TWHydraBand7 Imaging for CASA 3.3
Prepare Continuum Data for Further Processing
In order to speed up imaging/self-calibration for the continuum data we will average channels together, and then flag the few spectral features. This method works well in the case of only a few spectral lines, while retaining enough channelization to aid in multi-frequency synthesis and being able to remove spectral features in the averaged data. The amount of channel averaging should be adjusted to your particular data. Note for wide bandwidth data it is never a good idea to use the continuum estimate generated by uv-continuum subtraction as your continuum data. This worked OK in the past with the narrow band systems like the VLA and low dynamic range instruments like the SMA, but is not advised for ALMA wide bandwidth, high dynamic range data.
One can also flag the spectral features first but in that case make a backup of the file that is analogous to TWHydra_corrected.ms as you will need it later (unflagged) for spectral line imaging.
# In CASA
split(vis='TWHydra_corrected.ms',outputvis='TWHydra_cont.ms',
spw='0~3:22~3820',width=100,datacolumn='data')
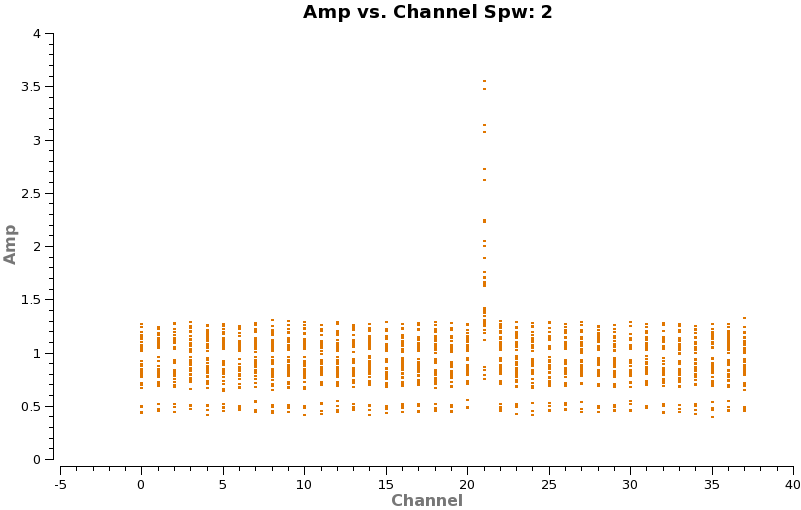
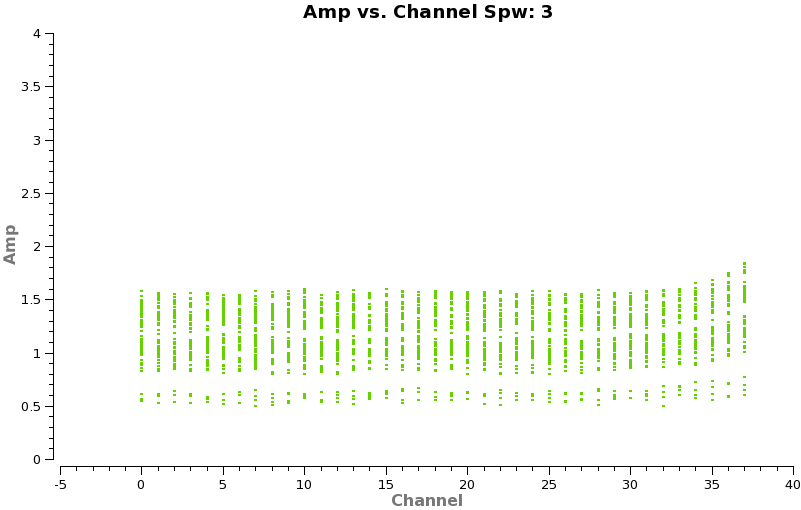
Now make a plot of amplitude vs. channel to see what needs to be flagged
# In CASA
plotms(vis='TWHydra_cont.ms',spw='0~3',xaxis='channel',yaxis='amp',
avgtime='1e8',avgscan=T,coloraxis='spw',iteraxis='spw',xselfscale=T)
The CO(3-2) and HCO+(4-3) lines are obvious. The 4th spw; spw=3 also shows an upswing on the highest channels that is problematic, it was seen on the calibrators too and is likely due to a weak atmospheric feature, we will flag the worst of it.
# In CASA
flagdata(vis='TWHydra_cont.ms',
spw=['0:16~16','2:21~21','3:33~37'])
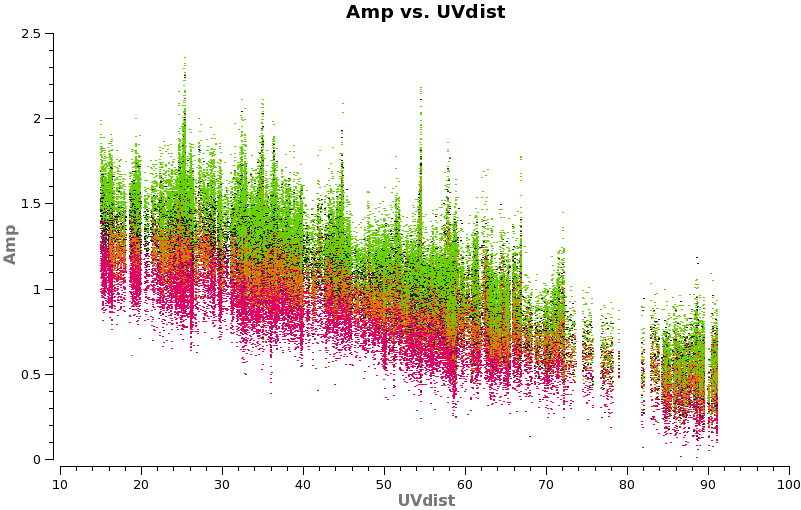
Have a look at the continuum as a function of uv-distance (in the plot below, the 4 spw are shown in different colors on top of each other). If the data are flat as a function of uv-distance the source is completely unresolved. If you see structure (in this case decreasing amplitude with uv-distance) the source is resolved.
# In CASA
plotms(vis='TWHydra_cont.ms',spw='',xaxis='uvdist',yaxis='amp',field='',avgchannel='38',
coloraxis='spw')
Notice that the spw do not have the same amplitude. Unfortunately, the red spw (highest frequency) should be higher than the green one (lowest frequency) because optically thin dust goes as nu^4, and optically thick as nu^2. This inconsistency is due to imperfect amplitude/absolute flux calibration. The self-calibration below will bring them into agreement with each other, but this is the sort of thing that results in considerable absolute flux uncertainty at submillimeter wavelengths. We are working on how to do this better.
Image and Self-Calibrate the Continuum
Make an initial image using clean. We use mode='mfs' to do multifrequency synthesis, in other words grid each uv-channel independently. Since the uv-spacing is a function of frequency, this will actually achieve greater uv-coverage than if all the channels and spws had been averaged in advance. The imsize=100 and cellsize=0.3arcsec were chosen to be appropriate for the Band and antenna configuration. At ALMA Band 7, the Full Width Half Power primary beam is about 20", so we want to make our image at least this big. We've made it a bit larger out to about the 25% power point. For the configuration used to take these data, the resolution is expected to be about 1.5" and it is a good idea to oversample the beam by a factor of 4 or 5 (we used 5 here).
For doing self-calibration it is essential that only "real" signal be included in the image model, thus it is important to make a clean mask, and clean the first iteration rather conservatively (shallowly) so that you are sure that you do not have clean components in your model that do not represent real source emission. In the example below we set interactive=T so you can interactively make the clean mask and stop clean when the signal from the source starts to approach the surrounding "noise" areas in the residual map.
Note: If you start an interactive clean, and then do not make a mask, clean will stop when you tell it to go on because it has nothing to clean. There is no default mask.
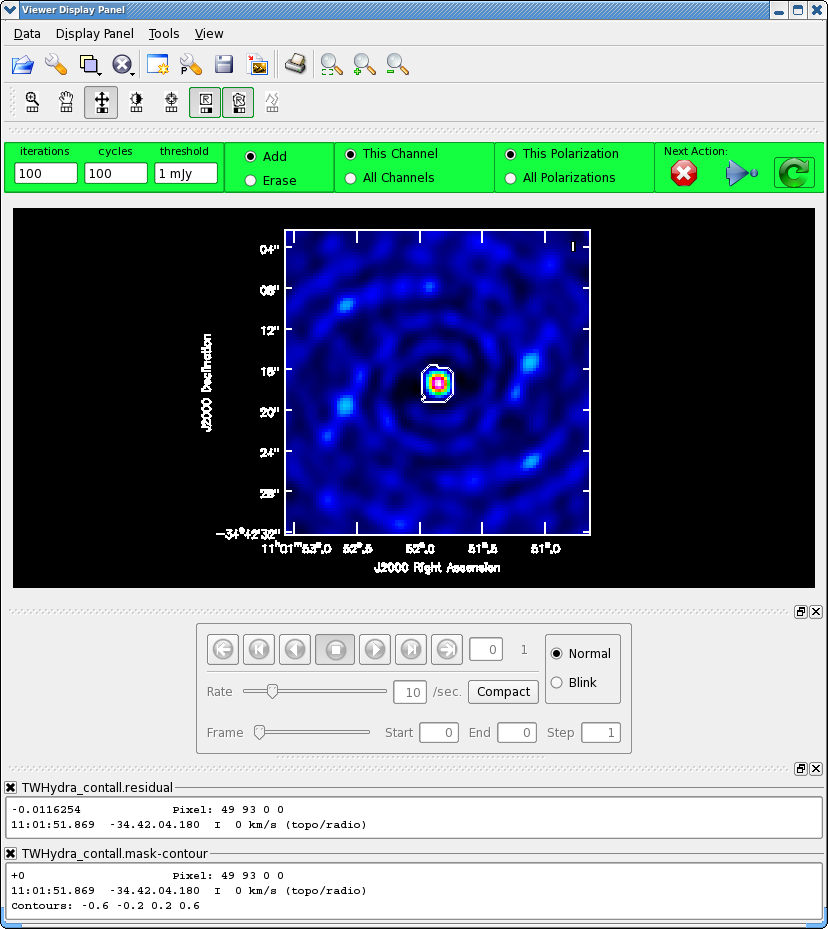
If you are unfamiliar with the CASA viewer it is a good idea to pause here and watch the demo at http://casa.nrao.edu/CasaViewerDemo/casaViewerDemo.html
# In CASA
clean(vis='TWHydra_cont.ms',imagename='TWHydra_contall',
mode='mfs',calready=T,imagermode='csclean',
imsize=100,cell=['0.3arcsec'],spw='',
weighting='briggs',robust=0.5,
mask='',
interactive=T,threshold='1mJy',niter=10000)
Once the Clean Viewer opens, click on the polygon tool with your left mouse button, and draw a polygon around the continuum eimssion with the left mouse button. Double click inside the region with the left mouse button when you are happy with it. It will turn from green to white when the clean mask is accepted. To continue with clean use the "Next action" buttons in the green area on the Viewer Display GUI: The red X will stop clean where you are, the blue arrow will stop the interactive part of clean, but continue to clean non-interactively until reaching the stopping niter or threshold (whichever comes first), and the green circle arrow will clean until it reaches the "iterations" parameter on the left side of the green area. You should chose the green circle arrow. It will clean 100 iterations (number under iteration in the green area on the Viewer gui) and then show the residual image again. The middle mouse button is automatically assigned to the "plus" icon, this adjust the colorscale. If you cannot easily access multiple mouse buttons, you can always assign your one mouse button the icon you want by clicking on it. Adjust the color so you can see it better. Also note that the logger is telling the amount of clean flux recovered so far, the min/max residuals, and the synthesize beam:
Model 0: max, min residuals = 0.0203202, 0.00159066 clean flux 1.31662 Threshhold not reached yet. Beam used in restoration: 1.69902 by 1.59083 (arcsec) at pa 22.3441 (deg)
Already with only 100 iterations, the residuals outside the clean mask are larger than that inside. So we will stop here by hitting the Red X symbol.
Next we can use the clean model to self-calibrate the uv-data. The process of self-calibration tries to adjust the data to better match the model you give it. That is why it is important to make the model as good as you can. The parameter calready=T ensured that a uv-model of the resulting image was placed in the model data column. This model can then be used to self-calibrate the data using gaincal. We use gaintype='T' to tell it to average the polarizations before deriving solutions which gains us a sqrt(2) in sensitivity. A critical parameter to play with is solint, we use solint=30s to gain a bit of sensitivity. From listobs the integration time of these data are about 10 seconds, using solint=30s then gives us a sqrt(3) in sensitivity. It is often best to start with a larger solint and then work your way down to the integration time if the S/N of the permits. For the same reasons as described in the TWHydraBand7_Calibration tutorial we want to fix up the phases before tackling amplitude by setting calcode='p'.
# In CASA
gaincal(vis='TWHydra_cont.ms',caltable='self_1.pcal',
solint='30s',combine='',gaintype='T',
refant='DV06',spw='',minblperant=4,
calmode='p',minsnr=2)
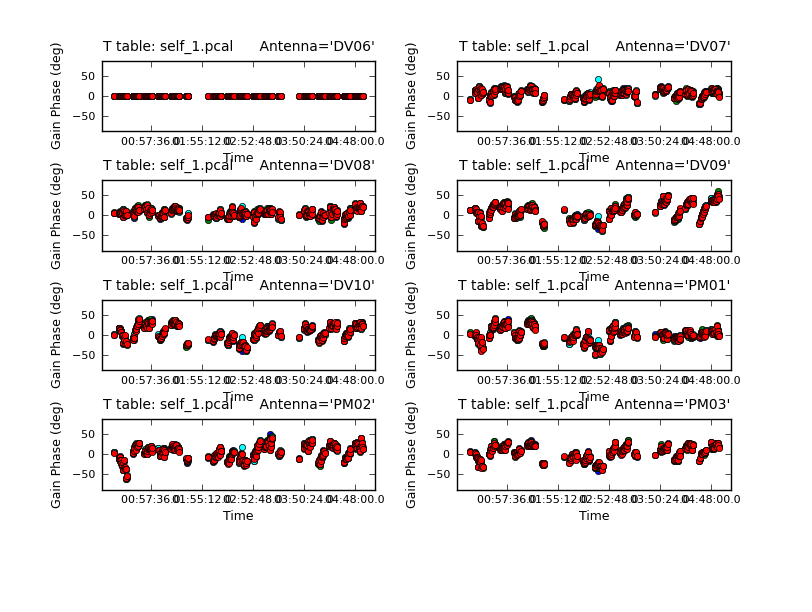
Because combine=, we are getting independent solutions for each spw. If you are desperate for S/N, you can try combining them with combine='spw'. We have plenty of S/N in these data so we keep them separate. Look at the solutions.
# In CASA
plotcal(caltable='self_1.pcal',xaxis='time',yaxis='phase',
spw='',field='',antenna='1~8',iteration='antenna',
subplot=421,plotrange=[0,0,-80,80],figfile='self_1_phase.png')
The magnitude of corrections gives you a sense of how accurate the phase transfer from the calibrators was. You are checking to see that the phases are being well tracked by the solutions, i.e. smoothly varying functions of time within a given scan. Also the solutions for each spw are lying basically on top of each other which indicates that the spw to spw phase offsets are well calibrated already. Overall, these solutions look quite good. If they didn't you could try increasing the solint and/or combining the spw into a single solution.
After checking that you are happy with the solutions, apply them with applycal
# In CASA
applycal(vis='TWHydra_cont.ms',field='',
gaintable=['self_1.pcal'],calwt=F)
Next clean the source again, to save time we can start with the clean mask from the last iteration. If you make adjust the clean mask they will be written to the new mask TWHydra_contall_1pcal.mask, and not the old one.
# In CASA
clean(vis='TWHydra_cont.ms',imagename='TWHydra_contall_1pcal',
mode='mfs',calready=T,imagermode='csclean',
imsize=100,cell=['0.3arcsec'],spw='',
weighting='briggs',robust=0.5,
mask='TWHydra_contall.mask',
interactive=T,threshold='1mJy',niter=10000)
Now after the first 100 iterations you see that the residuals inside the clean mask are higher than those outside. This is because flux that had been spread out in the map due to poorly correlated phases has been increased where it belongs on the source. You may also notice that your original clean mask is a little too small. If so you can make it a bit larger, just by drawing again around where you want the new mask to be. When you double click it will incorporate the new mask. You might want to use the magnifying glass to zoom in. Again you assign mouse buttons but clicking with it on the icon you want. Clean another 100 iterations and stop.
Now open
Now that we have a better model, we can try making the solint smaller. We'll try doing it on the integration time by setting solint='int'
# In CASA
gaincal(vis='TWHydra_cont.ms',caltable='self_2.pcal',
solint='int',combine='',gaintype='T',
refant='DV06',spw='',minblperant=4,
calmode='p',minsnr=2)
Now plot the new solutions.
# In CASA
plotcal(caltable='self_2.pcal',xaxis='time',yaxis='phase',
spw='',field='',antenna='1~8',iteration='antenna',
subplot=421,plotrange=[0,0,-80,80],figfile='self_2_phase.png')
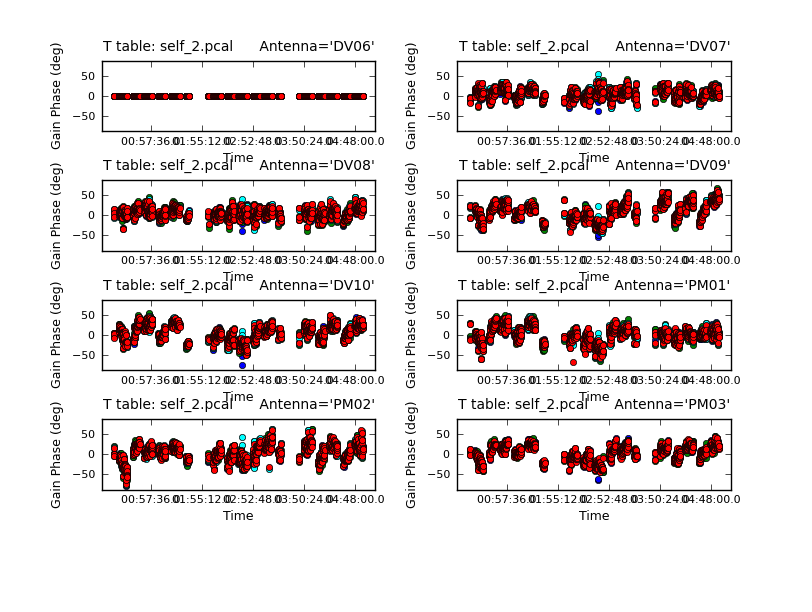
This shows a bit more structure in the phase, but there does appear to be enough S/N to use the shorter solint. It may help to zoom in on one of the plotcal panels with the magnify icon so you can see it better. Alternatively you can set subplot=111 and page through each plot.
Having decided this new iteration of phase-only self-cal was sucessful, we apply it.
# In CASA
applycal(vis='TWHydra_cont.ms',field='',
gaintable=['self_2.pcal'],calwt=F)
Now clean again, updating to start with the most recent mask file.
# In CASA
clean(vis='TWHydra_cont.ms',imagename='TWHydra_contall_2pcal',
mode='mfs',calready=T,imagermode='csclean',
imsize=100,cell=['0.3arcsec'],spw='',
weighting='briggs',robust=0.5,
mask='TWHydra_contall_1pcal.mask',
interactive=T,threshold='1mJy',niter=10000)
This time clean a total of about 250 iterations before stopping (adjust niter in the gui).
Model 0: max, min residuals = 0.00407018, -0.00397106 clean flux 1.48179
# In CASA
gaincal(vis='TWHydra_cont.ms',caltable='self_ap.cal',
solint='inf',combine='',gaintype='T',
refant='DV06',spw='',minblperant=4,
gaintable=['self_2.pcal'],
calmode='ap',minsnr=2,)
clean(vis='TWHydra_cont.ms',imagename='TWHydra_contall_2pcal',
mode='mfs',calready=T,imagermode='csclean',
imsize=100,cell=['0.3arcsec'],spw=,
weighting='briggs',robust=0.5,
mask='TWHydra_contall_1pcal.mask',
interactive=T,threshold='1mJy',niter=10000)
# In CASA
plotcal(caltable='self_ap.cal',xaxis='time',yaxis='phase',
spw='',field='',antenna='1~8',
iteration='antenna',subplot=421,plotrange=[0,0,-1,1])
# In CASA
plotcal(caltable='self_ap.cal',xaxis='time',yaxis='amp',
spw='',field='',antenna='1~8',
iteration='antenna',subplot=421)
# In CASA
applycal(vis='TWHydra_cont.ms',field='',
gaintable=['self_2.pcal','self_ap.cal'],calwt=F)
# In CASA
plotms(vis='TWHydra_cont.ms',xaxis='uvdist',yaxis='amp',
avgchannel='38',ydatacolumn='corrected',iteraxis='spw')
# In CASA
plotms(vis='TWHydra_cont.ms',xaxis='time',yaxis='amp',iteraxis='spw',
ydatacolumn='corrected')
# In CASA
clean(vis='TWHydra_cont.ms',imagename='TWHydra_contall_apcal',
mode='mfs',calready=T,imagermode='csclean',
imsize=100,cell=['0.3arcsec'],spw='',
weighting='briggs',robust=0.5,
mask='TWHydra_contall_2pcal.mask',
interactive=T,threshold='1mJy',niter=10000)
Continuum Subtraction and Spectral Line Imaging
Make channel plot to get line-free channel ranges. Since the line emission is simple in this case and there are more than enough line-free channels we do not need to be very precise just stay well away from the line and best to chose ranges on either side.
# In CASA
split(vis='TWHydra_corrected.ms',outputvis='TWHydra_CO3_2.ms',
datacolumn='corrected',spw='2')
# In CASA
uvcontsub2(vis='TWHydra_CO3_2.ms',fitorder=1,fitspw='0:20~2000,0:2400~3800')
# In CASA
clean(vis='TWHydra_CO3_2.ms.contsub',imagename='TWHydra_CO3_2line',
imagermode='csclean',calready=T,spw='',
imsize=100,cell=['0.3arcsec'],
mode='velocity',start='-4km/s',width='0.12km/s',nchan=118,
restfreq='345.79599GHz',outframe='LSRK',
weighting='briggs',robust=0.5,
mask='',
interactive=T,threshold='1mJy',niter=100000)
The viewer can be used to make nice figures. In this example we opened the CO(3-2) cube as a raster image and then the continuum as a contour image. Then we used the "wrench" icon and "P wrench" icons to set up the channel images, contour levels etc.
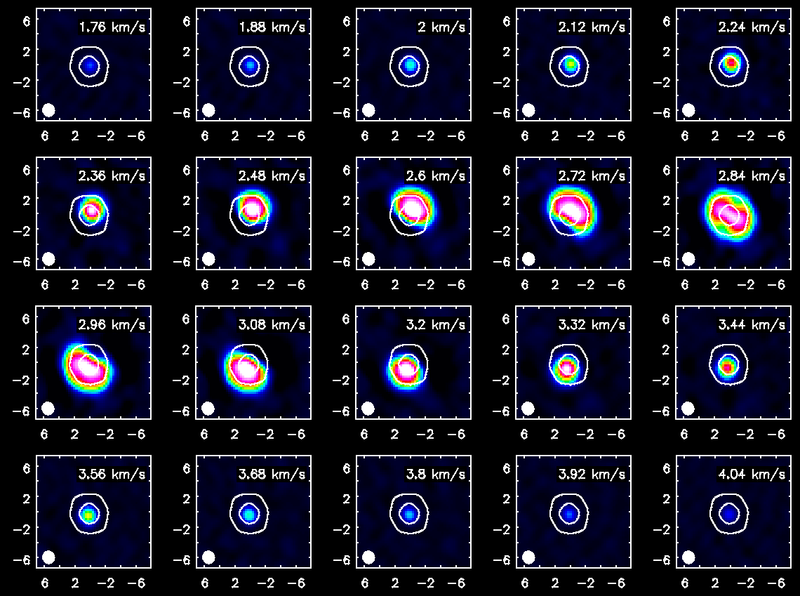
Channel maps of the CO(3-2) emission with white continuum contours at 3 and 100 sigma.
Now repeat for the HCO+(4-3) line.
# In CASA
split(vis='TWHydra_corrected.ms',outputvis='TWHydra_HCOplus.ms',
datacolumn='corrected',spw='0')
# In CASA
uvcontsub2(vis='TWHydra_HCOplus.ms',fitorder=1,fitspw='0:20~1500,0:1900~3800')
# In CASA
clean(vis='TWHydra_HCOplus.ms.contsub',imagename='TWHydra_HCOplusline',
imagermode='csclean',calready=T,spw='',
imsize=100,cell=['0.3arcsec'],
mode='velocity',start='-4km/s',nchan=118,width='0.12km/s',
restfreq='356.7342GHz',outframe='LSRK',
weighting='briggs',robust=0.5,
mask='',
interactive=T,threshold='1mJy',niter=100000)
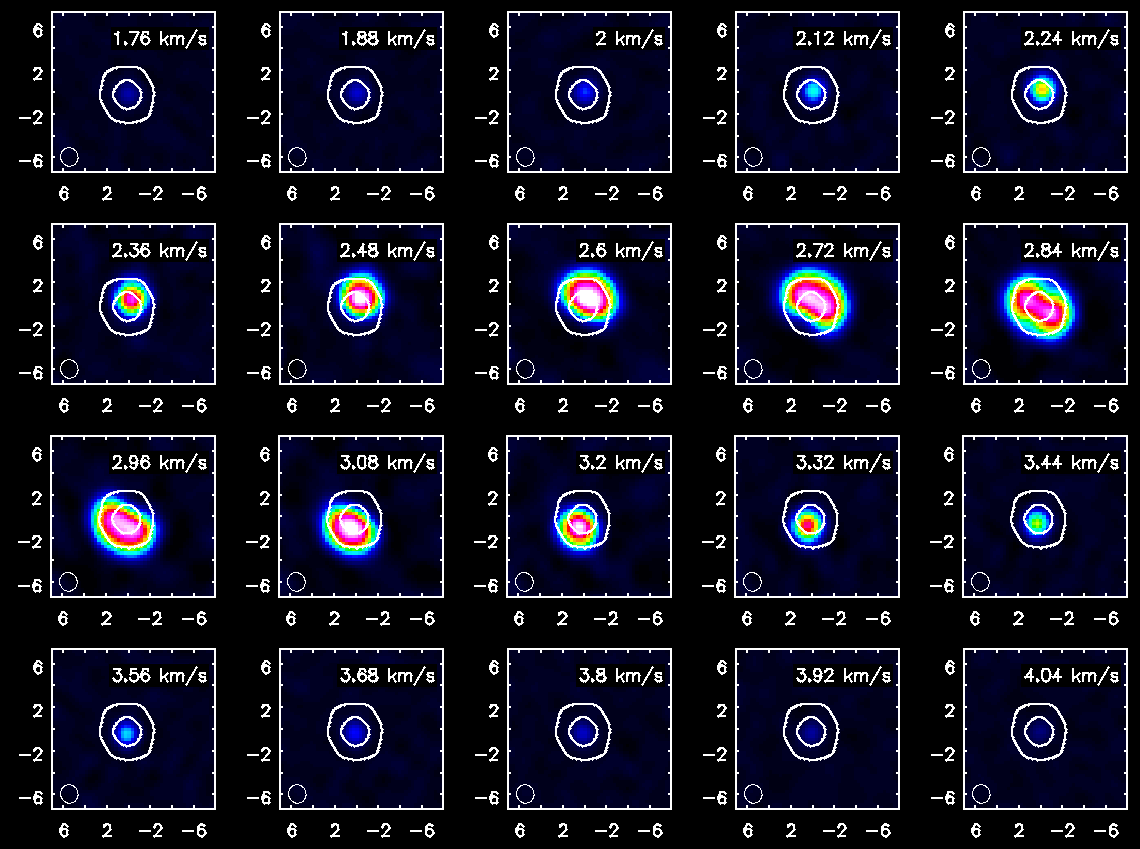
Channel maps of the HCO+(4-3) emission with white continuum contours at 3 and 100 sigma. The color intensity scale is the same as that used for CO(3-2).
Further Image Analysis
To determine channel ranges it is useful to open cube in viewer as a raster. Using the statistics tool the rms in a line free channel is about 25 mJy/beam. Now open the cube again but as a contour map and set the Unit Contour Level to 0.025 (this is in Jy/beam) and in Relative Contour levels set [3]. i.e. this is showing 3sigma contours. Now find out where the CO(3-2) emission drops below the 3sigma level.
For moment 0 (integrated intensity) maps you do not typically want to set a flux threshold because this will tend to noise bias your integrated intensity.
# In CASA
immoments(imagename='TWHydra_CO3_2line.image',moments=[0],
outfile='TWHydra_CO3_2line.image.mom0',
chans='40~76')
For higher order moments it is very important to set a conservative flux threshold. Typically something like 10sigma works well.
# In CASA
immoments(imagename='TWHydra_CO3_2line.image',moments=[1,2],
outfile='TWHydra_CO3_2line.image.mom',
chans='40~76',includepix=[0.25,100])
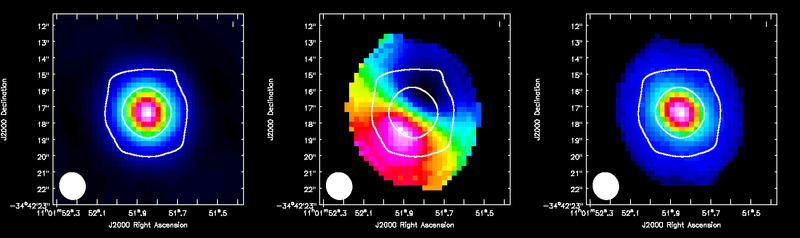
ALMA CO(3-2) moment maps, with white continuum contours at 3 and 100 sigma. From left to right: integrated intensity, intensity weighted velocity field, intensity weighted velocity dispersion are shown.
Repeat for HCO+(4-3) which has a bit narrower velocity extent than the CO(3-2), but similar rms noise.
# In CASA
immoments(imagename='TWHydra_CO3_2line.image',moments=[0],
outfile='TWHydra_CO3_2line.image.mom0',
chans='43~74')
# In CASA
immoments(imagename='TWHydra_CO3_2line.image',moments=[1,2],
outfile='TWHydra_CO3_2line.image.mom',
chans='43~74',includepix=[0.25,100])
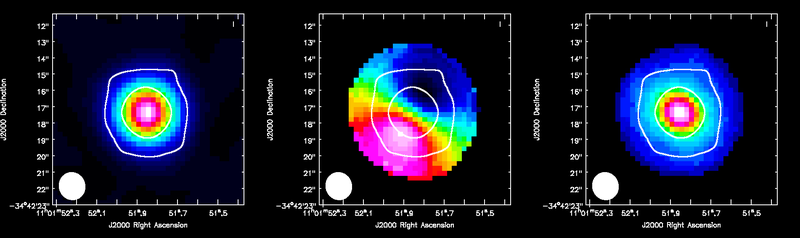
ALMA HCO+(4-3) moment maps, with white continuum contours at 3 and 100 sigma. From left to right: integrated intensity, intensity weighted velocity field, intensity weighted velocity dispersion are shown.