AntennaeBand7 North Imaging 3.4
- This tutorial picks up where AntennaeBand7_North_Calibration_3.4 leaves off: with a fully calibrated, split science target measurement set (Antennae_North.cal.ms). If you wish to skip the Calibration guide: obtain the calibrated data from AntennaeBand7_3.4#Obtaining_the_Data; extract it using tar -xzvf FILENAME; and cd into the extracted directory.
- Details of the ALMA data are provided at AntennaeBand7_3.4.
- This casaguide covers only the Northern Mosaic of the Antennae.
Confirm your version of CASA
This guide has been written for CASA release 3.4.0. Please confirm your version before proceeding.
# In CASA
version = casalog.version()
print "You are using " + version
if (int(version.split()[4][1:-1]) < 19407):
print "\033[91m YOUR VERSION OF CASA IS TOO OLD FOR THIS GUIDE."
print "\033[91m PLEASE UPDATE IT BEFORE PROCEEDING."
else:
print "Your version of CASA is appropriate for this guide."
Imaging Mosaics
If you are unfamiliar with the basic concepts of deconvolution and clean, pause here and review for example http://www.aoc.nrao.edu/events/synthesis/2010/lectures/wilner_synthesis10.pdf
Mosaics like other kinds of images are created in the CASA task clean. To invoke mosaic mode, you simply set the parameter imagermode='mosaic'. The default subparameter ftmachine='mosaic' is then automatically set. This is a joint deconvolution algorithm that works in the uv-plane. A convolution of the primary beam patterns for each pointing in the mosaic is created: the primary beam response function. The corresponding image of the mosaic response function will be called <imagename>.flux.pbcoverage and <imagename>.flux (where the latter differs from the former only if the sensitivity of each field in the mosaic varies).
Well-sampled mosaics like the patterns observed for the Northern and Southern Antennae mosaics shown on the Overview page are well-suited to joint deconvolution. However, if you have a poorly-sampled, or very irregular mosaic you may need to use the slower ftmachine='ft' which combines data in the image plane, similar to what is done in other packages like MIRIAD.
Additionally, for mosaics it is essential to pick the center of the region to be imaged explicitly using the phasecenter parameter. Otherwise it will default to the first pointing included in the field parameter -- since this is often at one corner of the mosaic, the image will not be centered. For the Northern mosaic, the center pointing corresponds to field id 12. Note that during the final split in the calibration section that selected only the Antennae fields, the field ids were renumbered, so that the original centers (shown in the Overview) have changed: field id 14 becomes 12 for the Northern mosaic and field 18 becomes 15 for the Southern mosaic. You can also set an explicit coordinate (see the clean help for syntax).
If you want to learn more about mosaicing, pause here and review for example http://www.aoc.nrao.edu/events/synthesis/2010/lectures/jott-mosaicking-school-04.pdf
Setting up the Dataset
At the end of the AntennaeBand7_North_Calibration_3.4 walk-through you should have ended up with the concatenated calibrated averaged dataset Antennae_North.cal.ms. We strongly recommend that you start this casaguide tutorial in a new directory or sub-directory and make a copy from your original Antennae_North.cal.ms here (in case you somehow corrupt your working verion).
If you are doing this tutorial on its own, then you can get the MS using the instructions given above.
Continuum Imaging
<figure id="Antennae_North-AMPvsCH.png">
</figure>
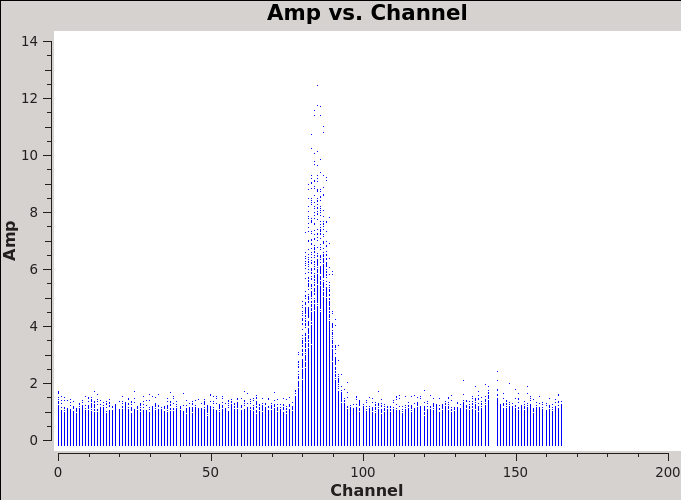
We will make 345 GHz continuum images for the region covered by the Northern mosaic. We use the task clean over the channels that are free of the line emission; we avoid the edge channels which tend to be noisier due to bandpass rolloff effects. The line-free channels are found by plotting the average spectrum (all fields). We find the CO(3-2) line from channels 70 to 100 in the northern mosaic (<xr id="Antennae_North-AMPvsCH.png"/>).
# In CASA
os.system('rm -rf Antennae_North-AMPvsCH.png')
plotms(vis='Antennae_North.cal.ms',
xaxis='channel',yaxis='amp',
avgtime='1e8',avgscan=T,
plotfile='Antennae_North-AMPvsCH.png')
The avgtime is set to a large value so that it averages over all the integrations, and avgscan is set to allow averaging of the different scans.
Next we create continuum images from the line-free channels.
Cleaning the Northern Continuum Mosaic
For illustrative purposes we first make a dirty image to see if there is emission and what the exact beam size is. It should be on the order of 1" but this will vary a bit according to the uv-coverage in the actual data. We will start with a cell size of 0.2" to oversample the beam by a factor of 5. The imsize needs to be large enough given the cell size to comfortably encompass the mosaic. From the mosaic footprints shown in the overview, we can see that the Northern mosaic imsize needs to be about 1 arcmin. With 0.2" pixels, this requires imsize=300.
Other essential clean parameters for this case include:
- vis='Antennae_North.cal.ms' : The calibrated dataset on the science target. clean will always use the CORRECTED DATA column (if it exists).
- imagename='Antennae_North.Cont.Dirty' : The base name of the output images:
- <imagename>.image # the final restored image
- <imagename>.flux # the effective primary beam response (where pbcor=True)
- <imagename>.flux.pbcoverage # the primary beam coverage (ftmachine=’mosaic’ only)
- <imagename>.model # the model image
- <imagename>.residual # the residual image
- <imagename>.psf # the synthesized (dirty) beam
- spw='0:1~50;120~164' : To specify only the line-free channels of spectral window 0.
- mode='mfs' : Multi-Frequency Synthesis: The default mode, which produces one image from all the specified data combined. This will grid each channel independently before imaging. For wide bandwidths this will give a far superior result compared to averaging the channels and then imaging.
- restfreq='345.79599GHz' : The rest frequency of CO(3-2) can be found with splatalogue.
- niter=0: Maximum number of clean iterations. (niter=0 will do no cleaning)
- usescratch=False: Do not force clean to create an explicit MODEL_DATA scratch column, but to store the model and gridding information in the MS header for later use.
# In CASA
os.system('rm -rf Antennae_North.Cont.Dirty*')
clean(vis='Antennae_North.cal.ms',
imagename='Antennae_North.Cont.Dirty',
field='',phasecenter='12',
mode='mfs',restfreq='345.79599GHz',
spw='0:1~50;120~164',
imagermode='mosaic',usescratch=False,
imsize=300,cell='0.2arcsec',
interactive=F,niter=0)
The reported beam size is about 1.27" x 0.71" P.A. 86.142 degrees.
# In CASA
viewer('Antennae_North.Cont.Dirty.image')
If you are unfamiliar with the CASA viewer it is a good idea to pause here and watch the demo at http://casa.nrao.edu/CasaViewerDemo/casaViewerDemo.html
<figure id="North_cont_clean.png">
</figure>
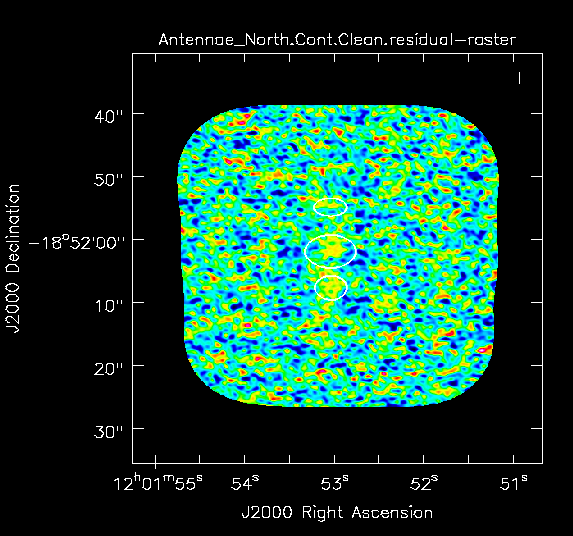
Yes there is definitely a detection in the vicinity of the Northern nucleus (see <xr id="North_contstat.png"/> and <xr id="Antennae_North.Cont.Clean.image.png"/> below). Using the square region icon, and drawing a box near but not including the emission, we find the rms noise is about 0.4 mJy/beam in the dirty image.
Next we switch to refined values of cell='0.13arcsec' and imsize=500 based on the observed beam size. We also switch to interactive mode so that you can create a clean mask using the polygon tool (note you need to double click inside the polygon region to activate the mask). See TWHya casaguide for a more complete description of interactive clean and mask creation.
- niter=1000: Maximum number of clean iterations -- we will stop interactively
- threshold='0.4mJy' : Stop cleaning if the maximum residual is below this value (the dirty rms noise)
- interactive=T: Clean will be periodically interrupted to show the residual clean image. Interactive clean mask can be made. If no mask is created, no cleaning is done.
# In CASA
os.system('rm -rf Antennae_North.Cont.Clean*')
clean(vis='Antennae_North.cal.ms',
imagename='Antennae_North.Cont.Clean',
field='',phasecenter='12',
mode='mfs',restfreq='345.79599GHz',
spw='0:1~50;120~164',
imagermode='mosaic',usescratch=False,
imsize=500,cell='0.13arcsec',
interactive=T,
niter=1000, threshold='0.4mJy')
The residuals are "noise-like" after only 100 iterations (see <xr id="North_cont_clean.png"/>), so hit the red X symbol in the interactive window to stop cleaning here.
Note that if you run the clean task again with the same imagename, without deleting the existing <imagename>.* files, clean assumes that you want to continue cleaning the existing images. We put an rm command before the clean command to guard against this, but sometimes this is what you want.
Image Statistics
<figure id="North_contstat.png">
</figure>
You can determine statistics for the images using the task imstat:
# In CASA
mystat = imstat('Antennae_North.Cont.Clean.image')
where the result dictionary is captured in the mystat variable. From this we find, that for the Northern continuum image the peak is 3.56 mJy/beam and the rms is 0.37 mJy/beam. You might get slightly different values but they should be in the ballpark of these.
However, the calculation of the rms comes with a couple of caveats. First, the mosaic primary beam response rolls off toward the edges of the mosaic, as do correspondingly the flux density and rms. Thus if you don't restrict the measurement to areas of full sensitivity, the apparent rms is skewed downward. Second, since there is real emission in the image, the rms will be skewed upward (with the error increasing with brighter emission). Both can be solved by picking boxes that exclude the edges of the mosaic and the real emission.
It is often easier to use either the viewer directly or imview (a task wrapper for the viewer; see help imview for more info) to display the image and then interactively use the region tools to get the statistics.
# In CASA
viewer('Antennae_North.Cont.Clean.image')
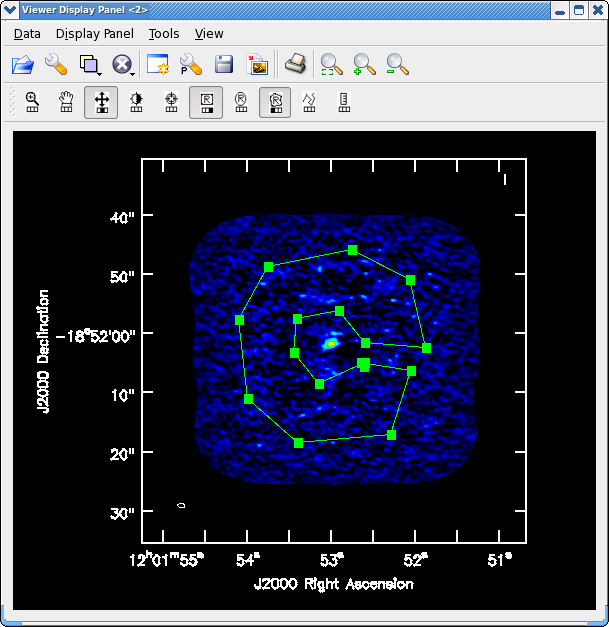
Adjust colorscale using the middle mouse button (or reassign to a different mouse button by clicking on "plus" symbol icon). Next, select the polygon tool by clicking on its symbol with a mouse button. Then draw a region that avoids edges and emission (see <xr id="North_contstat.png"/>), then double click inside the polygon (using the left mouse button in 3.4) to have the statistic printed to the screen. We find rms=0.40 mJy/beam, somewhat larger than the raw image statistics value.
Note for CASA 3.4 Note that the region drawing and managing mechanics have been changed in CASA 3.4. When you draw a region for the first time, a new panel will appear on your viewer for the region information (probably displacing much of your image). You can dismiss this panel (and bring it back using the View menu) or undock it and move it off your main viewer display (use the undock icon in upper right of that panel next to the "x" dismiss button). You start drawing vertices by clicking on points in the image in succession, when you draw the final vertex then you double-click to connect and close the region. When you mouse is inside the region a bounding box will appear with the vertices shown as draggable solid squares. If you want to adjust the vertices you can do so! These new region features are still under development and sometimes you might find that you get incomplete regions, or there is a lag when you start drawing and the lines appear, and other oddities. If you find you don't like your region you can dismiss it with with ESC key or using the remove region "X" button in lower right of the panel.
<figure id="Antennae_North.Cont.Clean.image.png">
</figure>
How does this compare to theory? You can find out using the ALMA sensitivity calculator. The continuum bandwidth of the Northern mosaic is about 1 GHz and about 0.85 GHz for the Southern mosaic. The number of antennas is about 12 and the time on a single pointing is about 300s. This yields an expected rms of about 1 mJy/beam. However, this needs to be decreased by about a factor of 2.5 near the center of the mosaic due to the hexagonal Nyquist sampling of the mosaic (radial spacing~0.37*HPBW) for a theoretical rms of about 0.4 mJy/beam, in good agreement with observation.
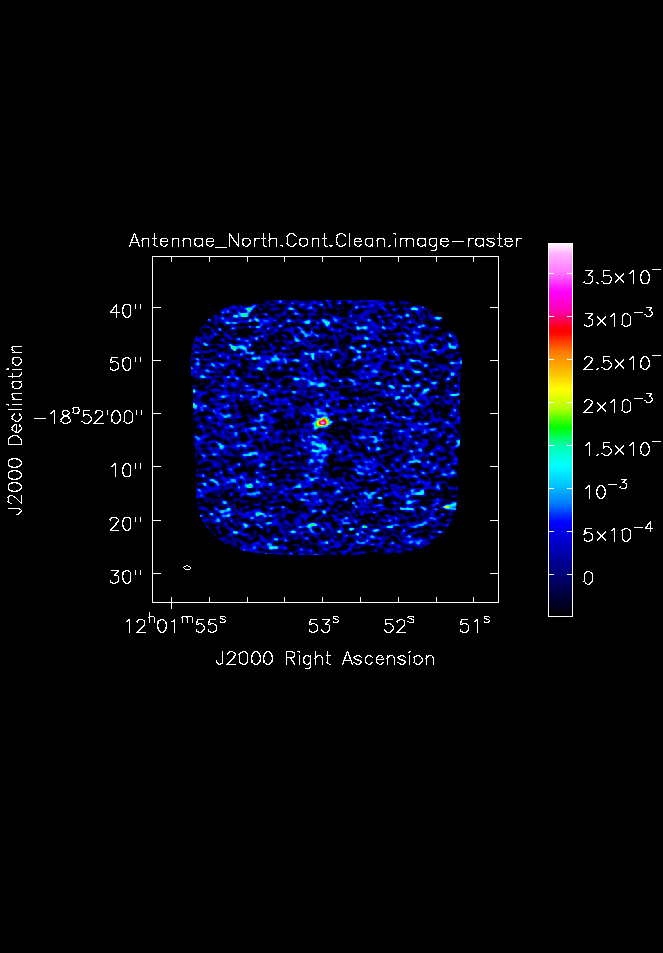
Next make hardcopy plots of the continuum images using imview. To make the contrast a bit better, set the data range from -1 x sigma to the peak determined above.
# In CASA
imview(raster={'file': 'Antennae_North.Cont.Clean.image',
'colorwedge':T,'range':[-0.00048,0.00380]},
zoom=1, out='Antennae_North.Cont.Clean.image.png')
You can see the output in <xr id="Antennae_North.Cont.Clean.image.png"/>.
Continuum subtraction
In these data, the continuum emission is too weak to contaminate the line emission (i.e. the peak continuum emission is less than the rms noise in the spectral line channels). Nevertheless, for illustrative purposes we demonstrate how to subtract the continuum emission in the uv-domain using the task uvcontsub2.
# In CASA
uvcontsub2(vis='Antennae_North.cal.ms',fitspw='0:1~50;120~164',fitorder=1)
Here, fitspw gives the line-free channels for each mosaic and fitorder=1. Higher order fits are not recommended. If you do not have line-free channels on both sides of the line fitorder=0 is recommended. The output MS will have .contsub appended to the name.
CO(3-2) Imaging
<figure id="North_CO3_2_vel.png">
</figure>
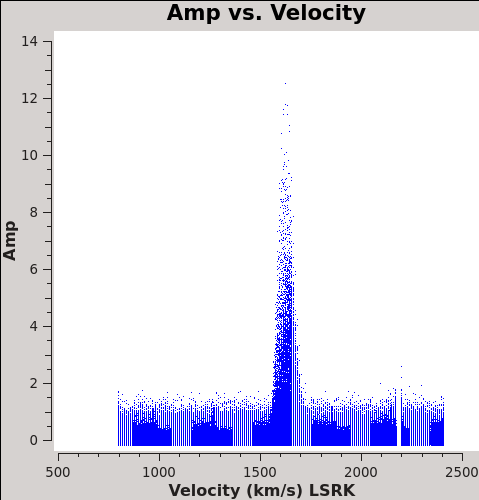
Now we are ready to make cubes of the line emission. The imaging parameters are similar to the continuum except for those dealing with the spectral setup: mode, start, width, nchan, restfreq, and outframe parameters. When making spectral images you have three choices for the mode parameter: channel, velocity, and frequency. Data are taken using constant frequency channels. For spectral line analysis it's often more useful to have constant velocity channels, and this is also the best way to make images of multiple lines with the exact same channelization for later comparison. For mode='velocity', the desired start and width also need to be given in velocity units for the desired output frame.
It is important to note that ALMA does not do on-line Doppler Tracking and the native frame of the data is TOPO. If you do not specify outframe the output cube will also be in TOPO, which is not very useful for scientific analysis. The Doppler Shift is taken out during the regridding to the desired outframe in clean or alternatively it can be done separately by the cvel task which would need to be run before clean.
To see what velocity parameters you want to set in clean, it is useful to make a plot in plotms in the desired output frame. Note these plots take a little longer because of the frame shift. In order to compare with recent SMA data, we chose LSRK, but it should be noted that many papers of this source are in the BARY frame.
# In CASA
os.system('rm -rf North_CO3_2_vel.png')
plotms(vis='Antennae_North.cal.ms.contsub/',
xaxis='velocity',yaxis='amp',
avgtime='1e8',avgscan=T,transform=T,freqframe='LSRK',
restfreq='345.79599GHz',plotfile='North_CO3_2_vel.png')
Cleaning the Northern Mosaic Spectral Cube
Notable parameters to be included in the clean call for our spectral cube are:
- mode='velocity' ,outframe='LSRK' ,restfreq='345.79599GHz' . We use 'velocity' mode to set velocity information in the Local Standard of Rest frame (kinematic definition), and use the rest frequency of the CO(3-2) line.
- nchan=60,start='1300km/s' ,width='10km/s' : To produce a data cube with 60 channels ("nchan"=60), starting at 1300km/s and with velocity widths of 10 km/s. That will include all the CO(3-2) emission in both mosaics.
- imsize=500,cell='0.13arcsec' : An image size of 65 arcsec, with pixels of 0.13 arcsec (about one-fifth the minor axis of the synthesized beam). We make the imsize larger than the mosaic to increase the dirty beam image size, as the emission is quite extended along the mosaic.
- weighting='briggs' ,robust=0.5: Weighting to apply to visibilities. We use Briggs' weighting and robustness parameter 0.5 (in between natural and uniform weighting).
- niter=10000: Maximum number of clean iterations.
- threshold='5.0mJy' : Stop cleaning if the maximum residual is below this value.
The threshold is set to roughly the rms of a single line-free channel for each dataset.
<figure id="screenshot.AntennaeNorthInteractiveClean_initial.png">
</figure>
<figure id="screenshot.AntennaeNorthInteractiveClean_n1000.png">
</figure>
# In CASA
os.system('rm -rf Antennae_North.CO3_2Line*')
clean(vis='Antennae_North.cal.ms.contsub',
imagename='Antennae_North.CO3_2Line.Clean',
spw='0',field='',phasecenter='12',
mode='velocity',outframe='LSRK',restfreq='345.79599GHz',
nchan=70,start='1200km/s',width='10km/s',
imagermode='mosaic',
imsize=500,cell='0.13arcsec',minpb=0.2,
interactive=T,usescratch=False,
weighting='briggs',robust=0.5,
niter=10000, threshold='5.0mJy')
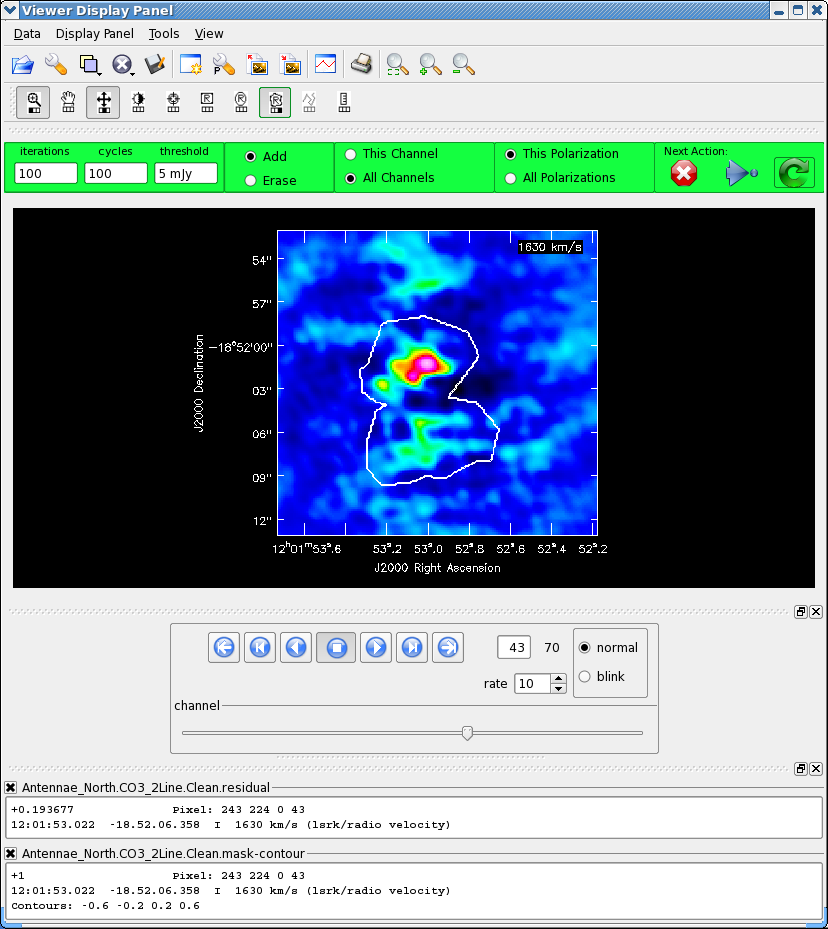
As before, it is very important that you make a clean mask. There are many ways to do this ranging from the complicated to simple. For this example we will make a single clean mask that encompasses the line emission in every channel and apply it to all channels. This is much better than no clean mask, though not quite as good as making an individual mask for each channel. When you reach the interactive clean display, use the tapedeck "play" button to play through the channels. You should see emission appear and peak around channel 42. On a playthrough you can "stop" the movie and step back and forth around the peak. Make sure the "All Channels" radio button is selected (so the mask applies to all channels). Use the polygon region tool to draw around the emission. Be sure to rock the channel displayed back and forth to make sure you encompass the emission you want and adjust the region accordingly by adding new area. For this initial stage just box up the brightest emission in the center, the outer stuff may just be sidelobes from the central emission, if its real we can pick it up later on. See <xr id="screenshot.AntennaeNorthInteractiveClean_initial.png"/> for an example of an initial clean region ready to go.
CASA 3.4 note: The interactive clean "viewer" is still the older 3.3 version and does not have the new region tools described earlier.
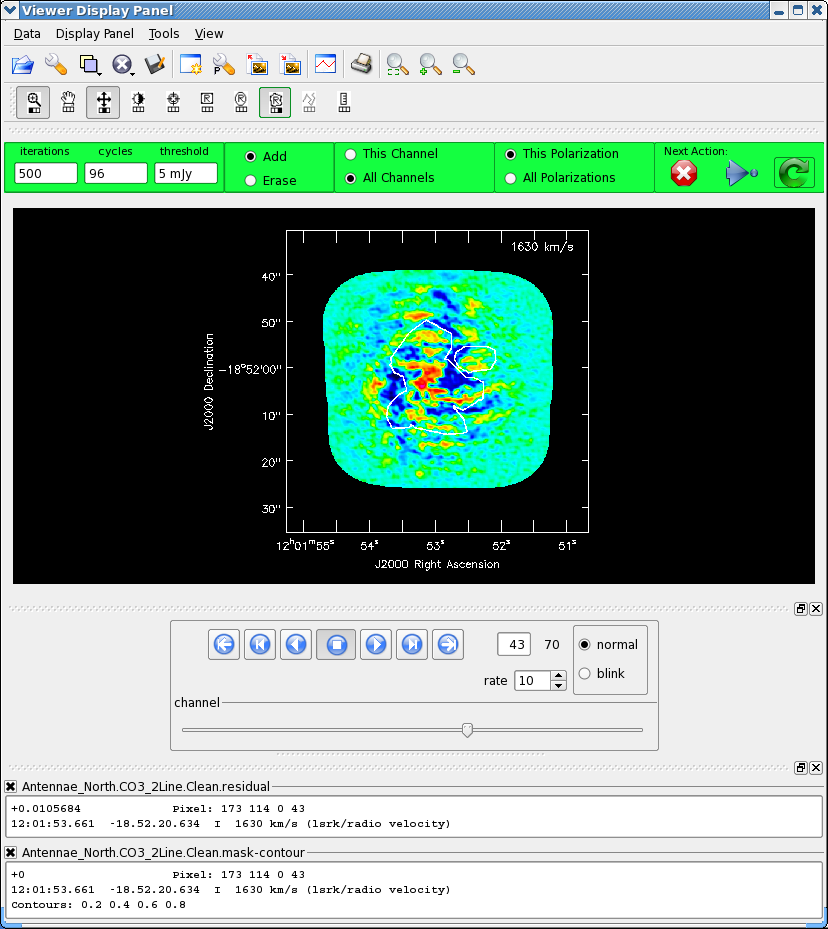
Now clean using the green circle arrow. You can watch the progress in the logger. When 100 iterations are done, the viewer will show the residual map for each channel. Cycle through the channels and see whether you are still happy with the clean box in each channel with significant signal. The "erase button" can help you fix mistakes. If necessary adjust. It is often useful to adjust the colorscale with the "plus" symbol icon. To make it go a bit faster, you can increase iteration interactively, to 200 or 300, but don't overdo it. <xr id="screenshot.AntennaeNorthInteractiveClean_n1000.png"/> shows what our clean box looked like after around 1000 iterations after which we stopped (see below).
These data are pretty severely dynamic range limited, in part due to the sparse uv coverage. In other words, the noise in bright channels is set by a maximum signal-to-noise. This effectively prevents us from stopping clean based on an rms based threshold because the effective rms changes as a function of the brightest signal in each channel. Thus, in this case we need to stop clean interactively. Below an approximate number of iterations is given to help you decide when to quit. NOTE: the threshold that is set in the CO(3-2) clean commands are about equal to the noise in a line-free channel and is only there to prevent clean running forever if you fail to stop it. You are not intended to clean to this threshold.
Also, we are going to self-calibrate the data so it's best to be conservative here - it can be difficult in images like these to discern real features from artifacts. If in doubt, don't include it in the clean mask. Keep cleaning and see if the feature becomes weaker. You cannot lose real emission by not masking it, but you can create a brighter feature by masking an artifact.
Stop cleaning after about 900-1000 iterations (Red X), when the artifacts start to look as bright as the residual.
Inspect the resulting data cube:
# In CASA
viewer('Antennae_North.CO3_2Line.Clean.image')
For the mosaic use the viewer polygon tool as described above for the continuum, to find the image statistics in both a line-free channel and the channel with the strongest emission.
The line-free channel rms for the northern mosaic is about 3.1-3.5 mJy/beam. However in a bright channel this degrades to about 6-9 mJy/beam. This is the aforementioned dynamic range limit.
Using the ALMA sensitivity calculator, a bandwidth of 10 km/s, dual polarization, 12 antennas, and the time on a single pointing of about 300s yields an rms of about 9.0 mJy/beam. As described above the Nyquist sampled hexagonal mosaic improves this by a factor of about 2.5 for an expected rms noise of about 3.6 mJy. So in line-free channels we are doing close to theoretical.
Self Calibration
Next we attempt to self-calibrate the uv-data. The continuum is far too weak so we will use the CO(3-2) line emission. The process of self-calibration tries to adjust the data to better match the model image that you give it. That is why it is important to make the model as good as you can by making clean masks. In the clean above we set usescratch=False so the model image and gridding information was placed in the header of the MS. This model can then be used to self-calibrate the data using gaincal.
Mosaics can be tricky to self-calibrate. The reason is that typically only a few of the pointings may have strong emission. The stronger the signal, the stronger the signal-to-noise will be for a given self-calibration solution interval (solint). This means that typically only a few fields are strong enough for self-calibration, though it can still bring about overall improvement to apply these solutions to all fields. The fine tuning of each field in the mosaic over small solutions intervals is often impossible. Indeed fine-tuning only a few fields in the mosaic on small time scales can result in a mosaic with dramatically different noise levels as a function of position. Below, we will simply attempt to self-calibrate on the strongest field in the mosaic, obtaining one average solution per original input dataset.
Another wrinkle is the self-calibration of spectral line data. To increase signal-to-noise it is often helpful to limit solutions to the range of channels containing the strongest line emission.
We will also use gaintype='T' to tell CASA to average the polarizations before deriving solutions which gains us a sqrt(2) in sensitivity.
Self-Calibration and Re-Imaging of the Northern Mosaic
<figure id="north_selfchan_data.png">
</figure>
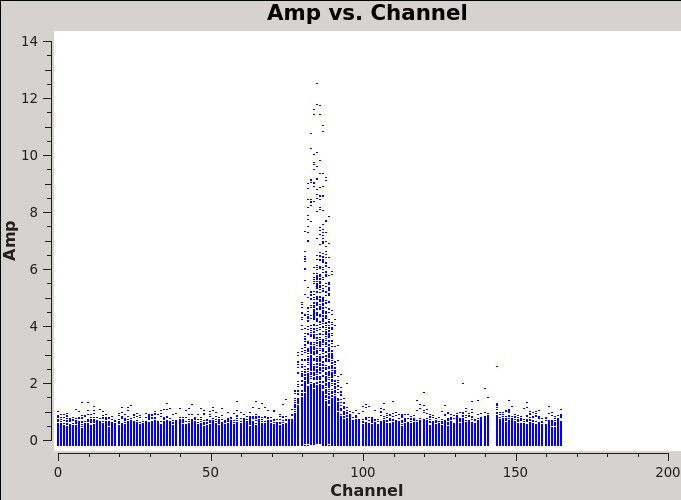
The strongest emission in the Northern mosaic comes from the center pointing, already identified above as field='12' . Below we make a uv-plot of the spectral line emission from this field to chose only the strongest channels to include in the self-calibration. It is essential that the self-calibration channel range be chosen based on the UV data and not on the image cube. This is because the channels in the image cube were modified by the velocity/channel specifications in the first invocation of clean. This plot shows the channel range of the uv data.
# In CASA
os.system('rm -rf North_selfchan_data.png')
plotms(vis='Antennae_North.cal.ms.contsub',
xaxis='channel',yaxis='amp',ydatacolumn='data',
spw='',field='12',avgtime='1e8',avgscan=T,
plotfile='North_selfchan_data.png')
We also want to check the channel amplitudes in the model column of the measurement set. The depth of the initial clean will determine what model channels have emission. (A very shallow clean could potentially produce a model with emission over a small range of channels.) Self calibration will attempt to make the data look like the model. Therefore, it is important that we select a channel range where the model is realistic.
<figure id="North_selfchan_model.png">
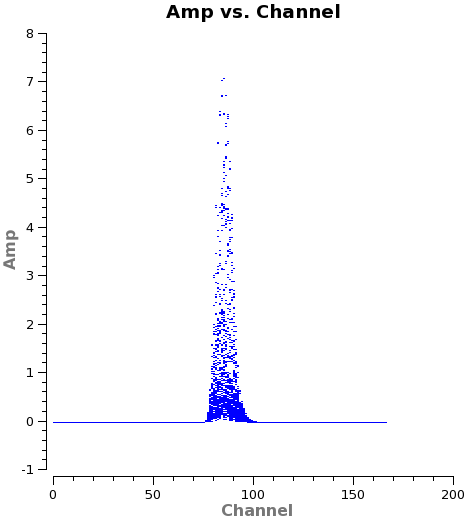
</figure>
# In CASA
os.system('rm -rf North_selfchan_model.png')
plotms(vis='Antennae_North.cal.ms.contsub',
xaxis='channel',yaxis='amp',ydatacolumn='model',
spw='',field='12',avgtime='1e8',avgscan=T,
plotfile='North_selfchan_model.png')
Next remind ourselves of the timing of the various observations.
# In CASA
plotms(vis='Antennae_North.cal.ms.contsub',
xaxis='time',yaxis='amp',field='12',
spw='',avgchannel='167',coloraxis='scan')
Zooming in on each dataset we see that they are about 1 hour long (you can also check the listobs output), and that each dataset has 2 or 3 observations of field 12. Each time, it's observed for about 25 seconds. We set solint='3600s' to get one solution per dataset.
<figure id="plotcal.North.selfcal1_phase.png">
</figure>
# In CASA
gaincal(vis='Antennae_North.cal.ms.contsub',
caltable='North.selfcal1',
solint='3600s',combine='scan',gaintype='T',field='12',
refant='DV09',spw='0:82~89',minblperant=4,
calmode='p',minsnr=2)
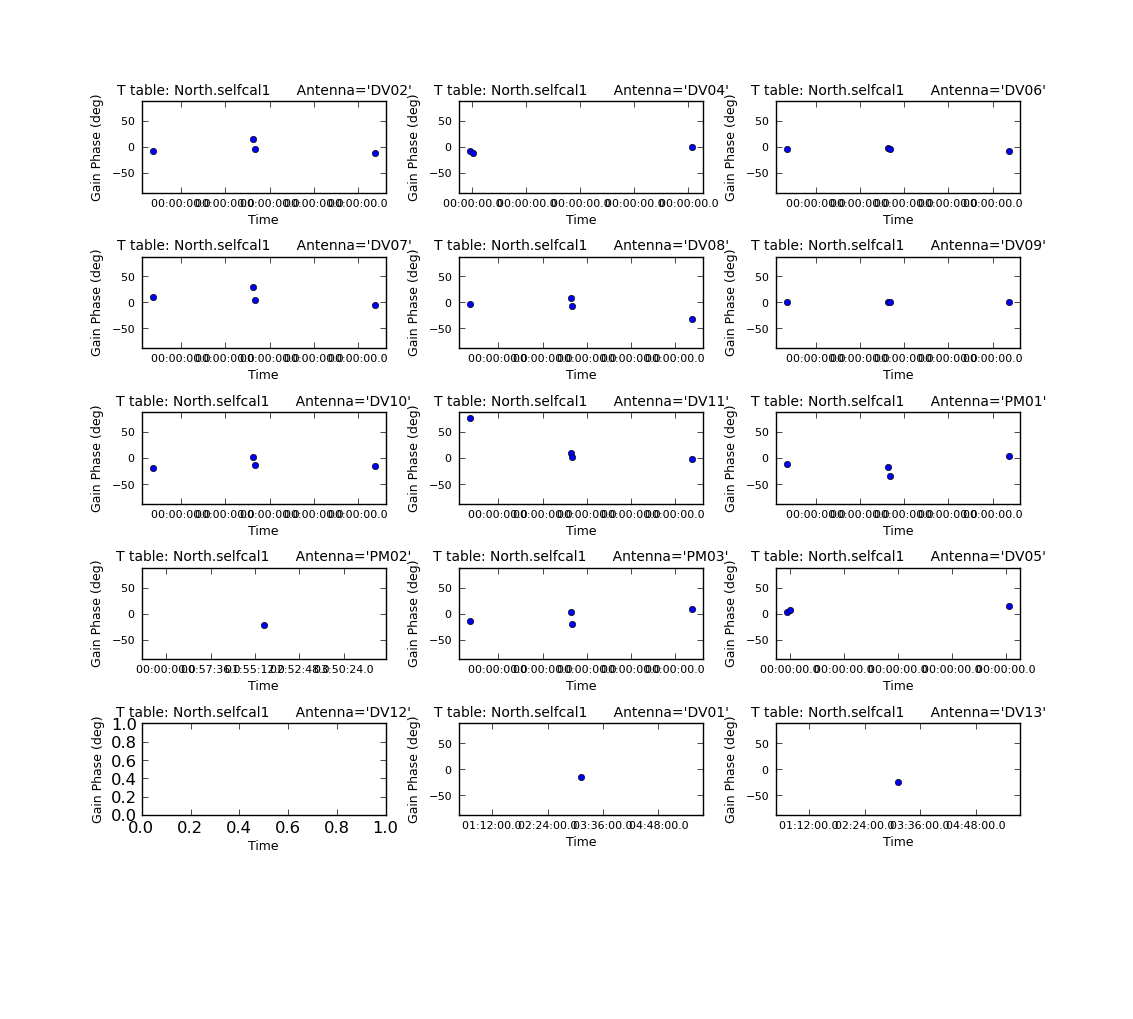
Plot the calibration solutions:
# In CASA
plotcal(caltable='North.selfcal1',
xaxis='time',yaxis='phase',
spw='',field='',antenna='',
iteration='antenna',subplot=531,
plotrange=[0,0,-80,80],
figfile='plotcal.North.selfcal1_phase.png')
Apply the solutions to all fields and channels with applycal, which will overwrite the corrected data column:
# In CASA
applycal(vis='Antennae_North.cal.ms.contsub',
gaintable=['North.selfcal1'],
field='',calwt=F)
Note that because we never created the scratch columns for the MS, it will do so the first time we run applycal.
<figure id="screenshot.AntennaeNorthInteractiveClean_selfcalfinal.png">
</figure>
Re-image the data with the selfcal applied. Remember that clean always uses the corrected data column if it exists:
# In CASA
os.system('rm -rf Antennae_North.CO3_2Line.Clean.selfcal1*')
clean(vis='Antennae_North.cal.ms.contsub',
imagename='Antennae_North.CO3_2Line.Clean.selfcal1',
spw='0',field='',phasecenter='12',
mode='velocity',outframe='LSRK',restfreq='345.79599GHz',
nchan=70,start='1200km/s',width='10km/s',
imagermode='mosaic',usescratch=False,
imsize=500,cell='0.13arcsec',minpb=0.2,
interactive=T,
mask='Antennae_North.CO3_2Line.Clean.mask',
weighting='briggs',robust=0.5,
niter=10000, threshold='5.0mJy')
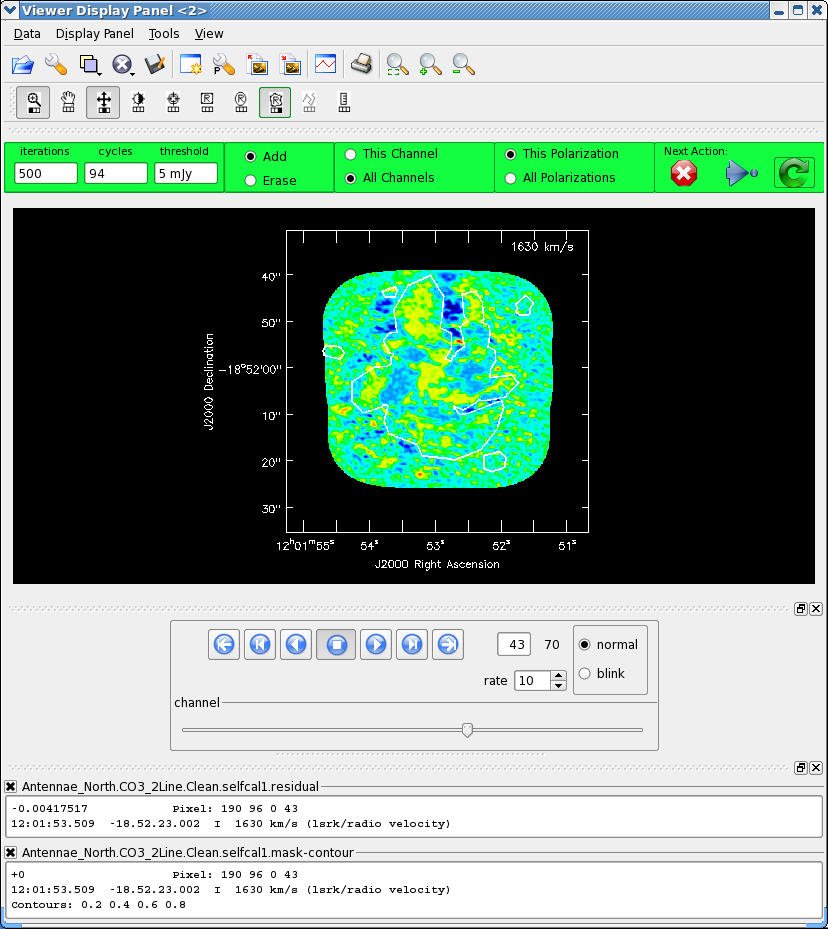
After 600 iterations or so, inspect each channel to see if there is more emission that needs to be included in the mask. Remember to select the "all channels" toggle. Stop when you feel you have cleaned most of the real emission out. This took us a few thousand iterations - we were not so careful to tightly box things and proceed slowly, as we are not going to self-cal again. If you were then you would want to be much more conservative!
Compare the un-selfcal'ed and selfcal'ed data to determine if there has been improvement:
# In CASA
imview (raster=[{'file': 'Antennae_North.CO3_2Line.Clean.image',
'range': [-0.07,0.4]},
{'file': 'Antennae_North.CO3_2Line.Clean.selfcal1.image',
'range': [-0.07,0.4]}],
zoom={'channel': 43})
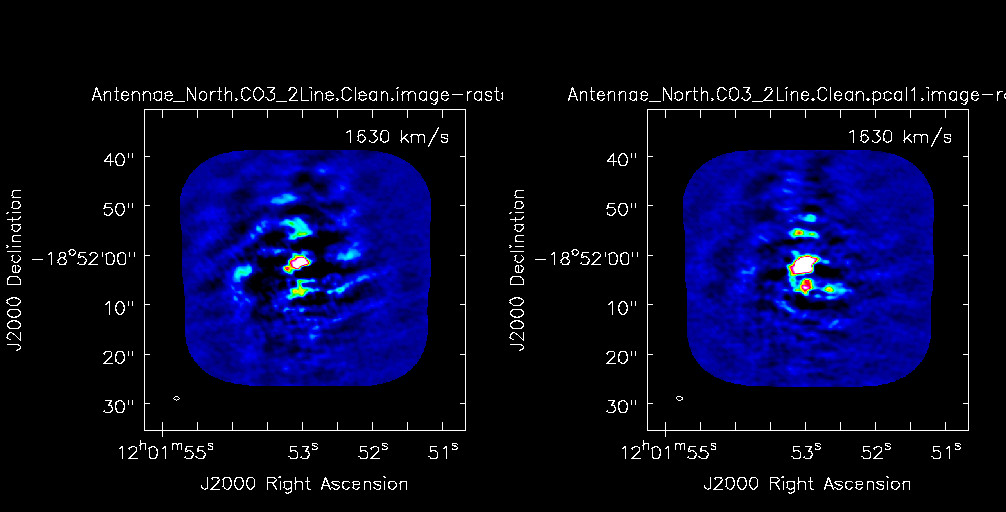
Then in the viewer, select the blink toggle in the "tapedeck", now the tapedeck buttons cycle between the two images at the selected channel. To change the channel toggle back to "normal", change the channel number, and then toggle back to "blink". To make the figure shown in <xr id="north_ch43compare.png"/>, you can select the "p wrench" icon and change the number of panels in x to 2. You will need to drag your viewer window rather wide to get both panels to fit well. If you want to compare other channels, you need to toggle blink off, then use the tapedeck to move to the new channel of interest, and then turn blink back on to see both cubes again.
Note: In "blink" mode, statistics will be reported for the image currently displayed. In "normal" mode, all open raster images will have their statistics reported.
What you should notice is that the peak flux density has increased substantially while the rms noise is about the same. This is reasonable for this self-calibration case because we have mostly adjusted relative position offsets between the datasets and not taken out any short-term phase variations which would reduce the rms.
Note that attempts to push the selfcal to shorter solint (for example to each scan of field=12) did not yield additional improvement.
<figure id="north_ch43compare.png">
</figure>
Image Analysis
Primary Beam Correction
<figure id="viewer.Antennae_North.CO3_2Line.Clean.selfcal1.flux.ch42.png">
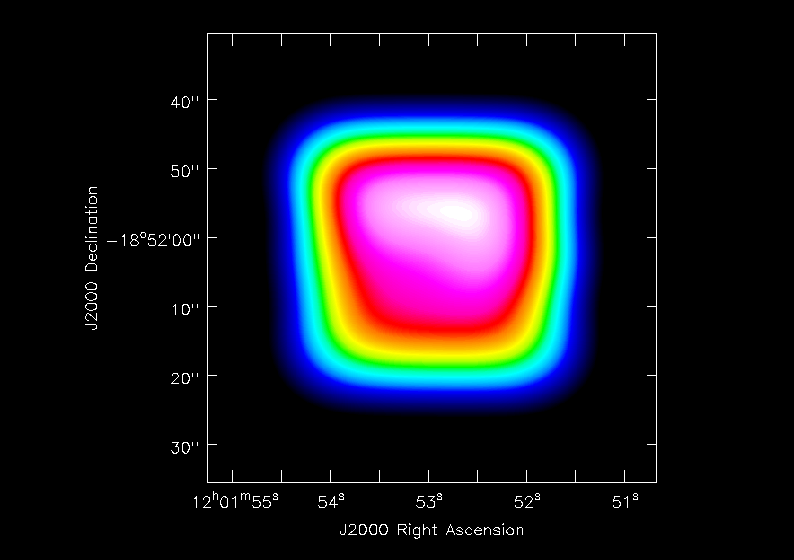
</figure>
The images that came out of clean were made to have "flag-noise" across the mosaic. Before doing any analysis where we care about the actual flux densities we need to correct for the effective mosaic "primary beam" fall-off (which includes the mosaic field sampling, weighting, and the telescope primary beam). This is stored in the .flux cube produced by clean:
# In CASA
viewer('Antennae_North.CO3_2Line.Clean.selfcal1.flux',
channel=42,
outfile='viewer.Antennae_North.CO3_2Line.Clean.selfcal1.flux.ch42.png')
]
Inspection of this (see <xr id="viewer.Antennae_North.CO3_2Line.Clean.selfcal1.flux.ch42.png"/> for the channel 42 plot) shows that there are indeed significant variations across the mosaic that need to be accounted for.
We use immath to do this:
# In CASA
os.system('rm -rf Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage')
immath(outfile='Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage',
mode='evalexpr',
imagename=['Antennae_North.CO3_2Line.Clean.selfcal1.image',
'Antennae_North.CO3_2Line.Clean.selfcal1.flux'],
expr="IM0[IM1>0.2]/IM1")
The expr tells the task to divide the .image cube by the .flux cube.
If you throw this up in the viewer and get statistics in a line-filled channel (e.g. channel 42) in a relatively quiet area of the central part of the mosaic, you find a standard deviation of around 18 mJy (much if this is real emission).
Moment Maps
<figure id="Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom0.png">
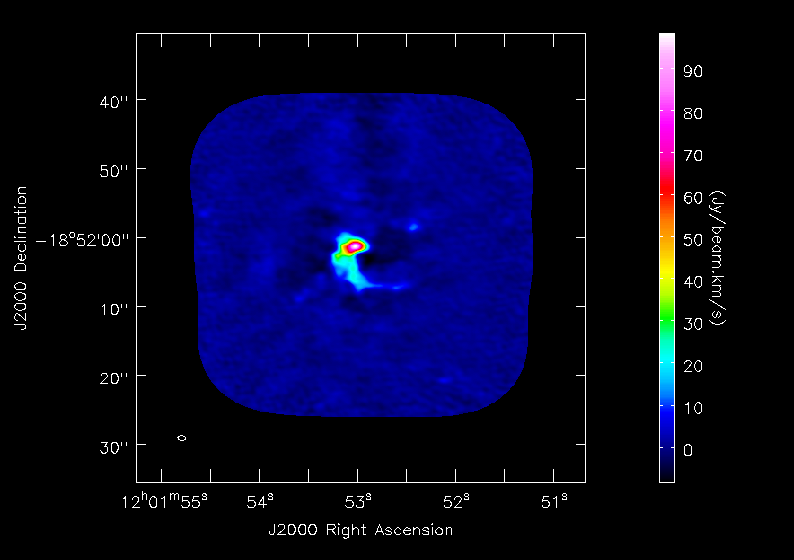
</figure>
<figure id="Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom1.png">
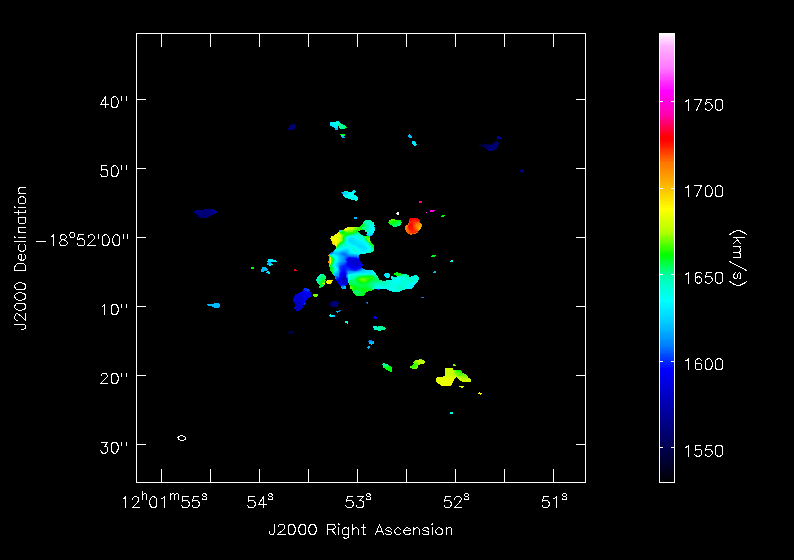
</figure>
<figure id="Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom2.png">
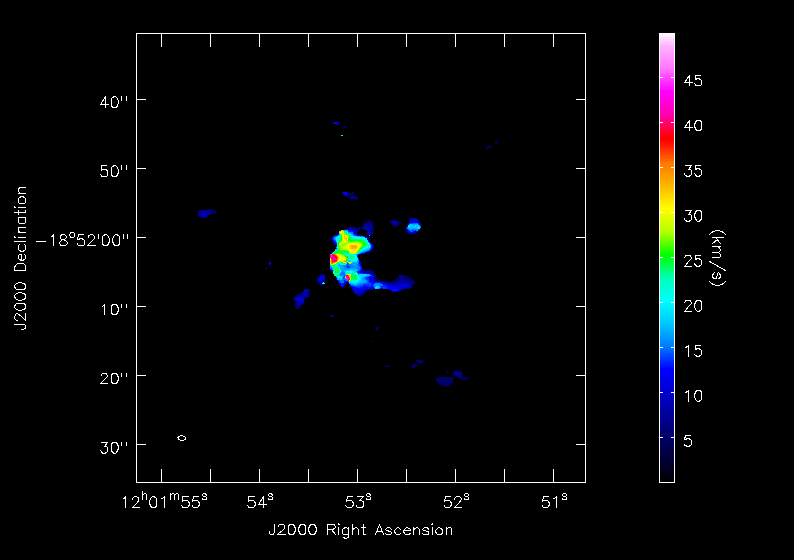
</figure>
Next we will make moment maps for the CO(3-2) emission: Moment 0 is the integrated intensity; Moment 1 is the intensity weighted velocity field; and Moment 2 is the intensity weighted velocity dispersion.
Above we determined the rms noise levels for the pb-corrected North mosaic in a line-bright channel (18 mJy). We want to limit the channel range of the moment calculations to those channels with significant emission. One good way to do this is to open the cube in the viewer overlaid with 3-sigma contours:
# In CASA
imview(raster={'file': 'Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage',
'range': [-0.04,0.4]},
contour={'file': 'Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage',
'levels': [3],'unit': 0.018})
We find a channel range for significant emission of 33~63. Note that because of the pb-correction there are occasional areas of emission even at 3-sigma near the edge where the noise is lifted.
For moment 0 (integrated intensity) maps you do not typically want to set a flux threshold because this will tend to noise bias your integrated intensity.
# In CASA
immoments('Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage',
moments=[0],chans='33~63',
outfile='Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom0')
For higher order moments it is very important to set a conservative flux threshold. For our image, we use 3-sigma, using sigma of 18 mJy from a bright line channel. We do this with the mask parameter in the commands below. When making multiple moments, immoments appends the appropriate file name suffix to the value of outfile.
# In CASA
immoments('Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage',
moments=[1,2], chans='33~63',
mask='Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage>0.018*3',
outfile='Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom')
Next we can display the moment maps (<xr id="Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom0.png"/>, <xr id="Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom1.png"/>, <xr id="Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom2.png"/>, from these images using imview.
# In CASA
os.system('rm -f Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom0.png')
imview (raster={'file': 'Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom0',
'colorwedge':T,'scaling': -0.5},
zoom=1,out='Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom0.png')
#
os.system('rm -f Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom1.png')
imview (raster={'file': 'Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom.weighted_coord',
'colorwedge':T},
zoom=1,out='Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom1.png')
#
os.system('rm -f Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom2.png')
imview (raster={'file': 'Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom.weighted_dispersion_coord',
'colorwedge':T},
zoom=1,out='Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom2.png')
If you want to look at these interactively, leave off the out file parameter from the call.
Exporting Images to Fits
If you want to analyze the data using another software package it is easy to convert from CASA format to FITS.
# In CASA
exportfits(imagename='Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage',
fitsimage='Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.fits')
exportfits(imagename='Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom0',
fitsimage='Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom0.fits')
exportfits(imagename='Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom.weighted_coord',
fitsimage='Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom1.fits')
exportfits(imagename='Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom.weighted_dispersion_coord',
fitsimage='Antennae_North.CO3_2Line.Clean.selfcal1.pbcorimage.mom2.fits')
Although "FITS format" is supposed to be a standard, in fact most packages expect slightly different things from a FITS image. If you are having difficulty, try setting velocity=T and/or dropstokes=T.
Comparison with previous SMA CO(3-2) data
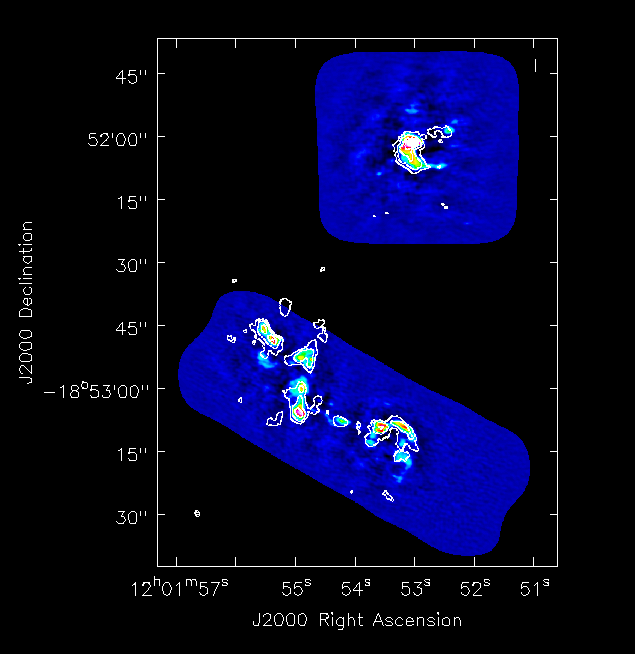
We compare with SMA CO(3-2) data (Ueda, Iono, Petitpas et al. 2012, ApJ, 745, 65). <xr id="alma_sma_compare.png"/> shows a comparison plot between the moment 0 maps of ALMA and SMA data using the viewer. The fluxes, peak locations, and large scale structure are consistent. Note that in this figure both southern and northern mosaic components have been combined.
<figure id="alma_sma_compare.png">
</figure>
Last checked on CASA Version 3.4.0.