Imaging Prep: Difference between revisions
| Line 52: | Line 52: | ||
<source lang="python"> | <source lang="python"> | ||
# or in CASA | # or in CASA | ||
For i in glob.glob( | For i in glob.glob('*.tar'): | ||
os.system( | os.system('tar -xvf %s' % (i)) | ||
</source> | </source> | ||
Revision as of 11:56, 11 December 2019
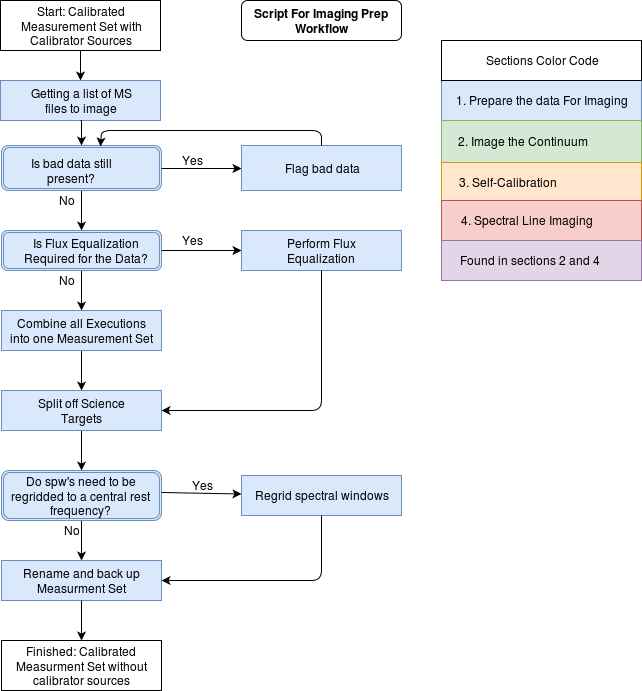
Imaging Prep Workflow
Commands for this page can be found in scriptForImagingPrep_template.py available on github.
Use the Imaging Prep workflow to determine which sections of this guide are applicable to your dataset.
Splitting off the calibrated data (Pipeline Calibration)
If your measurement set (ms) is pipeline calibrated, use this step to split off the calibrated data. After restoring the calibration using scriptForPI.py, you should have *.ms files which contain the calibrated data in the CORRECTED column. If you already have files with the name *.ms.split.cal, you can proceed to the next step. This block of code will split off the science spws for the calibrators and science target.
# in CASA
import glob
vislist = glob.glob('*[!_t].ms') # match full ms, not target.ms
for myvis in vislist:
msmd.open(myvis)
targetspws = msmd.spwsforintent('OBSERVE_TARGET*')
sciencespws = []
for myspw in targetspws:
if msmd.nchan(myspw)>4:
sciencespws.append(myspw)
sciencespws = ','.join(map(str,sciencespws))
msmd.close()
split(vis=myvis,outputvis=myvis+'.split.cal',spw=sciencespws)
Get a list of ms files to image
You should now have *.ms.split.cal files for each execution no matter how your dataset was calibrated. Run the following commands to grab the names of all files in the directory that have the extension ‘.ms.split.cal’ and place them into a list so you can easily check the calibration and concatenate the data further on in the script.
# in CASA
import glob
vislist=glob.glob('*.ms.split.cal')
Review the Calibration
<figure id="Weblog_Home.png">
</figure> Once the list of ms files have been collected, review the calibration to make sure it appears reasonable. A summary of the calibration are in the qa directory of your delivered package. If the dataset was pipeline calibrated, download and untar the pipeline weblog in the qa directory by using <figure id="Weblog_MS_Overview.png">
</figure>
# in bash
for i in $(ls *.tar); do tar -xvf $i; done
# or in CASA
For i in glob.glob('*.tar'):
os.system('tar -xvf %s' % (i))
Once this is done, look within the /html directory for the index.html file. This contains the html code for displaying the contents of the qa directory in a web browser. The command below opens the main page of the weblog in firefox.
firefox index.html
<figure id="Weblog_By_Task.png">
</figure> The standard opening page has three main sections: observation overview, pipeline summary, and observation summary. This is shown in Figure 1.
From the opening page, you can navigate to view the calibration By Topic and By Task and open observation specific information by clicking on the ms name. Figure 2 shows an example of an ms summary page.
Figure 3 shows the By Task view. Each task can be opened to view important calibration plots, tables, and flagging statistics.
If the data was manually calibrated, you can review the calibration with the *.png and *.txt files in the qa directory. These display helpful plots and statistics about the calibration.
In addition, the tasks plotms and plotcal are useful if you need to explore the data beyond the plots that the calibration pipeline or QA2 process generates. The TW Hydra guide gives some nice examples showing how to use these tasks.
Flag bad data (optional)
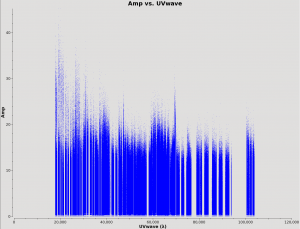
This section uses plotms to manually review the uvwave vs amp and channel vs amp plots of all field and spectral windows in the MS file. Outliers seen in the calibrator fields of these plots can be flagged either in the flagdata() command below or interactively within the GUI of plotms().
Save the original flags of each ms prior to flagging any additional data. Without this step, it will be very difficult if you need to remove flags generated later on.
# in CASA
for vis in vislist:
flagmanager(vis=vis,
mode='save',
versionname='original_flags')
The plotms task can be used to find bad data by iterating through fields and spws for the *.ms.split.cal files in the calibrated directory. You can also use the GUI interface to iterate through spectral windows (spws). Figure 4 shows an example of TW Hydra science spw 2.
# in CASA
fieldlist = ['3'] # change to list of science data fields to inspect
spwlist = ['1'] # change to list of science spws to inspect
for vis in vislist:
for field in fieldlist:
for spw in spwlist:
plotms(vis=vis,xaxis='uvwave',yaxis='amp',avgtime='3e8',
field=field,spw=spw)
raw_input("push enter to continue")
plotms(vis=vis,xaxis='chan',yaxis='amp',avgtime='3e8',
field=field,spw=spw)
raw_input("push enter to continue")
Use flagdata to flag bad antennas, spws, timeranges, etc. You can use the locate buttons in plotms and plotcal to identify what needs to be flagged. Locate will print the information about the selected points into the casalogger. Note that flagging in plotms is generally not recommended because a plotms crash will lose all flagging information.
To flag data, modify the flagdata command with the ms you want to flag and the flagging criteria. Flagging criteria can be any antenna, spw, timerange, etc. Once the data is flagged, check the flagging using plotms. The flagged data points should not appear on the plot. You can also plot the flagged data points using the "display" table on the left hand side of the plotms GUI. <figure id="TWHydra_corrected_Spw2.png">
</figure>
# in CASA
flagdata(vis='',mode='manual',action='apply',flagbackup=False)
If you need to restore original flags, use the following command. You will need to update the ms name in the vis parameter.
# in CASA
flagmanager(vis='',mode='restore',versionname='original_flags')
Flux Equalization (optional)
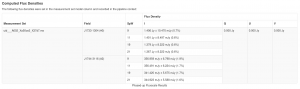
If you have multiple executions taken at a similar time with the same phase calibrator, you can rescale the derived fluxes for the phase calibrator so that they are the same in all executions. If the executions are spaced by more than a day or two, it is likely that the flux of the phase calibrator has changed and thus this step is usually unnecessary. You can compare the derived flux densities of your calibrators for each execution block using the "hifa_gfluxscale" step of the weblog (pipeline calibrations) or the *.fluxscale file (manual calibration).
The following guides may be useful for assessing the derived fluxes of your phase cailbrators. ALMA Memo 434 outlines various uncertainties relevant to amplitude calibration. You can refer to the respective cycle’s Proposer’s Guide (Cycle 4) for expected uncertainties. In addition, the ALMA Calibrator Source Catalog can be used to compare variations detected from source monitoring.
<figure id="Gfluxscale table.png">
</figure>
If you have determined flux equalization is required for your data, please file a Helpdesk ticket and a NAASC staff member can assist you by generating a script for your measurement sets. In general, the first step in flux equalization is to set the flux density of the phase calibrator in each spectral window to the desired value using setjy. Then amplitude calibration tables are generated and applied using gaincal and applycal respectively. Finally, all calibrated measurement sets will be concatenated using concat.
If you have performed flux equalization, you can skip to the Splitting off science target data section below.
Combining measurement sets from multiple executions
If there are multiple *.ms.split.cal files within the calibrated directory, combine them all into one single ms file in order to make later steps in the imaging process easier. Skip this section if you have only one execution.
Each execution of the scheduling block will generate multiple spectral windows with different sky frequencies, but the same rest frequency, due to the motion of the Earth. Thus, the resulting concatenated file will contain n spws, where n is (number of original science spws) x (number of executions). In other words, the multiple spws associated with a single rest frequency will not be regridded to a single spectral window in the ms. Combine all executions into one ms file, called calibrated.ms, using the concat command.
# in CASA
concatvis='calibrated.ms'
rmtables(concatvis)
os.system('rm -rf ' + concatvis + '.flagversions')
concat(vis=vislist,
#forcesingleephemfield='Uranus', # uncomment this line and insert source name if imaging an ephemeris object
concatvis=concatvis)
Splitting off science target data
The calibrated.ms produced in the Combining measurement sets from multiple executions section, or your original ms file if you only have one execution, will contain the science target(s) as well as all the calibrator sources. At this point, you are only concerned with imaging your science targets so you can split them off from the calibrated.ms, which has both the science and calibrator sources. This will reduce the size of the file and make it more manageable during imaging. Depending on your project, there may be several executions or only one. Uncomment the line that fits with your data to set the concatvis variable properly.
# in CASA
# Uncomment following line for single executions
# concatvis = vislist[0]
# Uncomment following line for multiple executions
# concatvis='calibrated.ms'
Split out all target fields from the file and copy them, with only the data column, into a new ms file called calibrated_source.ms. Follow this link for more information regarding the structure of measurement sets.
# in CASA
sourcevis='calibrated_source.ms'
rmtables(sourcevis)
os.system('rm -rf ' + sourcevis + '.flagversions')
split(vis=concatvis,
intent='*TARGET*', # split off the target sources
outputvis=sourcevis,
datacolumn='data')
At this point, it is convenient to create a listobs file with information about the sources and spectral windows to reference during imaging. Inspect the listobs to make sure the split and concat worked as desired.
# in CASA
listobs(vis=sourcevis,listfile=sourcevis+'.listobs.txt')
An example listobs file is below.
================================================================================
Observer: Unknown Project: T.B.D.
Observation: ALMA
Telescope Observation Date Observer Project
ALMA [ 4.81015e+09, 4.81015e+09]Unknown T.B.D.
ALMA [ 4.81015e+09, 4.81016e+09]Unknown T.B.D.
ALMA [ 4.81016e+09, 4.81017e+09]Unknown T.B.D.
Data records: 126900 Total elapsed time = 16902.1 seconds
Observed from 22-Apr-2011/00:15:36.7 to 22-Apr-2011/04:57:18.8 (UTC)
ObservationID = 0 ArrayID = 0
Date Timerange (UTC) Scan FldId FieldName nRows SpwIds Average Interval(s) ScanIntent
22-Apr-2011/00:15:36.7 - 00:16:07.0 9 0 TW Hya 540 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
00:20:57.8 - 00:30:43.2 13 0 TW Hya 8640 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
00:36:49.2 - 00:44:43.5 18 0 TW Hya 7020 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
00:49:33.9 - 00:59:19.3 22 0 TW Hya 8640 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
01:05:46.8 - 01:13:41.2 27 0 TW Hya 7020 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
01:20:07.2 - 01:29:52.6 31 0 TW Hya 8640 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
01:37:15.6 - 01:40:13.9 36 0 TW Hya 2700 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
ObservationID = 1 ArrayID = 0
Date Timerange (UTC) Scan FldId FieldName nRows SpwIds Average Interval(s) ScanIntent
22-Apr-2011/02:02:10.9 - 02:02:41.1 44 0 TW Hya 540 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
02:07:36.7 - 02:17:22.0 48 0 TW Hya 8640 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
02:23:35.7 - 02:31:30.0 53 0 TW Hya 7020 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
02:36:23.6 - 02:46:09.0 57 0 TW Hya 8640 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
02:52:40.2 - 03:00:34.6 62 0 TW Hya 7020 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
03:05:14.0 - 03:14:59.4 66 0 TW Hya 8640 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
03:21:21.4 - 03:24:19.6 71 0 TW Hya 2700 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
ObservationID = 2 ArrayID = 0
Date Timerange (UTC) Scan FldId FieldName nRows SpwIds Average Interval(s) ScanIntent
22-Apr-2011/03:44:46.9 - 03:45:17.1 79 0 TW Hya 540 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
03:50:08.4 - 03:59:53.7 83 0 TW Hya 8640 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
04:06:03.7 - 04:13:58.1 88 0 TW Hya 7020 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
04:18:49.1 - 04:28:34.4 92 0 TW Hya 8640 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
04:34:59.5 - 04:42:53.9 97 0 TW Hya 7020 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
04:47:33.5 - 04:57:18.8 101 0 TW Hya 8640 [0,1,2,3] [10.1, 10.1, 10.1, 10.1] [OBSERVE_TARGET#ON_SOURCE]
(nRows = Total number of rows per scan)
Fields: 1
ID Code Name RA Decl Epoch SrcId nRows
0 none TW Hya 11:01:51.844983 -34.42.17.16088 J2000 0 126900
Spectral Windows: (4 unique spectral windows and 1 unique polarization setups)
SpwID Name #Chans Frame Ch0(MHz) ChanWid(kHz) TotBW(kHz) CtrFreq(MHz) Corrs
0 3840 TOPO 356497.936 122.070 468750.0 356732.2500 XX YY
1 3840 TOPO 357734.314 122.070 468750.0 357968.6279 XX YY
2 3840 TOPO 346034.314 122.070 468750.0 345800.0000 XX YY
3 3840 TOPO 343955.936 122.070 468750.0 343721.6221 XX YY
Sources: 4
ID Name SpwId RestFreq(MHz) SysVel(km/s)
0 TW Hya 0 - -
0 TW Hya 1 - -
0 TW Hya 2 - -
0 TW Hya 3 - -
Antennas: 9:
ID Name Station Diam. Long. Lat. Offset from array center (m) ITRF Geocentric coordinates (m)
East North Elevation x y z
0 DV04 J505 12.0 m -067.45.18.0 -22.53.22.8 -7.2141 -541.3485 15.0178 2225061.036842 -5440128.036234 -2481534.422455
1 DV06 T704 12.0 m -067.45.16.2 -22.53.22.1 42.8987 -520.1911 15.0694 2225110.551677 -5440116.726350 -2481514.951072
2 DV07 J510 12.0 m -067.45.17.8 -22.53.23.5 -0.3652 -563.8032 14.9605 2225064.049398 -5440117.310745 -2481555.086720
3 DV08 T703 12.0 m -067.45.16.2 -22.53.23.9 42.8798 -575.6928 14.6278 2225102.207484 -5440096.375809 -2481565.910698
4 DV09 N602 12.0 m -067.45.17.4 -22.53.22.3 8.8012 -527.8598 15.0513 2225077.857538 -5440126.858119 -2481522.008896
5 DV10 N606 12.0 m -067.45.17.1 -22.53.23.6 19.1981 -566.5667 14.9520 2225081.746122 -5440108.902762 -2481557.629365
6 PM01 T702 12.0 m -067.45.18.6 -22.53.24.1 -23.6269 -582.3103 14.9195 2225039.780230 -5440119.418780 -2481572.120530
7 PM02 T701 12.0 m -067.45.18.8 -22.53.22.2 -29.1268 -522.7917 15.0566 2225043.501617 -5440143.045000 -2481517.341976
8 PM03 J504 12.0 m -067.45.17.0 -22.53.23.0 22.2015 -550.2548 14.9948 2225086.942689 -5440113.674706 -2481542.618537
Regridding spectral window (optional)
ALMA does what is called doppler setting, which means it adjusts the observed sky frequency for the motion of the source and the motion of the Earth around the Sun at the beginning of every execution block. Therefore, different execution blocks will have different observed sky frequencies because the vector of the motion of the Earth around the Sun will be different at different times.
In general, we strongly recommend using the tclean command to regrid spectral windows to a common velocity/rest frequency scale on the fly. However, you may also regrid the frequency axis of the visibility data using cvel prior to imaging. The most common use case for this is where the motion of the Earth around the Sun causes the sky frequencies of the lines to shift too much between executions to identify a common channel range for continuum subtraction. If you regrid using cvel prior to imaging, you should make sure to use the same input parameters to the tclean task that you used for cvel. In other words, you shouldn't regrid the data using both cvel and tclean.
You will likely need to regrid if you are imaging an ephemeris object. In this case, the outframe should be set to SOURCE, for example, outframe = 'SOURCE'.
Due to a bug in all versions of CASA prior to 5.0, mstransform and cvel should not be used to average greater than 2 channels together. This averaging should be done using tclean if using a CASA version less than 5.0
We will start by setting the parameters specific to your observations.
# in CASA
sourcevis='calibrated_source.ms'
regridvis='calibrated_source_regrid.ms'
veltype = 'radio' # Keep set to radio. See notes in imaging section.
width = '0.23km/s' # due to bug in cvel/mstransform, do not regrid > 2 channels
nchan = -1 # leave this as the default
mode='velocity' # see science goals in the OT
start='' # leave this as the default
outframe = 'bary' # velocity reference frame. see science goals in the OT.
restfreq='115.27120GHz' # rest frequency of primary line of interest.
field = '4' # select science fields.
spw = '0,5,10' # spws associated with a single rest frequency. Do not attempt to combine spectral windows associated with different rest frequencies. This will take a long time to regrid and most likely isn't what you want.
Regridding here uses the cvel command to regrid multiple spws associated with a single rest frequency into a single spw. To avoid tclean regridding the image a second time you will need to use the same parameters used for cvel for tclean.
# in CASA
rmtables(regridvis)
os.system('rm -rf ' + regridvis + '.flagversions')
cvel(vis=sourcevis,
field=field,
outputvis=regridvis,
spw=spw,
mode=mode,
nchan=nchan,
width=width,
start=start,
restfreq=restfreq,
outframe=outframe,
veltype=veltype)
The resulting ms will contain a single spw. You should regrid spws at different rest frequencies separately.
Rename and backup data set
Depending on the different tasks that were performed above, the current measurement set will have a variety of names. For ease of imaging with a common measurement set, rename the file to calibrated_final.ms.
If you haven’t regridded, your current ms should be named calibrated_source.ms.
# in CASA
# uncomment if you haven’t regridded
# os.system('mv -i ' + sourcevis + ' ' + 'calibrated_final.ms')
If you have regridded, your current ms should be named calibrated_source_regrid.ms.
# in CASA
# uncomment if you have regridded
# os.system('mv -i ' + regridvis + ' ' + 'calibrated_final.ms')
Now that you have a fully calibrated measurement set containing all your science sources to image, we recommend creating a backup of the dataset. It is possible to corrupt a measurement set if you kill clean, tclean, or uvcontsub while they are running.
# in CASA
os.system('cp -ir calibrated_final.ms calibrated_final.ms.backup')
Continue to Imaging
This guide is continued at Image the Continuum Template.