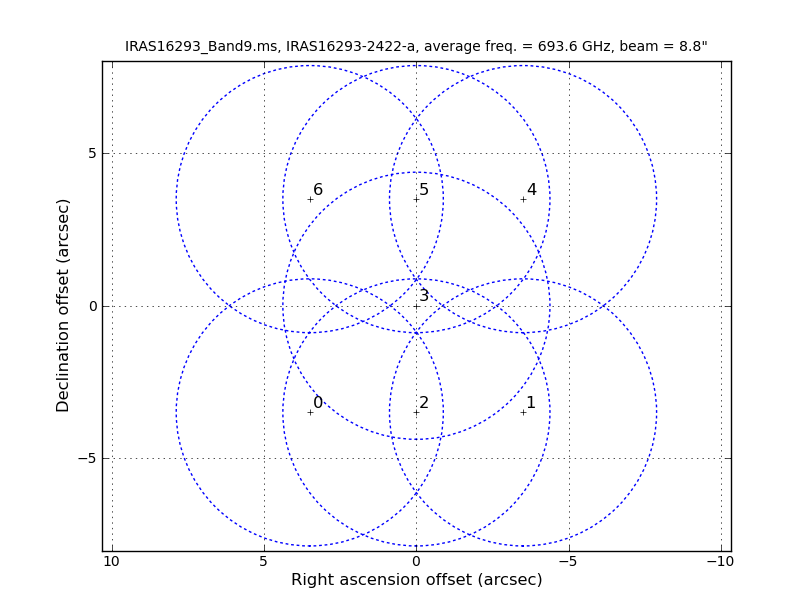
[[File:mosaic_pattern.png|200px|thumb|right|'''Fig. 1.''' Pointings showing the mosaic used for the observations.]] | [[File:mosaic_pattern.png|200px|thumb|right|'''Fig. 1.''' Pointings showing the mosaic used for the observations. The circles mark the FWHM of the primary beam response.]] | ||
<source lang="python"> | <source lang="python"> | ||
# Make a plot of the mosaic pattern with field ids identified | # Make a plot of the mosaic pattern with field ids identified | ||
au.plotMosaic(vis='IRAS16293_Band9.fixed.rebin.ms',sourceid='0', | |||
doplot=True,figfile='mosaic_pattern.png') | doplot=True,figfile='mosaic_pattern.png') | ||
</source> | </source> | ||
You might | You might wonder which of these pointings or fields have stronger line emission. You can determine this by running the next command. We set spw='1' which contains CO (6-5), and you can check all the plots for all the fields by just hitting the arrow for "next". Also see Figure 2 for an example of the output. | ||
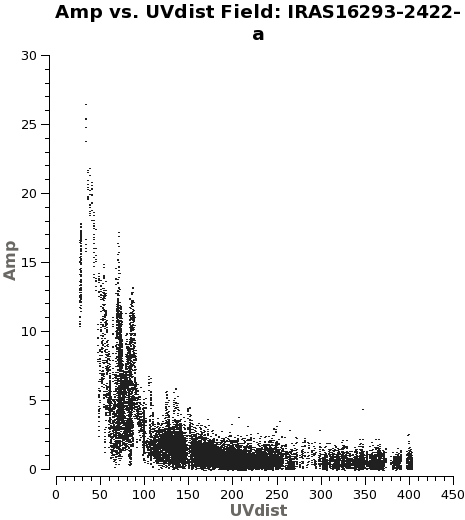
[[File:IRAS16293_Band9.AVG.ms_uvamp_f0_spw1.png|200px|thumb|right|'''Fig. 2.''' UV-amp distribution for the field 0 (spw 1) in the mosaic.]] | [[File:IRAS16293_Band9.AVG.ms_uvamp_f0_spw1.png|200px|thumb|right|'''Fig. 2.''' UV-amp distribution for the field 0 (spw 1) in the mosaic.]] | ||
| Line 59: | Line 102: | ||
<source lang="python"> | <source lang="python"> | ||
# See which mosaic fields have the brightest line emission | # See which mosaic fields have the brightest line emission | ||
plotms(vis='IRAS16293_Band9.rebin.ms', | plotms(vis='IRAS16293_Band9.fixed.rebin.ms', | ||
field='',xaxis='uvdist', yaxis='amp', | field='',xaxis='uvdist', yaxis='amp', | ||
spw='1', avgchannel='3840',coloraxis='field', | spw='1', avgchannel='3840',coloraxis='field', | ||
| Line 71: | Line 114: | ||
<source lang="python"> | <source lang="python"> | ||
# Iterate over spws all fields | # Iterate over spws all fields | ||
plotms(vis='IRAS16293_Band9.rebin.ms', | plotms(vis='IRAS16293_Band9.fixed.rebin.ms', | ||
field='',xaxis='uvdist', yaxis='amp', | field='',xaxis='uvdist', yaxis='amp', | ||
spw='', avgchannel='3840',coloraxis='field', | spw='', avgchannel='3840',coloraxis='field', | ||
| Line 79: | Line 122: | ||
We need to note the channels that do not present line emission. To identify them we can plot the amplitude vs frequency for all the spectral windows. Figure 4 shows the output from the next command for spw 0 (field 3). | We need to note the channels that do not present line emission. To identify them we can plot the amplitude vs frequency for all the spectral windows. Figure 4 shows the output from the next command for spw 0 (field 3). | ||
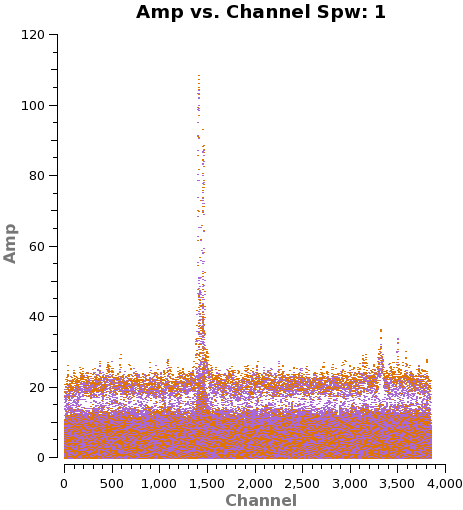
[[File: | [[File:amp_vs_channel.spw1.png|200px|thumb|right|'''Fig. 4.''' Frequency-amplitude plot for spw=1 of field=3. ]] | ||
<source lang="python"> | <source lang="python"> | ||
| Line 85: | Line 128: | ||
# increased S/N. | # increased S/N. | ||
plotms(vis='IRAS16293_Band9.rebin.ms', | plotms(vis='IRAS16293_Band9.fixed.rebin.ms', | ||
xaxis='channel', yaxis='amp',field='3', | xaxis='channel', yaxis='amp',field='3', | ||
spw='', avgtime='1e8',avgscan=T,coloraxis=' | spw='', avgtime='1e8',avgscan=T,coloraxis='corr', | ||
avgchannel=' | avgchannel='',iteraxis='spw',ydatacolumn='corrected', | ||
xselfscale=True,yselfscale=True) | xselfscale=True,yselfscale=True) | ||
</source> | </source> | ||
Using the plots from the previous command you can select | Using the plots from the previous command you can select a block of channels that are free of strong line emission. To do this in the most rigorous way, you would first need to make dirty image cubes and then examine the spectra on and near the continuum sources, because many more weaker lines will become apparent in the cubes. Here we provide you with the channel selection we used for continuum emission. | ||
Here we provide you with the channel selection we used for continuum emission. | |||
The next {{flagmanager}} command will save the flags state in case you might need to re-do the steps from this point. | The next {{flagmanager}} command will save the flags state in case you might need to re-do the steps from this point. | ||
<source lang="python"> | <source lang="python"> | ||
flagmanager(vis='IRAS16293_Band9.ms',mode='save',versionname='Original') | flagmanager(vis='IRAS16293_Band9.fixed.rebin.ms',mode='save',versionname='Original') | ||
</source> | </source> | ||
| Line 105: | Line 147: | ||
<source lang="python"> | <source lang="python"> | ||
# flagging the line channels | # flagging the line channels | ||
flagdata(vis='IRAS16293_Band9.rebin.ms', | flagdata(vis='IRAS16293_Band9.fixed.rebin.ms', | ||
spw='0:1700~2500,1:400~500;1100~1900;3000~3450,2:1700~2200 | spw='0:1700~2500,1:400~500;1100~1900;3000~3450,2:1700~2200;3100~3840,3:0~800;1600~2900;3300~3840',flagbackup=F) | ||
</source> | </source> | ||
| Line 112: | Line 154: | ||
<source lang="python"> | <source lang="python"> | ||
split(vis='IRAS16293_Band9.rebin.ms', | split(vis='IRAS16293_Band9.fixed.rebin.ms', | ||
outputvis=' | outputvis='IRAS16293_Band9.fixed.CONT.msAVG',field='', | ||
datacolumn='data',width=200,spw='0~3:20~3819') | datacolumn='data',width=200,spw='0~3:20~3819') | ||
</source> | </source> | ||
=== Clean and self-calibrate the continuum === | |||
=== Clean and self- | |||
Now we proceed with the cleaning itself. For the {{clean}} task you will need to choose the right cell size, map size, etc. Below are the parameters that were used in this casaguide. Note that the command will be interactive and you will have to set the cleaning boxes. | Now we proceed with the cleaning itself. For the {{clean}} task you will need to choose the right cell size, map size, etc. Below are the parameters that were used in this casaguide. Note that the command will be interactive and you will have to set the cleaning boxes. | ||
| Line 125: | Line 166: | ||
# cleaning 1 | # cleaning 1 | ||
os.system('rm -rf *CONTIN*') | os.system('rm -rf *CONTIN*') | ||
clean(vis=' | clean(vis='IRAS16293_Band9.fixed.CONT.msAVG', | ||
imagename='IRAS16293_Band9.CONTIN', | imagename='IRAS16293_Band9.fixed.CONTIN', | ||
spw='0~3',field='',phasecenter='3', | spw='0~3',field='',phasecenter='3', | ||
mode='mfs',imagermode='mosaic', | mode='mfs',imagermode='mosaic', | ||
| Line 135: | Line 176: | ||
</source> | </source> | ||
It is important to check if all fields have some emission in the model data column, if not, or very weak you would want to exclude from the gaincal below. Because the bright Source B is | It is important to check if all fields have some emission in the model data column, if not, or if it is very weak you would want to exclude from the gaincal below. Because the bright Source B is within the primary beam of all the observed fields, all fields have strong emission present in the image model. In Figure 5 you can see an example for the output from next command. | ||
[[File:IRAS16293_Band9.AVG.ms_tamp_f2_allspw.png|200px|thumb|right|'''Fig. 5.''' Time-amplitude plot for field 2. All spw are plotted.]] | [[File:IRAS16293_Band9.AVG.ms_tamp_f2_allspw.png|200px|thumb|right|'''Fig. 5.''' Time-amplitude plot for field 2. All spw are plotted.]] | ||
<source lang="python"> | <source lang="python"> | ||
# checking the model data for all the fields | # checking the model data for all the fields | ||
plotms(vis=' | plotms(vis='IRAS16293_Band9.fixed.CONT.msAVG', | ||
xaxis='time', yaxis='amp',field='',iteraxis='field', | xaxis='time', yaxis='amp',field='',iteraxis='field', | ||
spw='', avgchannel='4000',coloraxis='field', | spw='', avgchannel='4000',coloraxis='field', | ||
| Line 146: | Line 187: | ||
</source> | </source> | ||
Now we proceed with {{gaincal}} to perform self calibration (for phases) on the data. Even with minsnr=3.0 there will be failed solutions. | Now we proceed with {{gaincal}} to perform self calibration (for phases) on the data. Even with minsnr=3.0 there will be some failed solutions. | ||
<source lang="python"> | <source lang="python"> | ||
# get one scan per field | # get one scan per field | ||
os.system('rm -rf cont_pcal1') | os.system('rm -rf cont_pcal1') | ||
gaincal(vis=' | gaincal(vis='IRAS16293_Band9.fixed.CONT.msAVG',caltable='cont_pcal1',gaintype='T', | ||
refant='DV14',calmode='p',combine='',spw='',field='', | refant='DV14',calmode='p',combine='',spw='',field='', | ||
solint='inf',minsnr=3.0,minblperant=4) | solint='inf',minsnr=3.0,minblperant=4) | ||
| Line 158: | Line 199: | ||
The solutions for five antennas are shown in Figure 6, and are produced with the next {{plotcal}}. Be sure to check all the antennas. | The solutions for five antennas are shown in Figure 6, and are produced with the next {{plotcal}}. Be sure to check all the antennas. | ||
[[File: | [[File:cont_pcal1.png|200px|thumb|right|'''Fig. 6.''' Phase solutions for the first self-cal run.]] | ||
<source lang="python"> | <source lang="python"> | ||
| Line 172: | Line 213: | ||
<source lang="python"> | <source lang="python"> | ||
# application of the table | # application of the table | ||
applycal(vis=' | applycal(vis='IRAS16293_Band9.fixed.CONT.msAVG',spwmap=[],spw='', | ||
gaintable=['cont_pcal1'],calwt=F,flagbackup=F) | gaintable=['cont_pcal1'],calwt=F,flagbackup=F) | ||
</source> | </source> | ||
Now you are ready for the second cleaning. For this one we are using the mask resulting from the previous cleaning. In the next blocks of commands we will repeat this process (clean, viewer, gaincal, plotcal, applycal, clean, etc) to improve the overall quality of the image. | Now you are ready for the second cleaning. For this one we are using the mask resulting from the previous cleaning. In the next blocks of commands we will repeat this process (clean, viewer, gaincal, plotcal, applycal, clean, etc.) to improve the overall quality of the image. | ||
<source lang="python"> | <source lang="python"> | ||
# cleaning 2 | # cleaning 2 | ||
os.system('rm -rf *CONTIN_P1*') | os.system('rm -rf *CONTIN_P1*') | ||
clean(vis=' | clean(vis='IRAS16293_Band9.fixed.CONT.msAVG', | ||
imagename='IRAS16293_Band9.CONTIN_P1', | imagename='IRAS16293_Band9.fixed.CONTIN_P1', | ||
spw='0~3',field='',phasecenter='3', | spw='0~3',field='',phasecenter='3', | ||
mode='mfs',imagermode='mosaic', | mode='mfs',imagermode='mosaic', | ||
| Line 188: | Line 229: | ||
interactive=T, | interactive=T, | ||
weighting='briggs',robust=0.5, | weighting='briggs',robust=0.5, | ||
mask='IRAS16293_Band9.CONTIN.mask', | mask='IRAS16293_Band9.fixed.CONTIN.mask', | ||
niter=10000, threshold='0.3mJy') | niter=10000, threshold='0.3mJy') | ||
# visualize the image | # visualize the image | ||
imview (raster=[{'file':'IRAS16293_Band9.CONTIN.image', | imview (raster=[{'file':'IRAS16293_Band9.fixed.CONTIN.image', | ||
'range': [-0.01,0.5]}, | 'range': [-0.01,0.5]}, | ||
{'file': 'IRAS16293_Band9.CONTIN_P1.image', | {'file': 'IRAS16293_Band9.fixed.CONTIN_P1.image', | ||
'range': [-0.01,0.5]}]) | 'range': [-0.01,0.5]}]) | ||
</source> | </source> | ||
| Line 203: | Line 244: | ||
# second self-cal | # second self-cal | ||
os.system('rm -rf cont_pcal2') | os.system('rm -rf cont_pcal2') | ||
gaincal(vis=' | gaincal(vis='IRAS16293_Band9.fixed.CONT.msAVG',caltable='cont_pcal2',gaintype='T', | ||
refant='DV14',calmode='p',combine='',spw='',field='', | refant='DV14',calmode='p',combine='',spw='',field='', | ||
solint='inf',minsnr=3.0,minblperant=4) | solint='inf',minsnr=3.0,minblperant=4) | ||
| Line 213: | Line 254: | ||
# Applying results to the data | # Applying results to the data | ||
applycal(vis=' | applycal(vis='IRAS16293_Band9.fixed.CONT.msAVG',spwmap=[],spw='', | ||
gaintable=['cont_pcal2'],calwt=F,flagbackup=F) | gaintable=['cont_pcal2'],calwt=F,flagbackup=F) | ||
</source> | </source> | ||
| Line 222: | Line 263: | ||
# Cleaning 3 | # Cleaning 3 | ||
os.system('rm -rf *CONTIN_P2*') | os.system('rm -rf *CONTIN_P2*') | ||
clean(vis=' | clean(vis='IRAS16293_Band9.fixed.CONT.msAVG', | ||
imagename='IRAS16293_Band9.CONTIN_P2', | imagename='IRAS16293_Band9.fixed.CONTIN_P2', | ||
spw='0~3',field='',phasecenter='3', | spw='0~3',field='',phasecenter='3', | ||
mode='mfs',imagermode='mosaic', | mode='mfs',imagermode='mosaic', | ||
| Line 229: | Line 270: | ||
interactive=T, | interactive=T, | ||
weighting='briggs',robust=0.5, | weighting='briggs',robust=0.5, | ||
mask='IRAS16293_Band9.CONTIN.mask', | mask='IRAS16293_Band9.fixed.CONTIN.mask', | ||
niter=10000, threshold='0.3mJy') | niter=10000, threshold='0.3mJy') | ||
# to view the resulting image | # to view the resulting image | ||
imview (raster=[{'file':'IRAS16293_Band9.CONTIN_P1.image', | imview (raster=[{'file':'IRAS16293_Band9.fixed.CONTIN_P1.image', | ||
'range': [-0.01,0.5]}, | 'range': [-0.01,0.5]}, | ||
{'file': 'IRAS16293_Band9.CONTIN_P2.image', | {'file': 'IRAS16293_Band9.fixed.CONTIN_P2.image', | ||
'range': [-0.01,0.5]}]) | 'range': [-0.01,0.5]}]) | ||
| Line 241: | Line 282: | ||
# solint phase self-cal. For now, moving on to amplitude. | # solint phase self-cal. For now, moving on to amplitude. | ||
</source> | </source> | ||
[[File:cont_apcal.png|200px|thumb|right|'''Fig. 6.''' Amplitude self-calibration solutions.]] | |||
Next iteration with gaincal, now using amplitude and phase for "calmode". | Next iteration with gaincal, now using amplitude and phase for "calmode". | ||
| Line 247: | Line 290: | ||
# For amplitude self-cal we only want to get one solution per spw, per dataset | # For amplitude self-cal we only want to get one solution per spw, per dataset | ||
os.system('rm -rf cont_apcal') | os.system('rm -rf cont_apcal') | ||
gaincal(vis=' | gaincal(vis='IRAS16293_Band9.fixed.CONT.msAVG',caltable='cont_apcal',gaintype='T', | ||
refant='DV14',calmode='ap',combine='scan,field',spw='',field='', | refant='DV14',calmode='ap',combine='scan,field',spw='',field='', | ||
solint='8000s',minsnr=3.0,minblperant=4,gaintable='cont_pcal2') | solint='8000s',minsnr=3.0,minblperant=4,gaintable='cont_pcal2') | ||
| Line 257: | Line 300: | ||
# and applying to the data | # and applying to the data | ||
applycal(vis=' | applycal(vis='IRAS16293_Band9.fixed.CONT.msAVG',spwmap=[],spw='', | ||
gaintable=['cont_pcal2','cont_apcal'],calwt=F,flagbackup=F) | gaintable=['cont_pcal2','cont_apcal'],calwt=F,flagbackup=F) | ||
| Line 265: | Line 308: | ||
</source> | </source> | ||
Continuing the | Continuing the final cleaning step. | ||
<source lang="python"> | <source lang="python"> | ||
# Cleaning 4 | # Cleaning 4 | ||
os.system('rm -rf *CONTIN_AP*') | os.system('rm -rf *CONTIN_AP*') | ||
clean(vis=' | clean(vis='IRAS16293_Band9.fixed.CONT.msAVG', | ||
imagename='IRAS16293_Band9.CONTIN_AP', | imagename='IRAS16293_Band9.fixed.CONTIN_AP', | ||
spw='0~3',field='',phasecenter='3', | spw='0~3',field='',phasecenter='3', | ||
mode='mfs',imagermode='mosaic', | mode='mfs',imagermode='mosaic', | ||
| Line 277: | Line 320: | ||
interactive=T, | interactive=T, | ||
weighting='briggs',robust=0.5, | weighting='briggs',robust=0.5, | ||
mask='IRAS16293_Band9.CONTIN_P2.mask', | mask='IRAS16293_Band9.fixed.CONTIN_P2.mask', | ||
niter=10000, threshold='0.3mJy') | niter=10000, threshold='0.3mJy') | ||
# and to view the image | # and to view the image | ||
imview (raster=[{'file':'IRAS16293_Band9.CONTIN_P2.image', | imview (raster=[{'file':'IRAS16293_Band9.fixed.CONTIN_P2.image', | ||
'range': [-0.01,0.5]}, | 'range': [-0.01,0.5]}, | ||
{'file': 'IRAS16293_Band9.CONTIN_AP.image', | {'file': 'IRAS16293_Band9.fixed.CONTIN_AP.image', | ||
'range': [-0.01,0.5]}]) | 'range': [-0.01,0.5]}]) | ||
</source> | </source> | ||
We use now {{impbcor}} to produce an image corrected by the primary beam. | We use now {{impbcor}} to produce an image corrected by the primary beam response. We are | ||
outputting a sub-region of the cube because the area covered is large compared to | |||
the area of interest, and we want to keep the file size of the result as small | |||
as possible. | |||
<source lang="python"> | <source lang="python"> | ||
# | # correcting for primary beam | ||
impbcor(imagename='IRAS16293_Band9.CONTIN_AP.image', | impbcor(imagename='IRAS16293_Band9.fixed.CONTIN_AP.image', | ||
pbimage='IRAS16293_Band9.CONTIN_AP.flux', | pbimage='IRAS16293_Band9.fixed.CONTIN_AP.flux', | ||
outfile='IRAS16293_Band9.CONTIN_AP.image.pbcor.subim', | outfile='IRAS16293_Band9.fixed.CONTIN_AP.image.pbcor.subim', | ||
box=' | box='150,104,345,299') | ||
# and checking the image | # and checking the image | ||
imview (raster=[{'file':'IRAS16293_Band9.CONTIN_AP.image', | imview (raster=[{'file': 'IRAS16293_Band9.fixed.CONTIN_AP.image.pbcor.subim', | ||
'range': [-0.01, | 'range': [-0.01,1.5],'colorwedge':True}], | ||
out='IRAS16293_Band9.fixed.CONTIN_AP.image.pbcor.subim.png') | |||
</source> | </source> | ||
In Figure 7 we show the final image of | In Figure 7 we show the final image of the self-calibration process. Source A is the weaker source to the SE and Source B is the stronger source to the NW. The rms noise is about 18 mJy/beam, and the restoring beam is 0.29 x 0.17 arcsec, PA=-71 deg with robust=0.5 weighting. The peak intensities for the two components are about 3.19 (B) and 1.07 (A) Jy/beam, while the flux densities are about 11.9 (B) and 9 (A) Jy. | ||
[[File:IRAS16293_Band9.CONTIN_AP.image.pbcor.subim_c.png|200px|thumb|right|'''Fig. 7.''' Continuum image resulted from the complete process of self calibration for the mosaic of IRAS16293 Band 9.]] | [[File:IRAS16293_Band9.fixed.CONTIN_AP.image.pbcor.subim_c.png|200px|thumb|right|'''Fig. 7.''' Continuum image resulted from the complete process of self calibration for the mosaic of IRAS16293 Band 9.]] | ||
Now you can export your final image in | Now you can export your final image in FITS format, so you can analyze it in other software packages if desired. | ||
<source lang="python"> | <source lang="python"> | ||
exportfits(imagename='IRAS16293_Band9.CONTIN_AP.image.pbcor.subim', | exportfits(imagename='IRAS16293_Band9.fixed.CONTIN_AP.image.pbcor.subim', | ||
fitsimage='IRAS16293_Band9.CONTIN_AP.image.pbcor.subim.fits') | fitsimage='IRAS16293_Band9.fixed.CONTIN_AP.image.pbcor.subim.fits') | ||
</source> | </source> | ||
== Spectral | == Spectral Line Imaging == | ||
<source lang="python"> | <source lang="python"> | ||
# Restore the flags made for the continuum imaging | # Restore the flags made for the continuum imaging | ||
flagmanager(vis='IRAS16293_Band9.rebin.ms',mode='restore',versionname='Original') | flagmanager(vis='IRAS16293_Band9.fixed.rebin.ms',mode='restore',versionname='Original') | ||
</source> | </source> | ||
| Line 327: | Line 372: | ||
# Subtract the continuum from the line data | # Subtract the continuum from the line data | ||
contspw='0:20~1700;2500~3820,1:20~400;500~1100;1900~3000;3450~3800,2:20~1700;2200~3100,3:800~1600;2900~3300' | contspw='0:20~1700;2500~3820,1:20~400;500~1100;1900~3000;3450~3800,2:20~1700;2200~3100,3:800~1600;2900~3300' | ||
uvcontsub(vis='IRAS16293_Band9.rebin.ms',fitspw=contspw,fitorder=1,combine='spw') | uvcontsub(vis='IRAS16293_Band9.fixed.rebin.ms',fitspw=contspw,fitorder=1,combine='spw') | ||
</source> | </source> | ||
| Line 334: | Line 379: | ||
<source lang="python"> | <source lang="python"> | ||
# Apply final self-calibration to the line data | # Apply final self-calibration to the line data | ||
applycal(vis='IRAS16293_Band9.rebin.ms.contsub',spwmap=[],spw='', | applycal(vis='IRAS16293_Band9.fixed.rebin.ms.contsub',spwmap=[],spw='', | ||
gaintable=['cont_pcal2','cont_apcal'],calwt=F,flagbackup=F) | gaintable=['cont_pcal2','cont_apcal'],calwt=F,flagbackup=F) | ||
</source> | </source> | ||
| Line 340: | Line 385: | ||
To make your work more effective we recommend you to clean cubes 1/2 spw at a time to make the image cubes a manageable size. Also, average every 2 channels since this is the native velocity resolution. | To make your work more effective we recommend you to clean cubes 1/2 spw at a time to make the image cubes a manageable size. Also, average every 2 channels since this is the native velocity resolution. | ||
Some notes for this sections: the threshold was chosen after doing a couple of smaller test images. This is | Some notes for this sections: the threshold was chosen after doing a couple of smaller test images. This is about 3 sigma in channels with weak line emission. The mask was also chosen from a few test images. With this set, there is no need to do interactive cleaning. Note however, that this clean mask does not do a good job on the extended emission of CO(6-5). | ||
about 3 sigma in channels with weak line emission. The mask was also chosen from a few test images. With this set, there is no need to do | |||
=== | === Clean each spw === | ||
First clean the first half of each spw. | |||
<source lang="python"> | <source lang="python"> | ||
os.system('rm -rf * | # In CASA | ||
clean(vis='IRAS16293_Band9.rebin.ms.contsub', | os.system('rm -rf *spw*a*') | ||
imagename='IRAS16293_Band9.ms. | spw=[0,1,2,3] | ||
spw= | for i in spw: | ||
clean(vis='IRAS16293_Band9.fixed.rebin.ms.contsub', | |||
imagename='IRAS16293_Band9.fixed.ms.spw'+str(i)+'a', | |||
spw=str(i),field='',phasecenter='3', | |||
mode='channel',outframe='LSRK', | mode='channel',outframe='LSRK', | ||
nchan=950,start=20,width=2, | nchan=950,start=20,width=2, | ||
| Line 358: | Line 404: | ||
imsize=500,cell='0.04arcsec',minpb=0.25, | imsize=500,cell='0.04arcsec',minpb=0.25, | ||
interactive=F, | interactive=F, | ||
mask=['circle[[16h32m22. | mask=['circle[[16h32m22.61s,-24d28m32.49s ], 0.75arcsec]','circle[[16h32m22.86s,-24d28m36.57s ], 1.0arcsec]'], | ||
weighting='briggs',robust=0.5, | weighting='briggs',robust=0.5, | ||
niter=200, threshold='220.0mJy',usescratch= | niter=200, threshold='220.0mJy',usescratch=F) | ||
</source> | </source> | ||
Next clean the 2nd half of each spw. | |||
<source lang="python"> | <source lang="python"> | ||
os.system('rm -rf * | # In CASA | ||
clean(vis='IRAS16293_Band9.rebin.ms.contsub', | os.system('rm -rf *spw*b*') | ||
imagename='IRAS16293_Band9.ms. | spw=[0,1,2,3] | ||
spw= | for i in spw: | ||
clean(vis='IRAS16293_Band9.fixed.rebin.ms.contsub', | |||
imagename='IRAS16293_Band9.fixed.ms.spw'+str(i)+'b', | |||
spw=str(i),field='',phasecenter='3', | |||
mode='channel',outframe='LSRK', | mode='channel',outframe='LSRK', | ||
nchan=950,start=1918,width=2, | nchan=950,start=1918,width=2, | ||
| Line 375: | Line 424: | ||
imsize=500,cell='0.04arcsec',minpb=0.25, | imsize=500,cell='0.04arcsec',minpb=0.25, | ||
interactive=F, | interactive=F, | ||
mask=['circle[[16h32m22. | mask=['circle[[16h32m22.61s,-24d28m32.49s ], 0.75arcsec]','circle[[16h32m22.86s,-24d28m36.57s ], 1.0arcsec]'], | ||
weighting='briggs',robust=0.5, | weighting='briggs',robust=0.5, | ||
niter=200, threshold='220.0mJy',usescratch= | niter=200, threshold='220.0mJy',usescratch=F) | ||
</source> | </source> | ||
=== Create a velocity cube for H13CN (J=8-7) === | |||
= | In this section we provide an example for how to create a Doppler corrected velocity cube for one of the interesting lines in the data: H13CN (J=8-7). | ||
A good reference for finding rest frequencies is http://www.splatalogue.net/. Another good reference for Band 9 specifically is the 450 micron CSO survey of Orion: http://iopscience.iop.org/0067-0049/132/2/281/ | |||
First, determine the velocity range for H13CN 690.55207 GHz | |||
First | |||
with the help of the next {{plotms}} | with the help of the next {{plotms}} | ||
<source lang="python"> | <source lang="python"> | ||
plotms(vis='IRAS16293_Band9.ms.contsub', | plotms(vis='IRAS16293_Band9.fixed.rebin.ms.contsub', | ||
xaxis='velocity', yaxis='amp',field='', | xaxis='velocity', yaxis='amp',field='', | ||
spw='1', avgtime='1e8',avgscan=T,coloraxis='field', | spw='1', avgtime='1e8',avgscan=T,coloraxis='field', | ||
| Line 514: | Line 450: | ||
cleaning when noise like residuals remain. Number of iterations | cleaning when noise like residuals remain. Number of iterations | ||
between showing interactive window can be controlled within the | between showing interactive window can be controlled within the | ||
interactive viewer. | interactive viewer. We used about 400 iterations. | ||
<source lang="python"> | <source lang="python"> | ||
os.system('rm -rf IRAS16293_Band9.H13CN_8_7*') | os.system('rm -rf IRAS16293_Band9.fixed.H13CN_8_7*') | ||
clean(vis='IRAS16293_Band9.ms.contsub', | clean(vis='IRAS16293_Band9.fixed.rebin.ms.contsub', | ||
imagename='IRAS16293_Band9.H13CN_8_7', | imagename='IRAS16293_Band9.fixed.H13CN_8_7', | ||
spw='1',field='',phasecenter='3', | spw='1',field='',phasecenter='3', | ||
mode='velocity',outframe='LSRK',restfreq='690.55207GHz', | mode='velocity',outframe='LSRK',restfreq='690.55207GHz', | ||
| Line 527: | Line 463: | ||
interactive=T, | interactive=T, | ||
weighting='briggs',robust=0.5, | weighting='briggs',robust=0.5, | ||
niter= | niter=500, threshold='220.0mJy') | ||
</source> | </source> | ||
| Line 536: | Line 472: | ||
spw=[0,1,2,3] | spw=[0,1,2,3] | ||
for i in spw: | for i in spw: | ||
impbcor(imagename='IRAS16293_Band9.ms.spw'+str(i)+'a.image', | impbcor(imagename='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'a.image', | ||
pbimage='IRAS16293_Band9.ms.spw'+str(i)+'a.flux', | pbimage='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'a.flux', | ||
outfile='IRAS16293_Band9.ms.spw'+str(i)+'a.image.pbcor.subim', | outfile='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'a.image.pbcor.subim', | ||
box=' | box='150,104,345,299') | ||
spw=[0,1,2,3] | spw=[0,1,2,3] | ||
for i in spw: | for i in spw: | ||
impbcor(imagename='IRAS16293_Band9.ms.spw'+str(i)+'b.image', | impbcor(imagename='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'b.image', | ||
pbimage='IRAS16293_Band9.ms.spw'+str(i)+'b.flux', | pbimage='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'b.flux', | ||
outfile='IRAS16293_Band9.ms.spw'+str(i)+'b.image.pbcor.subim', | outfile='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'b.image.pbcor.subim', | ||
box=' | box='150,104,345,299') | ||
impbcor(imagename='IRAS16293_Band9.H13CN_8_7.image', | impbcor(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image', | ||
pbimage='IRAS16293_Band9.H13CN_8_7.flux', | pbimage='IRAS16293_Band9.fixed.H13CN_8_7.flux', | ||
outfile='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim', | outfile='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim', | ||
box=' | box='150,104,345,299') | ||
</source> | </source> | ||
To | To enable analysis in other software packages, you can export the data from CASA to FITS format using {{exportfits}} | ||
<source lang="python"> | <source lang="python"> | ||
spw=[0,1,2,3] | spw=[0,1,2,3] | ||
for i in spw: | for i in spw: | ||
exportfits(imagename='IRAS16293_Band9.ms.spw'+str(i)+'a.image.pbcor.subim', | exportfits(imagename='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'a.image.pbcor.subim', | ||
fitsimage='IRAS16293_Band9.ms.spw'+str(i)+'a.image.pbcor.subim.fits') | fitsimage='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'a.image.pbcor.subim.fits') | ||
spw=[0,1,2,3] | spw=[0,1,2,3] | ||
for i in spw: | for i in spw: | ||
exportfits(imagename='IRAS16293_Band9.ms.spw'+str(i)+'b.image.pbcor.subim', | exportfits(imagename='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'b.image.pbcor.subim', | ||
fitsimage='IRAS16293_Band9.ms.spw'+str(i)+'b.image.pbcor.subim.fits') | fitsimage='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'b.image.pbcor.subim.fits') | ||
exportfits(imagename='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim', | exportfits(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim', | ||
fitsimage='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.fits') | fitsimage='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.fits') | ||
</source> | </source> | ||
=== | === Moment Maps for H13CN (J=8-7) === | ||
To proceed with the moment maps generation, you will need to determine the channel range and pixel range for inclusion for higher order moments. | To proceed with the moment maps generation, you will need to determine the channel range and pixel range for inclusion for higher order moments. | ||
<source lang="python"> | <source lang="python"> | ||
viewer('IRAS16293_Band9.H13CN_8_7.image.pbcor.subim') | viewer('IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim') | ||
</source> | </source> | ||
| Line 583: | Line 519: | ||
<source lang="python"> | <source lang="python"> | ||
os.system('rm -rf IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.mom0') | os.system('rm -rf IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.mom0') | ||
immoments(imagename='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim',moments=[0], | immoments(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim',moments=[0], | ||
outfile='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.mom0', | outfile='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.mom0', | ||
chans='10~44') | chans='10~44') | ||
</source> | </source> | ||
which you can see in Figure 8. produce that figure run the | which you can see in Figure 8. To produce that figure run the following command | ||
[[File:IRAS16293_Band9.H13CN_8_7.mom0.png|200px|thumb|right|'''Fig. 8.''' Moment 0 map for the spectral line H13CN.]] | [[File:IRAS16293_Band9.fixed.H13CN_8_7.mom0.png|200px|thumb|right|'''Fig. 8.''' Moment 0 map for the spectral line H13CN J=8-7.]] | ||
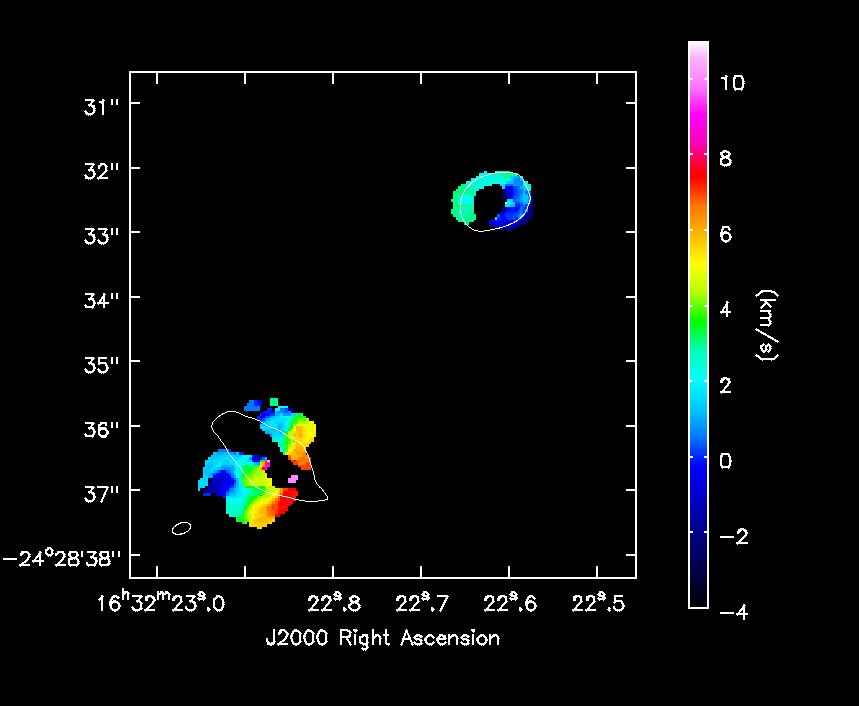
[[File:IRAS16293_Band9.fixed.H13CN_8_7.Positive_mom1.png|200px|thumb|right|'''Fig. 9.''' Positive part of the moment 1 map.]] | |||
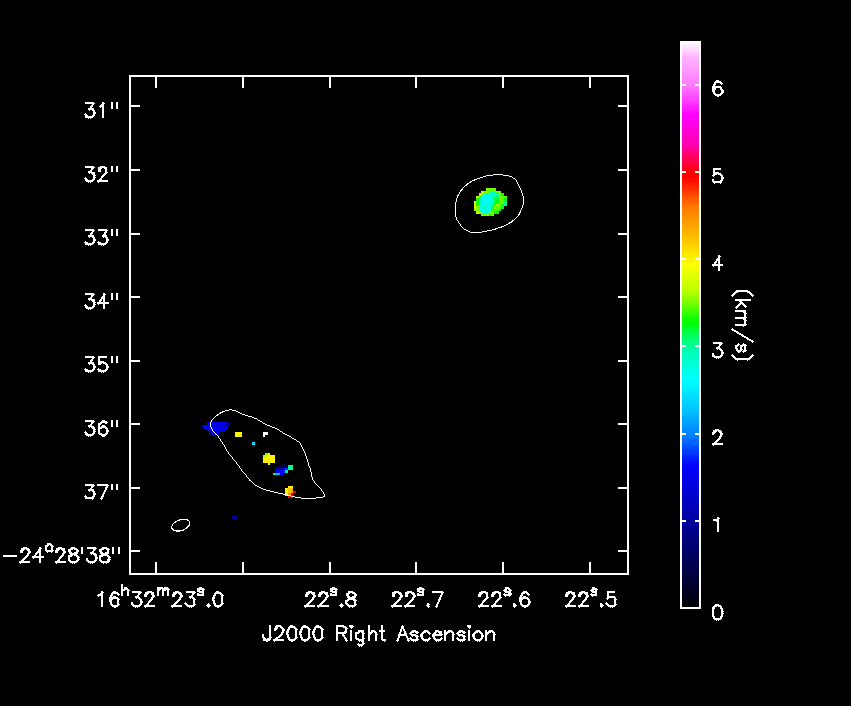
[[File:IRAS16293_Band9.fixed.H13CN_8_7.Negative_mom1.png|200px|thumb|right|'''Fig. 10.''' Negative part of the moment 1.]] | |||
<source lang="python"> | <source lang="python"> | ||
imview( raster=[ {'file':'IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.mom0', | imview( raster=[ {'file':'IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.mom0', | ||
'range': [-1., | 'range': [-1.,5.5]} ], | ||
contour={'file':'IRAS16293_Band9.CONTIN_AP.image.pbcor.subim', | contour={'file':'IRAS16293_Band9.fixed.CONTIN_AP.image.pbcor.subim', | ||
'base':0, 'unit':0.025, 'levels':[3]}, | 'base':0, 'unit':0.025, 'levels':[3]}, | ||
out='IRAS16293_Band9.H13CN_8_7.mom0.png') | out='IRAS16293_Band9.fixed.H13CN_8_7.mom0.png') | ||
</source> | </source> | ||
| Line 605: | Line 545: | ||
below. Also, we need to restrict the higher order moments to regions | below. Also, we need to restrict the higher order moments to regions | ||
of real signal. Below we demonstrate using the clean mask for this | of real signal. Below we demonstrate using the clean mask for this | ||
purpose. We must first subimage it to match the | purpose. We must first subimage it to match the primary beam corrected image. We | ||
also set the included pixels to about 5sigma. | also set the included pixels to about 5sigma. | ||
<source lang="python"> | <source lang="python"> | ||
immath(imagename='IRAS16293_Band9.H13CN_8_7.mask', | immath(imagename='IRAS16293_Band9.fixed.H13CN_8_7.mask', | ||
mode='evalexpr',expr='IM0',box=' | mode='evalexpr',expr='IM0',box='150,104,345,299', | ||
outfile='IRAS16293_Band9.H13CN_8_7.mask.subim') | outfile='IRAS16293_Band9.fixed.H13CN_8_7.mask.subim') | ||
</source> | </source> | ||
| Line 617: | Line 557: | ||
<source lang="python"> | <source lang="python"> | ||
os.system('rm -rf IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.Positive*') | os.system('rm -rf IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Positive*') | ||
immoments(imagename='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim',moments=[1,2], | immoments(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim',moments=[1,2], | ||
outfile='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.Positive', | outfile='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Positive', | ||
mask='IRAS16293_Band9.H13CN_8_7.mask.subim', | mask='IRAS16293_Band9.fixed.H13CN_8_7.mask.subim', | ||
chans='10~44',includepix=[0. | chans='10~44',includepix=[0.4,100]) | ||
os.system('rm -rf IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.Negative*') | os.system('rm -rf IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Negative*') | ||
immoments(imagename='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim',moments=[1,2], | immoments(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim',moments=[1,2], | ||
outfile='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.Negative', | outfile='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Negative', | ||
mask='IRAS16293_Band9.H13CN_8_7.mask.subim', | mask='IRAS16293_Band9.fixed.H13CN_8_7.mask.subim', | ||
chans='10~44',includepix=[-100,-0.4]) | chans='10~44',includepix=[-100,-0.4]) | ||
</source> | </source> | ||
Visualizing the maps | Visualizing the maps | ||
<source lang="python"> | <source lang="python"> | ||
imview( raster=[ {'file':'IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.Negative.weighted_coord', | imview( raster=[ {'file':'IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Negative.weighted_coord', | ||
'colorwedge':True}], | 'colorwedge':True}], | ||
contour={'file':'IRAS16293_Band9.CONTIN_AP.image.pbcor.subim', | contour={'file':'IRAS16293_Band9.fixed.CONTIN_AP.image.pbcor.subim', | ||
'base':0, 'unit':0.025, 'levels':[3]}, | 'base':0, 'unit':0.025, 'levels':[3]}, | ||
out='IRAS16293_Band9.H13CN_8_7.Negative_mom1.png') | out='IRAS16293_Band9.fixed.H13CN_8_7.Negative_mom1.png') | ||
</source> | </source> | ||
<source lang="python"> | <source lang="python"> | ||
imview( raster=[ {'file':'IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.Positive.weighted_coord', | imview( raster=[ {'file':'IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Positive.weighted_coord', | ||
'colorwedge':True}], | 'colorwedge':True}], | ||
contour={'file':'IRAS16293_Band9.CONTIN_AP.image.pbcor.subim', | contour={'file':'IRAS16293_Band9.fixed.CONTIN_AP.image.pbcor.subim', | ||
'base':0, 'unit':0.025, 'levels':[3]}, | 'base':0, 'unit':0.025, 'levels':[3]}, | ||
out='IRAS16293_Band9.H13CN_8_7.Positive_mom1.png') | out='IRAS16293_Band9.fixed.H13CN_8_7.Positive_mom1.png') | ||
</source> | </source> | ||
And finally export the | And finally export the result to FITS format | ||
<source lang="python"> | <source lang="python"> | ||
exportfits(imagename='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim', | exportfits(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim', | ||
fitsimage='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.fits') | fitsimage='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.fits') | ||
exportfits(imagename='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.mom0', | exportfits(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.mom0', | ||
fitsimage='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.mom0.fits') | fitsimage='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.mom0.fits') | ||
exportfits(imagename='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.Negative.weighted_coord', | exportfits(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Negative.weighted_coord', | ||
fitsimage='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.Negative.weighted_coord.fits') | fitsimage='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Negative.weighted_coord.fits') | ||
exportfits(imagename='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.Positive.weighted_coord', | exportfits(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Positive.weighted_coord', | ||
fitsimage='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.Positive.weighted_coord.fits') | fitsimage='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Positive.weighted_coord.fits') | ||
exportfits(imagename='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.Negative.weighted_dispersion_coord', | exportfits(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Negative.weighted_dispersion_coord', | ||
fitsimage='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.Negative.weighted_dispersion_coord.fits') | fitsimage='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Negative.weighted_dispersion_coord.fits') | ||
exportfits(imagename='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.Positive.weighted_dispersion_coord', | exportfits(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Positive.weighted_dispersion_coord', | ||
fitsimage='IRAS16293_Band9.H13CN_8_7.image.pbcor.subim.Positive.weighted_dispersion_coord.fits') | fitsimage='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Positive.weighted_dispersion_coord.fits') | ||
</source> | </source> | ||
{{Checked 3.4.0}} | {{Checked 3.4.0}} | ||
Latest revision as of 20:24, 9 October 2012
- This portion of the guide covers imaging of the fully calibrated data see: IRAS16293_Band9_-_Calibration_for_CASA_3.4.
WARNING: On June 15, 2012 the calibration guide and the final data products (calibrated science data: IRAS16293_Band9_CalibratedMS_FIXED.tgz and reference images: IRAS16293_Band9_ReferenceImages_FIXED.tgz)) were changed to correct for a 1.2" position error in the phase calibrator (1625-254). Without correction, the science images will suffer from a similar offset.
Overview
This section of the casa guide continues from IRAS16293_Band9_-_Calibration_for_CASA_3.4 . If you completed the calibration section, then you already have the final calibrated dataset. If you just started to work on this IRAS16293 B9 data, you can download the fully calibrated dataset IRAS16293_Band9_CalibratedMS from
This casa guide covers imaging of both continuum and spectral lines of IRAS16293 Band 9. This guide requires CASA 3.4. Among other things, CASA 3.4.0 contains an improved antenna primary beam model for ALMA. In this casa guide you will also need the Analysis Utils package, so make sure you have it imported into CASA.
Confirm your version of CASA
This guide has been written for CASA release 3.4.0. Please confirm your version before proceeding.
# In CASA
version = casalog.version()
print "You are using " + version
if (int(version.split()[4][1:-1]) < 19874):
print "\033[91m YOUR VERSION OF CASA IS TOO OLD FOR THIS GUIDE."
print "\033[91m PLEASE UPDATE IT BEFORE PROCEEDING."
else:
print "Your version of CASA is appropriate for this guide."
Install Analysis Utilities
Analysis Utilities (or analysisUtils for short) is a small set of Python scripts that provide a number of analysis and plotting utilities for ALMA data reduction. This guide uses a few of these utilities. They are very easy to install (just download and untar). See
http://casaguides.nrao.edu/index.php?title=Analysis_Utilities
for a full description and download instructions. If you do not wish to do this, see the CASA 3.3 version of the guide linked at the top of this page for alternative (but slow) plotting options. Analysis Utilities are updated frequently so if its been a while since you installed it, its probably worth doing it again. If you are at an ALMA site or ARC, the analysis utilities are probably already installed and up to date.
List the contents of the Data
Before we start with the imaging itself, do a listobs to the calibrated dataset, so you can check that everything is ok.
# listobs to check data
listobs(vis='IRAS16293_Band9.fixed.rebin.ms',
listfile='IRAS16293_Band9.fixed.rebin.ms.listobs',verbose=F)
You should see the next information in the .listobs file.
Fields: 7 ID Code Name RA Decl Epoch SrcId nRows 0 none IRAS16293-2422-a 16:32:22.99200 -24.28.36.0000 J2000 0 7811 1 none IRAS16293-2422-a 16:32:22.47925 -24.28.36.0000 J2000 0 7829 2 none IRAS16293-2422-a 16:32:22.73563 -24.28.36.0000 J2000 0 7829 3 none IRAS16293-2422-a 16:32:22.73563 -24.28.32.5000 J2000 0 7829 4 none IRAS16293-2422-a 16:32:22.47925 -24.28.29.0000 J2000 0 7429 5 none IRAS16293-2422-a 16:32:22.73563 -24.28.29.0000 J2000 0 7151 6 none IRAS16293-2422-a 16:32:22.99200 -24.28.29.0000 J2000 0 7151 Spectral Windows: (4 unique spectral windows and 1 unique polarization setups) SpwID #Chans Frame Ch1(MHz) ChanWid(kHz) TotBW(kHz) Corrs 0 3840 TOPO 703312.744 488.28125 1875000 XX YY 1 3840 TOPO 692237.256 -488.28125 1875000 XX YY 2 3840 TOPO 690437.256 -488.28125 1875000 XX YY 3 3840 TOPO 688437.256 -488.28125 1875000 XX YY Antennas: 13 'name'='station' ID= 0-3: 'DA41'='A003', 'DA43'='A075', 'DA47'='A026', 'DV02'='A077', ID= 4-8: 'DV03'='A137', 'DV07'='A076', 'DV09'='A046', 'DV10'='A071', ID= 9-13: 'DV12'='A011', 'DV13'='A072', 'DV14'='A025', 'DV17'='A138',
If you are using the calibrated data from the science portal, it is essential to do the following.
# In CASA
tb.open('IRAS16293_Band9.fixed.rebin.ms/POINTING', nomodify = False)
a = tb.rownumbers()
tb.removerows(a)
tb.close()
Continuum Emission Imaging
As shown in Figure 2 of IRAS16293Band9, these observations have a 7 pointing mosaic, which you can visualize with the next command (see also Figure 1).
# Make a plot of the mosaic pattern with field ids identified
au.plotMosaic(vis='IRAS16293_Band9.fixed.rebin.ms',sourceid='0',
doplot=True,figfile='mosaic_pattern.png')
You might wonder which of these pointings or fields have stronger line emission. You can determine this by running the next command. We set spw='1' which contains CO (6-5), and you can check all the plots for all the fields by just hitting the arrow for "next". Also see Figure 2 for an example of the output.
# See which mosaic fields have the brightest line emission
plotms(vis='IRAS16293_Band9.fixed.rebin.ms',
field='',xaxis='uvdist', yaxis='amp',
spw='1', avgchannel='3840',coloraxis='field',
iteraxis='field')
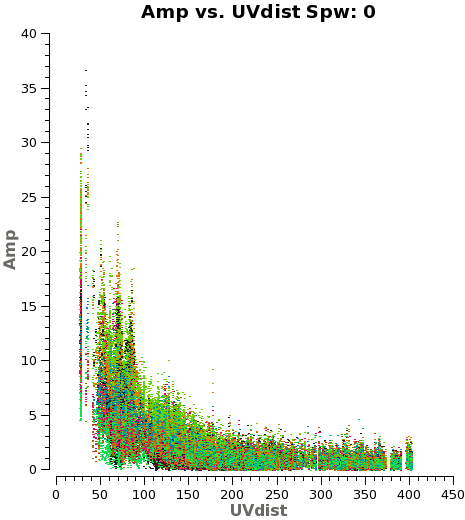
Another way to visualize the uv-amp distribution is with the next plotms. This will load all the fields or pointing into the plot, and you can change interactively between the spws using the appropriate button in plotms. See Figure 3 for a plot of spw 0.
# Iterate over spws all fields
plotms(vis='IRAS16293_Band9.fixed.rebin.ms',
field='',xaxis='uvdist', yaxis='amp',
spw='', avgchannel='3840',coloraxis='field',
iteraxis='spw')
We need to note the channels that do not present line emission. To identify them we can plot the amplitude vs frequency for all the spectral windows. Figure 4 shows the output from the next command for spw 0 (field 3).
# Select Channels for uvcontsub. Average every 8 channels for
# increased S/N.
plotms(vis='IRAS16293_Band9.fixed.rebin.ms',
xaxis='channel', yaxis='amp',field='3',
spw='', avgtime='1e8',avgscan=T,coloraxis='corr',
avgchannel='',iteraxis='spw',ydatacolumn='corrected',
xselfscale=True,yselfscale=True)
Using the plots from the previous command you can select a block of channels that are free of strong line emission. To do this in the most rigorous way, you would first need to make dirty image cubes and then examine the spectra on and near the continuum sources, because many more weaker lines will become apparent in the cubes. Here we provide you with the channel selection we used for continuum emission.
The next flagmanager command will save the flags state in case you might need to re-do the steps from this point.
flagmanager(vis='IRAS16293_Band9.fixed.rebin.ms',mode='save',versionname='Original')
Now we will flag the channels identified to have line emission, so we will be ready to proceed with the continuum imaging.
# flagging the line channels
flagdata(vis='IRAS16293_Band9.fixed.rebin.ms',
spw='0:1700~2500,1:400~500;1100~1900;3000~3450,2:1700~2200;3100~3840,3:0~800;1600~2900;3300~3840',flagbackup=F)
To help clean to be faster, you can average 200 channels, as you do not need to have a high spectral resolution for continuum imaging.
split(vis='IRAS16293_Band9.fixed.rebin.ms',
outputvis='IRAS16293_Band9.fixed.CONT.msAVG',field='',
datacolumn='data',width=200,spw='0~3:20~3819')
Clean and self-calibrate the continuum
Now we proceed with the cleaning itself. For the clean task you will need to choose the right cell size, map size, etc. Below are the parameters that were used in this casaguide. Note that the command will be interactive and you will have to set the cleaning boxes.
# cleaning 1
os.system('rm -rf *CONTIN*')
clean(vis='IRAS16293_Band9.fixed.CONT.msAVG',
imagename='IRAS16293_Band9.fixed.CONTIN',
spw='0~3',field='',phasecenter='3',
mode='mfs',imagermode='mosaic',
imsize=500,cell='0.04arcsec',minpb=0.25,
interactive=T,
weighting='briggs',robust=0.5,
niter=10000, threshold='0.3mJy',usescratch=F)
It is important to check if all fields have some emission in the model data column, if not, or if it is very weak you would want to exclude from the gaincal below. Because the bright Source B is within the primary beam of all the observed fields, all fields have strong emission present in the image model. In Figure 5 you can see an example for the output from next command.
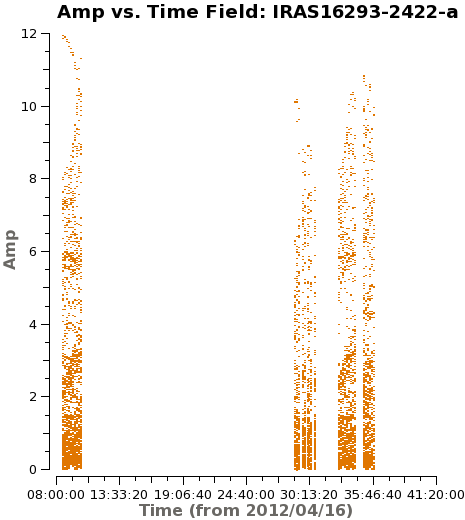
# checking the model data for all the fields
plotms(vis='IRAS16293_Band9.fixed.CONT.msAVG',
xaxis='time', yaxis='amp',field='',iteraxis='field',
spw='', avgchannel='4000',coloraxis='field',
ydatacolumn='model',xselfscale=True,yselfscale=True)
Now we proceed with gaincal to perform self calibration (for phases) on the data. Even with minsnr=3.0 there will be some failed solutions.
# get one scan per field
os.system('rm -rf cont_pcal1')
gaincal(vis='IRAS16293_Band9.fixed.CONT.msAVG',caltable='cont_pcal1',gaintype='T',
refant='DV14',calmode='p',combine='',spw='',field='',
solint='inf',minsnr=3.0,minblperant=4)
The solutions for five antennas are shown in Figure 6, and are produced with the next plotcal. Be sure to check all the antennas.

plotcal(caltable='cont_pcal1',xaxis='time',yaxis='phase',
spw='',iteration='antenna',subplot=511,plotrange=[0,0,-80,80],
antenna='',timerange='')
# The phases are remarkably stable for Band 9!
We then apply the table to the data to proceed right after with the proper cleaning, to check the improvement of the data.
# application of the table
applycal(vis='IRAS16293_Band9.fixed.CONT.msAVG',spwmap=[],spw='',
gaintable=['cont_pcal1'],calwt=F,flagbackup=F)
Now you are ready for the second cleaning. For this one we are using the mask resulting from the previous cleaning. In the next blocks of commands we will repeat this process (clean, viewer, gaincal, plotcal, applycal, clean, etc.) to improve the overall quality of the image.
# cleaning 2
os.system('rm -rf *CONTIN_P1*')
clean(vis='IRAS16293_Band9.fixed.CONT.msAVG',
imagename='IRAS16293_Band9.fixed.CONTIN_P1',
spw='0~3',field='',phasecenter='3',
mode='mfs',imagermode='mosaic',
imsize=500,cell='0.04arcsec',minpb=0.25,
interactive=T,
weighting='briggs',robust=0.5,
mask='IRAS16293_Band9.fixed.CONTIN.mask',
niter=10000, threshold='0.3mJy')
# visualize the image
imview (raster=[{'file':'IRAS16293_Band9.fixed.CONTIN.image',
'range': [-0.01,0.5]},
{'file': 'IRAS16293_Band9.fixed.CONTIN_P1.image',
'range': [-0.01,0.5]}])
Second gaincal step for self calibration.
# second self-cal
os.system('rm -rf cont_pcal2')
gaincal(vis='IRAS16293_Band9.fixed.CONT.msAVG',caltable='cont_pcal2',gaintype='T',
refant='DV14',calmode='p',combine='',spw='',field='',
solint='inf',minsnr=3.0,minblperant=4)
#Checking the table
plotcal(caltable='cont_pcal2',xaxis='time',yaxis='phase',
spw='',iteration='antenna',subplot=511,plotrange=[0,0,-80,80],
antenna='',timerange='')
# Applying results to the data
applycal(vis='IRAS16293_Band9.fixed.CONT.msAVG',spwmap=[],spw='',
gaintable=['cont_pcal2'],calwt=F,flagbackup=F)
We continue with the next cleaning.
# Cleaning 3
os.system('rm -rf *CONTIN_P2*')
clean(vis='IRAS16293_Band9.fixed.CONT.msAVG',
imagename='IRAS16293_Band9.fixed.CONTIN_P2',
spw='0~3',field='',phasecenter='3',
mode='mfs',imagermode='mosaic',
imsize=500,cell='0.04arcsec',minpb=0.25,
interactive=T,
weighting='briggs',robust=0.5,
mask='IRAS16293_Band9.fixed.CONTIN.mask',
niter=10000, threshold='0.3mJy')
# to view the resulting image
imview (raster=[{'file':'IRAS16293_Band9.fixed.CONTIN_P1.image',
'range': [-0.01,0.5]},
{'file': 'IRAS16293_Band9.fixed.CONTIN_P2.image',
'range': [-0.01,0.5]}])
# There is very little change, the bold could try another iteration using a shorter
# solint phase self-cal. For now, moving on to amplitude.
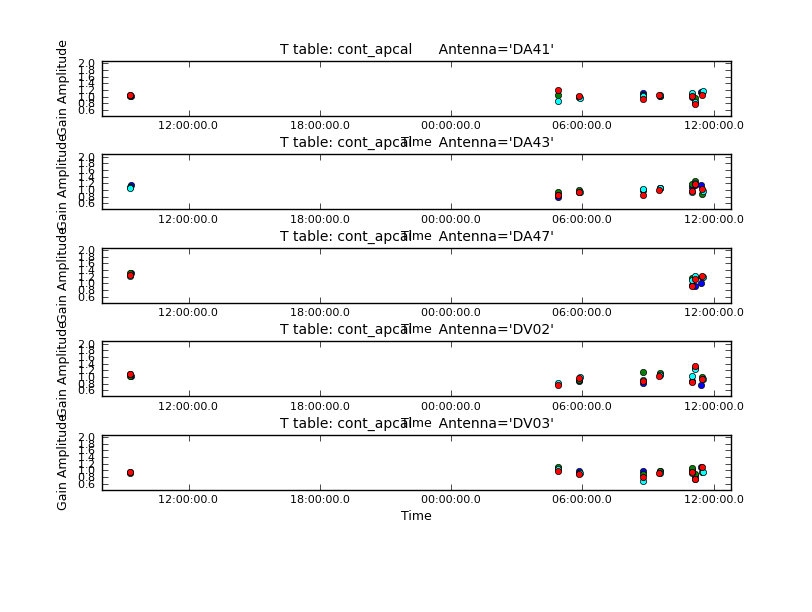
Next iteration with gaincal, now using amplitude and phase for "calmode".
# For amplitude self-cal we only want to get one solution per spw, per dataset
os.system('rm -rf cont_apcal')
gaincal(vis='IRAS16293_Band9.fixed.CONT.msAVG',caltable='cont_apcal',gaintype='T',
refant='DV14',calmode='ap',combine='scan,field',spw='',field='',
solint='8000s',minsnr=3.0,minblperant=4,gaintable='cont_pcal2')
# ploting the table
plotcal(caltable='cont_apcal',xaxis='time',yaxis='amp',
spw='',iteration='antenna',subplot=511,plotrange=[0,0,0.5,2],
antenna='',timerange='')
# and applying to the data
applycal(vis='IRAS16293_Band9.fixed.CONT.msAVG',spwmap=[],spw='',
gaintable=['cont_pcal2','cont_apcal'],calwt=F,flagbackup=F)
# A large correction is present for the 2nd dataset. This amplitude
# error was causing many of the artifacts noticeable in the previous
# images.
Continuing the final cleaning step.
# Cleaning 4
os.system('rm -rf *CONTIN_AP*')
clean(vis='IRAS16293_Band9.fixed.CONT.msAVG',
imagename='IRAS16293_Band9.fixed.CONTIN_AP',
spw='0~3',field='',phasecenter='3',
mode='mfs',imagermode='mosaic',
imsize=500,cell='0.04arcsec',minpb=0.25,
interactive=T,
weighting='briggs',robust=0.5,
mask='IRAS16293_Band9.fixed.CONTIN_P2.mask',
niter=10000, threshold='0.3mJy')
# and to view the image
imview (raster=[{'file':'IRAS16293_Band9.fixed.CONTIN_P2.image',
'range': [-0.01,0.5]},
{'file': 'IRAS16293_Band9.fixed.CONTIN_AP.image',
'range': [-0.01,0.5]}])
We use now impbcor to produce an image corrected by the primary beam response. We are outputting a sub-region of the cube because the area covered is large compared to the area of interest, and we want to keep the file size of the result as small as possible.
# correcting for primary beam
impbcor(imagename='IRAS16293_Band9.fixed.CONTIN_AP.image',
pbimage='IRAS16293_Band9.fixed.CONTIN_AP.flux',
outfile='IRAS16293_Band9.fixed.CONTIN_AP.image.pbcor.subim',
box='150,104,345,299')
# and checking the image
imview (raster=[{'file': 'IRAS16293_Band9.fixed.CONTIN_AP.image.pbcor.subim',
'range': [-0.01,1.5],'colorwedge':True}],
out='IRAS16293_Band9.fixed.CONTIN_AP.image.pbcor.subim.png')
In Figure 7 we show the final image of the self-calibration process. Source A is the weaker source to the SE and Source B is the stronger source to the NW. The rms noise is about 18 mJy/beam, and the restoring beam is 0.29 x 0.17 arcsec, PA=-71 deg with robust=0.5 weighting. The peak intensities for the two components are about 3.19 (B) and 1.07 (A) Jy/beam, while the flux densities are about 11.9 (B) and 9 (A) Jy.
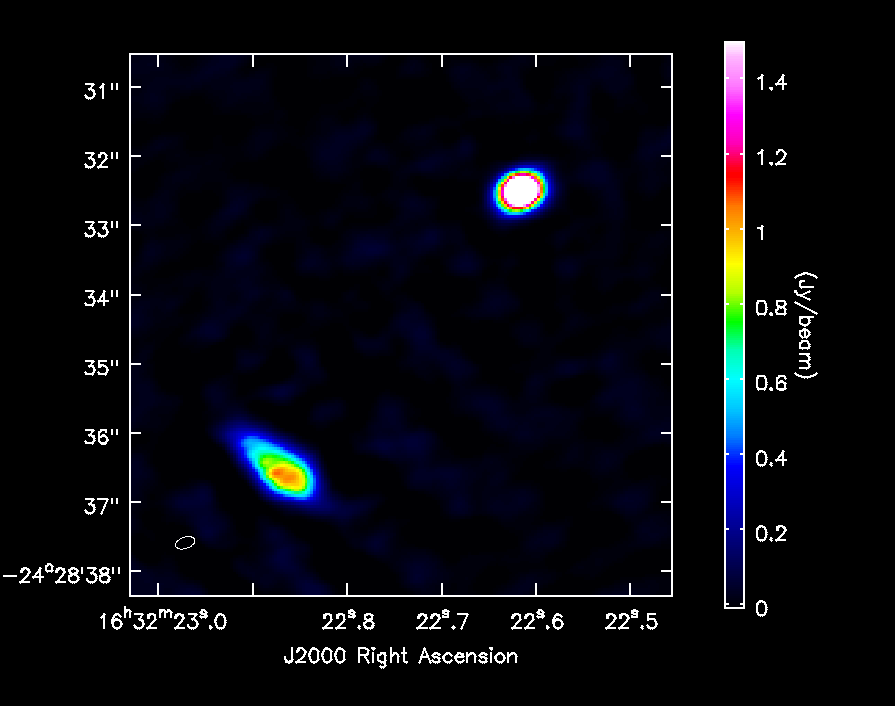
Now you can export your final image in FITS format, so you can analyze it in other software packages if desired.
exportfits(imagename='IRAS16293_Band9.fixed.CONTIN_AP.image.pbcor.subim',
fitsimage='IRAS16293_Band9.fixed.CONTIN_AP.image.pbcor.subim.fits')
Spectral Line Imaging
# Restore the flags made for the continuum imaging
flagmanager(vis='IRAS16293_Band9.fixed.rebin.ms',mode='restore',versionname='Original')
Then we need to subtract the continuum, using the line free channels
# Subtract the continuum from the line data
contspw='0:20~1700;2500~3820,1:20~400;500~1100;1900~3000;3450~3800,2:20~1700;2200~3100,3:800~1600;2900~3300'
uvcontsub(vis='IRAS16293_Band9.fixed.rebin.ms',fitspw=contspw,fitorder=1,combine='spw')
Before we commence with the cleaning we also need to apply the tables we produced in the self calibration steps.
# Apply final self-calibration to the line data
applycal(vis='IRAS16293_Band9.fixed.rebin.ms.contsub',spwmap=[],spw='',
gaintable=['cont_pcal2','cont_apcal'],calwt=F,flagbackup=F)
To make your work more effective we recommend you to clean cubes 1/2 spw at a time to make the image cubes a manageable size. Also, average every 2 channels since this is the native velocity resolution.
Some notes for this sections: the threshold was chosen after doing a couple of smaller test images. This is about 3 sigma in channels with weak line emission. The mask was also chosen from a few test images. With this set, there is no need to do interactive cleaning. Note however, that this clean mask does not do a good job on the extended emission of CO(6-5).
Clean each spw
First clean the first half of each spw.
# In CASA
os.system('rm -rf *spw*a*')
spw=[0,1,2,3]
for i in spw:
clean(vis='IRAS16293_Band9.fixed.rebin.ms.contsub',
imagename='IRAS16293_Band9.fixed.ms.spw'+str(i)+'a',
spw=str(i),field='',phasecenter='3',
mode='channel',outframe='LSRK',
nchan=950,start=20,width=2,
imagermode='mosaic',
imsize=500,cell='0.04arcsec',minpb=0.25,
interactive=F,
mask=['circle[[16h32m22.61s,-24d28m32.49s ], 0.75arcsec]','circle[[16h32m22.86s,-24d28m36.57s ], 1.0arcsec]'],
weighting='briggs',robust=0.5,
niter=200, threshold='220.0mJy',usescratch=F)
Next clean the 2nd half of each spw.
# In CASA
os.system('rm -rf *spw*b*')
spw=[0,1,2,3]
for i in spw:
clean(vis='IRAS16293_Band9.fixed.rebin.ms.contsub',
imagename='IRAS16293_Band9.fixed.ms.spw'+str(i)+'b',
spw=str(i),field='',phasecenter='3',
mode='channel',outframe='LSRK',
nchan=950,start=1918,width=2,
imagermode='mosaic',
imsize=500,cell='0.04arcsec',minpb=0.25,
interactive=F,
mask=['circle[[16h32m22.61s,-24d28m32.49s ], 0.75arcsec]','circle[[16h32m22.86s,-24d28m36.57s ], 1.0arcsec]'],
weighting='briggs',robust=0.5,
niter=200, threshold='220.0mJy',usescratch=F)
Create a velocity cube for H13CN (J=8-7)
In this section we provide an example for how to create a Doppler corrected velocity cube for one of the interesting lines in the data: H13CN (J=8-7).
A good reference for finding rest frequencies is http://www.splatalogue.net/. Another good reference for Band 9 specifically is the 450 micron CSO survey of Orion: http://iopscience.iop.org/0067-0049/132/2/281/
First, determine the velocity range for H13CN 690.55207 GHz with the help of the next plotms
plotms(vis='IRAS16293_Band9.fixed.rebin.ms.contsub',
xaxis='velocity', yaxis='amp',field='',
spw='1', avgtime='1e8',avgscan=T,coloraxis='field',
transform=True,freqframe='LSRK',restfreq='690.55207GHz',
avgchannel='8',
ydatacolumn='corrected',plotfile='Vel_H13CN.png')
Now do an interactive clean, making clean boxes for the emission. Stop cleaning when noise like residuals remain. Number of iterations between showing interactive window can be controlled within the interactive viewer. We used about 400 iterations.
os.system('rm -rf IRAS16293_Band9.fixed.H13CN_8_7*')
clean(vis='IRAS16293_Band9.fixed.rebin.ms.contsub',
imagename='IRAS16293_Band9.fixed.H13CN_8_7',
spw='1',field='',phasecenter='3',
mode='velocity',outframe='LSRK',restfreq='690.55207GHz',
nchan=60,start='-10km/s',width='0.5km/s',
imagermode='mosaic',
imsize=500,cell='0.04arcsec',minpb=0.25,
interactive=T,
weighting='briggs',robust=0.5,
niter=500, threshold='220.0mJy')
You need to correct for the primary beam attenuation and subimage the cubes to a smaller more manageable size since most of the mosaic is empty.
spw=[0,1,2,3]
for i in spw:
impbcor(imagename='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'a.image',
pbimage='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'a.flux',
outfile='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'a.image.pbcor.subim',
box='150,104,345,299')
spw=[0,1,2,3]
for i in spw:
impbcor(imagename='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'b.image',
pbimage='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'b.flux',
outfile='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'b.image.pbcor.subim',
box='150,104,345,299')
impbcor(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image',
pbimage='IRAS16293_Band9.fixed.H13CN_8_7.flux',
outfile='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim',
box='150,104,345,299')
To enable analysis in other software packages, you can export the data from CASA to FITS format using exportfits
spw=[0,1,2,3]
for i in spw:
exportfits(imagename='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'a.image.pbcor.subim',
fitsimage='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'a.image.pbcor.subim.fits')
spw=[0,1,2,3]
for i in spw:
exportfits(imagename='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'b.image.pbcor.subim',
fitsimage='IRAS16293_Band9.fixed.rebin.ms.spw'+str(i)+'b.image.pbcor.subim.fits')
exportfits(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim',
fitsimage='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.fits')
Moment Maps for H13CN (J=8-7)
To proceed with the moment maps generation, you will need to determine the channel range and pixel range for inclusion for higher order moments.
viewer('IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim')
Generation of moment 0
os.system('rm -rf IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.mom0')
immoments(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim',moments=[0],
outfile='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.mom0',
chans='10~44')
which you can see in Figure 8. To produce that figure run the following command
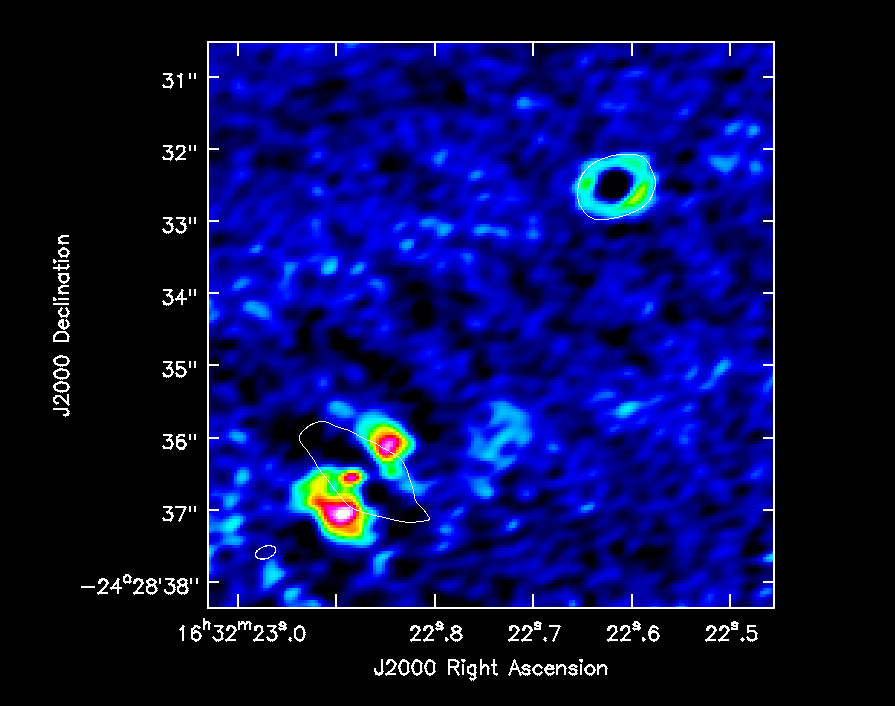
imview( raster=[ {'file':'IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.mom0',
'range': [-1.,5.5]} ],
contour={'file':'IRAS16293_Band9.fixed.CONTIN_AP.image.pbcor.subim',
'base':0, 'unit':0.025, 'levels':[3]},
out='IRAS16293_Band9.fixed.H13CN_8_7.mom0.png')
One cannot make higher order moments that simultaneously include both positive and negative emission. So we separate them below. Also, we need to restrict the higher order moments to regions of real signal. Below we demonstrate using the clean mask for this purpose. We must first subimage it to match the primary beam corrected image. We also set the included pixels to about 5sigma.
immath(imagename='IRAS16293_Band9.fixed.H13CN_8_7.mask',
mode='evalexpr',expr='IM0',box='150,104,345,299',
outfile='IRAS16293_Band9.fixed.H13CN_8_7.mask.subim')
Producing moment 1 and 2
os.system('rm -rf IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Positive*')
immoments(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim',moments=[1,2],
outfile='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Positive',
mask='IRAS16293_Band9.fixed.H13CN_8_7.mask.subim',
chans='10~44',includepix=[0.4,100])
os.system('rm -rf IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Negative*')
immoments(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim',moments=[1,2],
outfile='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Negative',
mask='IRAS16293_Band9.fixed.H13CN_8_7.mask.subim',
chans='10~44',includepix=[-100,-0.4])
Visualizing the maps
imview( raster=[ {'file':'IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Negative.weighted_coord',
'colorwedge':True}],
contour={'file':'IRAS16293_Band9.fixed.CONTIN_AP.image.pbcor.subim',
'base':0, 'unit':0.025, 'levels':[3]},
out='IRAS16293_Band9.fixed.H13CN_8_7.Negative_mom1.png')
imview( raster=[ {'file':'IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Positive.weighted_coord',
'colorwedge':True}],
contour={'file':'IRAS16293_Band9.fixed.CONTIN_AP.image.pbcor.subim',
'base':0, 'unit':0.025, 'levels':[3]},
out='IRAS16293_Band9.fixed.H13CN_8_7.Positive_mom1.png')
And finally export the result to FITS format
exportfits(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim',
fitsimage='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.fits')
exportfits(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.mom0',
fitsimage='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.mom0.fits')
exportfits(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Negative.weighted_coord',
fitsimage='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Negative.weighted_coord.fits')
exportfits(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Positive.weighted_coord',
fitsimage='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Positive.weighted_coord.fits')
exportfits(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Negative.weighted_dispersion_coord',
fitsimage='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Negative.weighted_dispersion_coord.fits')
exportfits(imagename='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Positive.weighted_dispersion_coord',
fitsimage='IRAS16293_Band9.fixed.H13CN_8_7.image.pbcor.subim.Positive.weighted_dispersion_coord.fits')
Last checked on CASA Version 3.4.0.