Fit a Gaussian to Visibility data and plot it over the data
From CASA Guides
Jump to navigationJump to search
- DRAFT*
B.Mason, K.Ward-Duong, & J.Patience July 2015 developed in CASA 4.2.2
This CASA guide illustrates how to run UVMODELFIT to fit a single Gaussian component to UV data (i.e. the raw visibilities); and subsequently how to make a plot of the binned, "azimuthally" averaged (in UV space) UV data together with the fitted model. In broad outlines, the steps required are:
1 Estimate the peak location of emission. shift the phase center here if desired. 2 Estimate starting values for the gaussian component fit 3 do the fit. 4 run ft() to put the results of the fit in the MS explicitly (as a MODEL column). 5 Extract MODEL and DATA, make the plot
PHASE SHIFT:
fixvis(vis="J04292165_cont.ms",outputvis="J04292165_cont_recenter.ms",field="",refcode="",reuse=True,phasecenter="J2000 4h29m21.66s +27d01m25.72s",datacolumn="all")
(note: not done in this example)
do the fit -- this is for all spws together:
default uvmodelfit vis=myvis field='0' comptype='G' sourcepar=[0.006,0.1,-0.05,0.4,0.3,0.0] varypar=[] outfile='J04292165_cont_allspw.uv.cl' inp go
in this example we had not phase shifted.
run FT -- you must have usescratch=True
default ft vis=myvis complist='J04292165_cont_allspw.uv.cl' usescratch=True inp go # did that work? plotms(vis=myvis,xaxis='uvwave',yaxis='real',ydatacolumn='model') # yes it did.
Finally, use VISSTAT() to extract model and data values in bins of uv-radius:
import numpy as np
import matplotlib.pyplot as plt
import glob
testms = 'J04292165_cont.ms'
# set the uvrange in kilo lambda
# NB: take care that the bin width and uvmin/max ranges ensure
# you get no bins without data! the appropriate values may change
# if you, e.g., change the SPW you are looking at
uvmin = 0
#uvmax = 1200
uvmax = 800
# define steps in klambda
duv=40
uvsteps = np.arange(uvmin, uvmax, duv)
avg_amps = [] # define an empty list in which to put the averaged amplitudes
stddev_amps = [] # define an empty list in which to put the amp stddev
numpoints = []
uvpoints = uvsteps[:-1] + np.diff(uvsteps)/2 # get array of uvmidpoints over which avg taken
model_amps = [] # define list to get model amplitudes after fourier transforming into the model data column
# iterate over the uvrange & build up
# the binned data values (looking only at the Real part -- a pretty
# good approach if the signal is approximately symmetric and you have
# recentered the phase center on the peak of emission):
for ii in range(len(uvsteps[:-1])):
tmp = visstat(vis = testms,axis = 'real',
uvrange = str(uvsteps[ii]) + '~' + str(uvsteps[ii+1]) + 'klambda',
datacolumn = 'data')
print str(uvsteps[ii]) + '~' + str(uvsteps[ii+1]) + 'klambda'
avg_amps.append(tmp['DATA']['mean'])
stddev_amps.append(tmp['DATA']['stddev'])
numpoints.append(tmp['DATA']['npts'])
# now build up the model values
for ii in range(len(uvsteps[:-1])):
tmp = visstat(vis = testms,
axis = 'real', # you may want to change this to real...?
uvrange = str(uvsteps[ii]) + '~' + str(uvsteps[ii+1]) + 'klambda',
datacolumn = 'model')
print str(uvsteps[ii]) + '~' + str(uvsteps[ii+1]) + 'klambda, from the model column'
model_amps.append(tmp['MODEL']['mean'])
error_amps = stddev_amps/(np.sqrt(numpoints) - 1)
plt.clf()
plt.cla()
avg_amps_mjy = [x * 1000 for x in avg_amps]
model_amps_mjy = [x * 1000 for x in model_amps]
error_amps_mjy = [x * 1000 for x in error_amps]
plt.errorbar(uvpoints, avg_amps_mjy, yerr = error_amps_mjy, mfc='k', fmt='o', label='data')
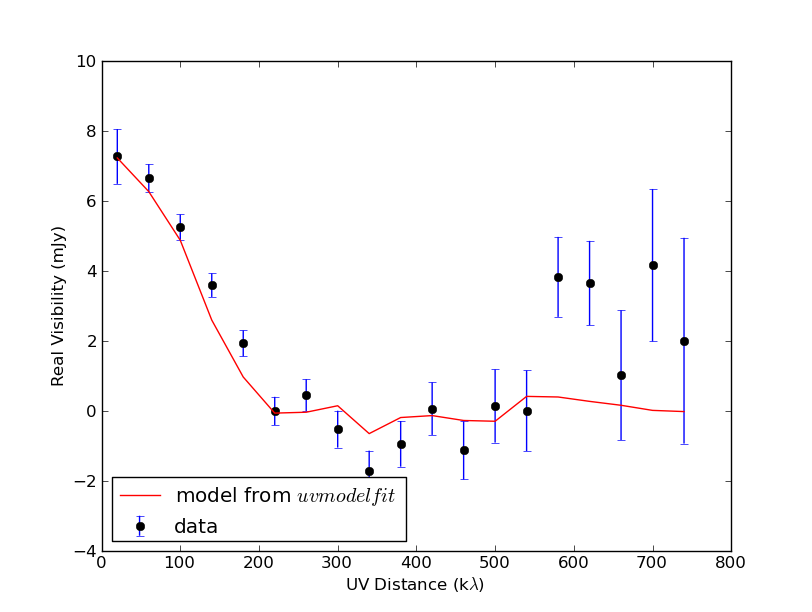
plt.plot(uvpoints, model_amps_mjy, 'r-', label=r'model from $uvmodelfit$')
plt.legend(loc = 3, numpoints = 1)
plt.xlabel(r'UV Distance (k$\lambda$)')
plt.ylabel('Real Visibility (mJy)')
plt.show()
The result--