NGC 5921: red-shifted HI emission 5.7.2: Difference between revisions
| Line 233: | Line 233: | ||
* [[#Bandpass Calibration | Determine bandpass corrections]] based on the primary calibrator. In the script that follows, the bandpass calibration is stored in ''ngc5921.demo.bcal'', which is itself referenced by the python variable ''btable.'' | * [[#Bandpass Calibration | Determine bandpass corrections]] based on the primary calibrator. In the script that follows, the bandpass calibration is stored in ''ngc5921.demo.bcal'', which is itself referenced by the python variable ''btable.'' | ||
* [[#Inspect the Bandpass Response Curve | Inspect the bandpass correction]] to determine viable channels for averaging and imaging. The bandpass response will look like a filter response curve, and we want to toss out channels where the response is poor. | * [[#Inspect the Bandpass Response Curve | Inspect the bandpass correction]] to determine viable channels for averaging and imaging. The bandpass response will look like a filter response curve, and we want to toss out channels where the response is poor. | ||
* [[Gain Calibration | Determine the gain calibrations]] on the bandpass-corrected and channel-averaged data. In this step, we effectively turn the spectral line data into a single-channel continuum data set and calibrate accordingly. The calibration is stored in ''ngc5921.demo.gcal'', which is itself referenced by the python variable ''gtable.'' | * [[#Gain Calibration | Determine the gain calibrations]] on the bandpass-corrected and channel-averaged data. In this step, we effectively turn the spectral line data into a single-channel continuum data set and calibrate accordingly. The calibration is stored in ''ngc5921.demo.gcal'', which is itself referenced by the python variable ''gtable.'' | ||
Revision as of 15:16, 7 December 2009
Overview
The technique used to calibrate and image continuum datasets generally applies to spectral line observations, except that an additional calibration step is required. Bandpass calibration flattens the spectral response of the observations, ensuring that spectral channel images are properly calibrated in amplitude and phase.
The following tutorial derives from an annotated script provided in the CASA Cookbook. The script is largely reproduced in sections and additionally annotated with figures and illustrations.
The data are included with the CASA installation.
Setting up the CASA Environment
Start up CASA in the directory you want to use.
# in bash
mkdir NGC5921
cd NGC5921
casapy
Now, in CASA, set up paths and global variables to facilitate the data reduction. These operations can be performed on-the-fly if you are reducing data interactively, but it's better to get them out of the way in the scripting environment.
import os
scriptmode = True
prefix = 'ngc5921.demo' # The prefix to use for all output files
# Set up some useful variables (these will be altered later on)
msfile = prefix + '.ms'
btable = prefix + '.bcal'
gtable = prefix + '.gcal'
ftable = prefix + '.fluxscale'
splitms = prefix + '.src.split.ms'
imname = prefix + '.cleanimg'
We'll use a python os command to get the appropriate CASA path for your installation. The use of os.environ.get is explained in the Appendix.
pathname=os.environ.get('CASAPATH').split()[0]
fitsdata=pathname+'/data/demo/NGC5921.fits'
Scripts are of course modified and repeated to the satisfaction of observer. To help clean up the bookkeeping and further avoid issues of write privileges, remove prior versions of the measurement set and calibration tables.
Tip: The first command in the following code block removes the measurement set. Depending on the size of the source data, refilling a measurement set can be time-consuming, and so you may want to consider editing a separate script that preserves the measurement set but skips the importuvfits command given below. This particular dataset is not very large and so refilling is fairly quick.
rmtables(msfile)
rmtables(btable)
rmtables(gtable)
rmtables(ftable)
rmtables(ftable)
rmtables(splitms+'*')
rmtables(imname+'.*')
rmtables(prefix+'.moments*')
# Final clean up of auxiliary files
os.system('rm -rf '+prefix+'*')
Import the Data
The next step is to import the multisource UVFITS data to a CASA measurement set via the importuvfits filler.
# Safest to start from task defaults
default('importuvfits')
# Set up the MS filename and save as new global variable
msfile = prefix + '.ms'
# Use task importuvfits
fitsfile = fitsdata
vis = msfile
saveinputs('importuvfits',prefix+'.importuvfits.saved')
importuvfits()
Saveinputs saves the parameters of a given command to specified text file, handy to debug a script and see what actually was run. Here, the parameters of importuvfits are saved to the file "ngc5921.demo.importuvfits.saved". Here's a listing of this file. Notice that it is executable with execfile in CASA (remove the # commenting symbol before importuvfits to have the execfile run the command).
CASA <71>: os.system('cat ngc5921.demo.importuvfits.saved')
taskname = "importuvfits"
fitsfile = "/usr/lib64/casapy/30.0.9709test-001/data/demo/NGC5921.fits"
vis = "ngc5921.demo.ms"
antnamescheme = "old"
#importuvfits(fitsfile="/usr/lib64/casapy/30.0.9709test-001/data/demo/NGC5921.fits",vis="ngc5921.demo.ms",antnamescheme="old")
A Summary of the Data
We'll need to have a look at the observing tables to learn the calibrator and source names. The relevant command is listobs.
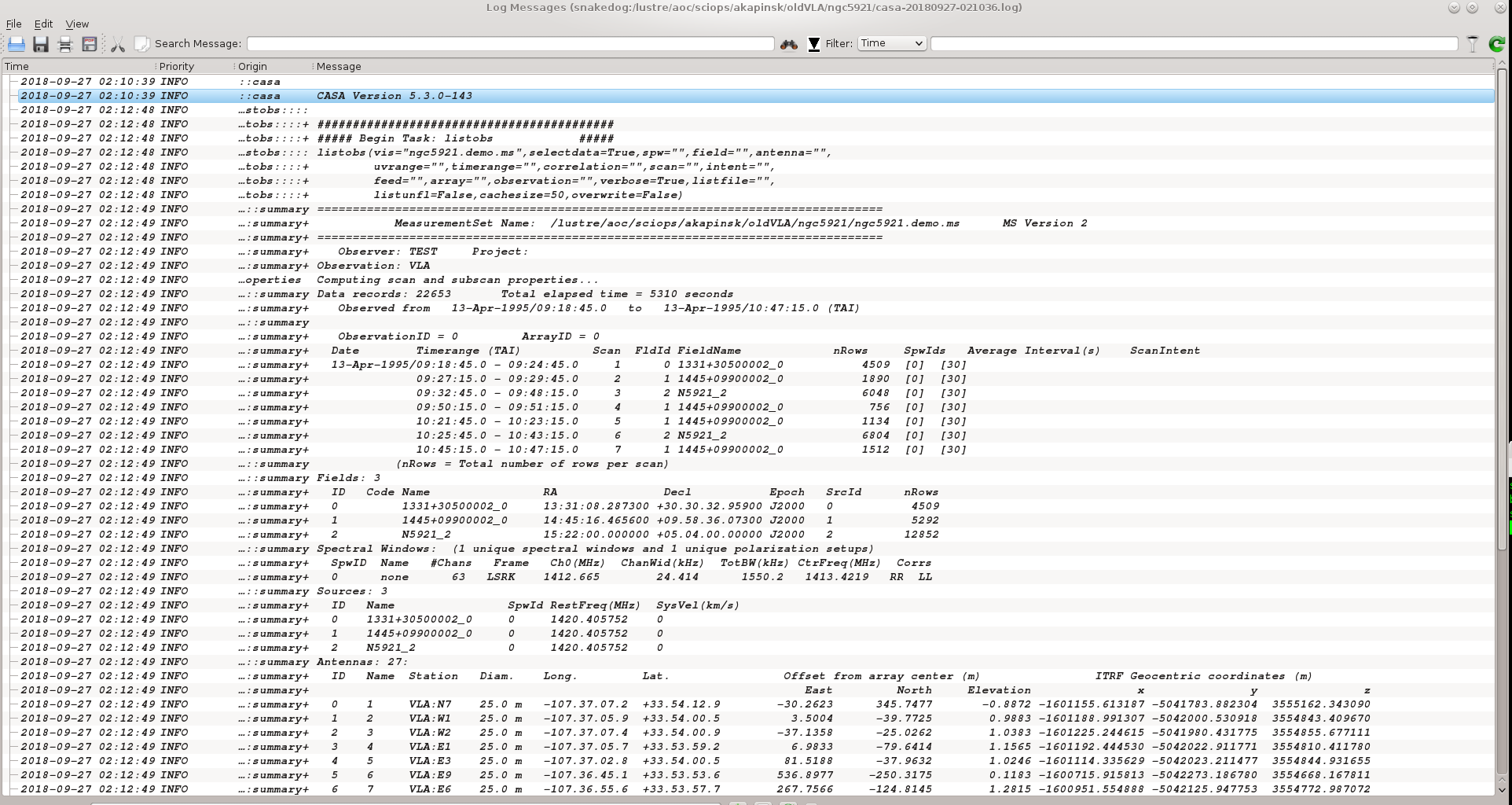
listobs(vis='ngc5921.demo.ms', verbose=True)
The output goes to the logger window; see the screenshot at right.
Tip: You can control the text size of the logger window using <ctrl>-A (smaller font) and <ctrl>-L (larger font).
A more complete listing of the listobs output follows.
INFO listobs ##### Begin Task: listobs #####
INFO listobs::::casa
================================================================================
MeasurementSet Name: /DATA/ASHLESHA_2/NGC5921/ngc5921.demo.ms MS Version 2
================================================================================
Observer: TEST Project:
Observation: VLA
Data records: 22653 Total integration time = 5280 seconds
Observed from 13-Apr-1995/09:19:00.0 to 13-Apr-1995/10:47:00.0 (TAI)
2009-12-03 16:15:38 INFO listobs::ms::summary
ObservationID = 0 ArrayID = 0
Date Timerange (TAI) Scan FldId FieldName nVis Int(s) SpwIds
13-Apr-1995/09:19:00.0 - 09:24:30.0 1 0 1331+305000* 4509 30 [0]
09:27:30.0 - 09:29:30.0 2 1 1445+099000* 1890 30 [0]
09:33:00.0 - 09:48:00.0 3 2 N5921_2 6048 30 [0]
09:50:30.0 - 09:51:00.0 4 1 1445+099000* 756 30 [0]
10:22:00.0 - 10:23:00.0 5 1 1445+099000* 1134 30 [0]
10:26:00.0 - 10:43:00.0 6 2 N5921_2 6804 30 [0]
10:45:30.0 - 10:47:00.0 7 1 1445+099000* 1512 30 [0]
(nVis = Total number of time/baseline visibilities per scan)
Fields: 3
ID Code Name RA Decl Epoch SrcId nVis
0 C 1331+305000* 13:31:08.2873 +30.30.32.9590 J2000 0 4509
1 A 1445+099000* 14:45:16.4656 +09.58.36.0730 J2000 1 5292
2 N5921_2 15:22:00.0000 +05.04.00.0000 J2000 2 12852
(nVis = Total number of time/baseline visibilities per field)
Spectral Windows: (1 unique spectral windows and 1 unique polarization setups)
SpwID #Chans Frame Ch1(MHz) ChanWid(kHz)TotBW(kHz) Ref(MHz) Corrs
0 63 LSRK 1412.66507 24.4140625 1550.19688 1413.42801 RR LL
Sources: 3
ID Name SpwId RestFreq(MHz) SysVel(km/s)
0 1331+305000* 0 1420.405752 0
1 1445+099000* 0 1420.405752 0
2 N5921_2 0 1420.405752 0
Antennas: 27:
ID Name Station Diam. Long. Lat.
0 1 VLA:N7 25.0 m -107.37.07.2 +33.54.12.9
1 2 VLA:W1 25.0 m -107.37.05.9 +33.54.00.5
2 3 VLA:W2 25.0 m -107.37.07.4 +33.54.00.9
3 4 VLA:E1 25.0 m -107.37.05.7 +33.53.59.2
4 5 VLA:E3 25.0 m -107.37.02.8 +33.54.00.5
5 6 VLA:E9 25.0 m -107.36.45.1 +33.53.53.6
6 7 VLA:E6 25.0 m -107.36.55.6 +33.53.57.7
7 8 VLA:W8 25.0 m -107.37.21.6 +33.53.53.0
8 9 VLA:N5 25.0 m -107.37.06.7 +33.54.08.0
9 10 VLA:W3 25.0 m -107.37.08.9 +33.54.00.1
10 11 VLA:N4 25.0 m -107.37.06.5 +33.54.06.1
11 12 VLA:W5 25.0 m -107.37.13.0 +33.53.57.8
12 13 VLA:N3 25.0 m -107.37.06.3 +33.54.04.8
13 14 VLA:N1 25.0 m -107.37.06.0 +33.54.01.8
14 15 VLA:N2 25.0 m -107.37.06.2 +33.54.03.5
15 16 VLA:E7 25.0 m -107.36.52.4 +33.53.56.5
16 17 VLA:E8 25.0 m -107.36.48.9 +33.53.55.1
17 18 VLA:W4 25.0 m -107.37.10.8 +33.53.59.1
18 19 VLA:E5 25.0 m -107.36.58.4 +33.53.58.8
19 20 VLA:W9 25.0 m -107.37.25.1 +33.53.51.0
20 21 VLA:W6 25.0 m -107.37.15.6 +33.53.56.4
21 22 VLA:E4 25.0 m -107.37.00.8 +33.53.59.7
23 24 VLA:E2 25.0 m -107.37.04.4 +33.54.01.1
24 25 VLA:N6 25.0 m -107.37.06.9 +33.54.10.3
25 26 VLA:N9 25.0 m -107.37.07.8 +33.54.19.0
26 27 VLA:N8 25.0 m -107.37.07.5 +33.54.15.8
27 28 VLA:W7 25.0 m -107.37.18.4 +33.53.54.8
2009-12-03 16:15:38 INFO listobs::::casa
##### End Task: listobs #####
##########################################
Key Information from Listobs
Certainly the output of listobs is dense with information, but there are some particularly vital data that we'll need for the calibration.
- The calibrators are 1331+305* (3C286, the flux and bandpass calibrator) and 1445+099* (the phase calibrator). We can use wild-cards 1331* and 1445* since they uniquely identify the sources.
- The calibrator field indices are field='0' (1331+305) and field='1' (1445+099).
- The name of the source in the observations list is N5921_2, or field = '2'.
- The data were taken in a single IF (a single spectral window, SpwID = 0), divided into 63 channels.
- Only RR and LL correlations are present; cross-pols are absent.
Flagging
Flag the autocorrelations
We don't need the autocorrelation data, and flagautocorr gets rid of them. You shouldn't have to specify the measurement set, because the variable vis is already set, but it never hurts to be cautious.
flagautocorr(vis="ngc5921.demo.ms")
Interactive Flagging
Casaplotms is a good tool for flagging spectral line data. Check out the tutorial that describes editing VLA continuum data. The only difference for spectral line data is that it's worth averaging channels, at least for an initial look.
Casaplotms currently runs outside the CASA python shell, but we can use os.system to start it from within CASA. In the following code snippet, the command is commented out to provide a non-interactive run of the script. Uncomment to include interactive data editing as part of the script.
# os.system("casaplotms")
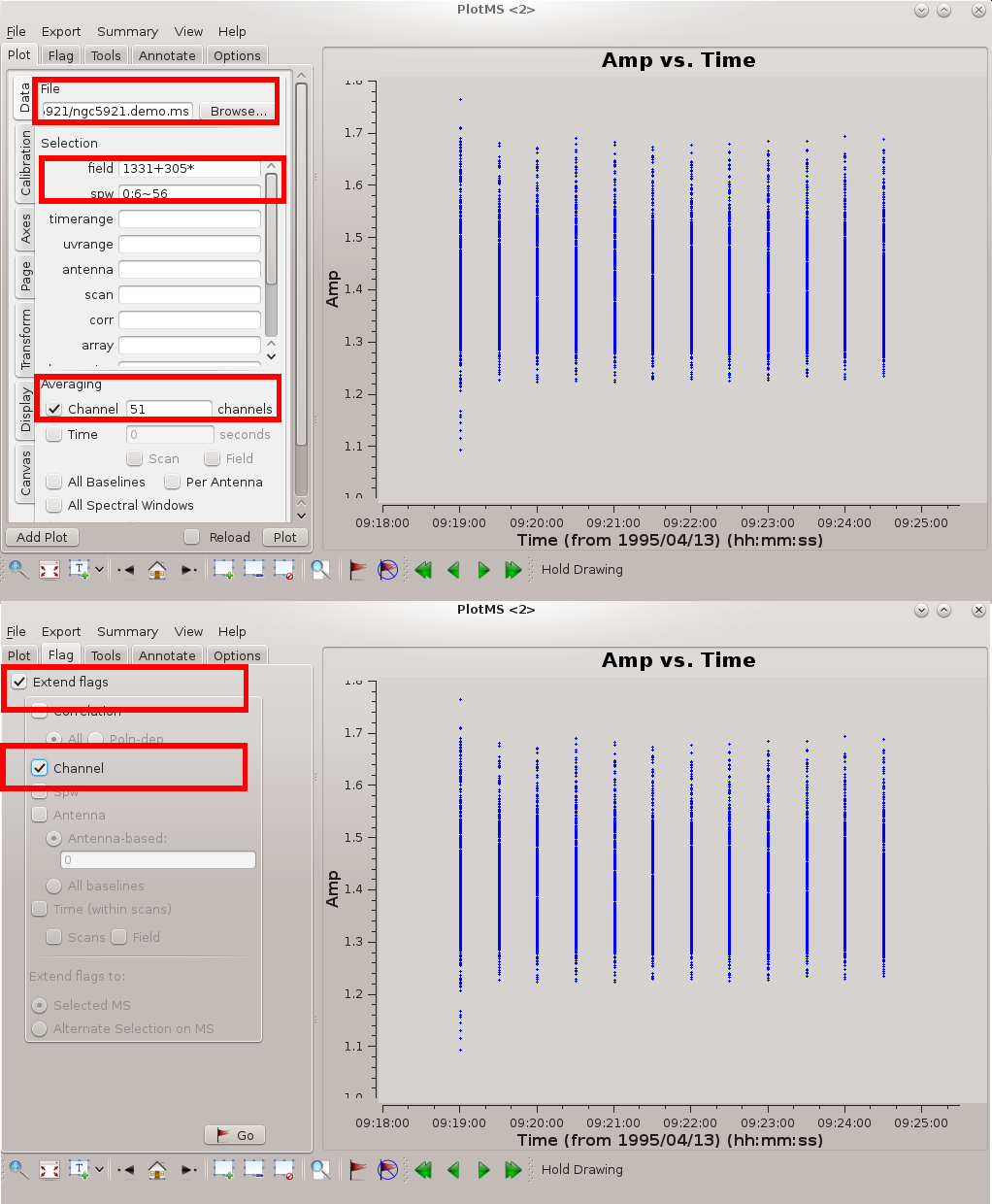
The figure at right highlights the settings needed for effective editing of a spectral line data set. The key settings are as follows.
- Specify the measurement set in File Location; the Browse button allows you to hunt down the measurement set.
- It's better to edit one source at a time. In the illustrated example, the flux / bandpass calibrator 1331+305* is displayed.
- Specify channel averaging. At the time of this writing casaplotms was not particularly versatile at specifying channel ranges, and so for now the specification is just an average over all channels. Ultimately you should be able to target specific ranges of channels.
- Ideally you want the channels to have the same (u, v) coverage (projected baseline spacings as viewed from the source); otherwise, the beam (point spread function) will be different for each channel. Therefore, if you flag data from a given channel it's usually a good idea to flag those data from all channels. Under the Flagging tab, specify Extend flags to Channel.
With these settings, interactive flagging proceeds as for continuum data. When you're satisfied with the edits, File → Quit to return to the CASA prompt.
Calibration
Calibration of spectral line data broadly follows the approach for continuum data, except that the amplitude and phase corrections are a function of frequency and so must be corrected by bandpass calibration. The basic calibration steps follow.
- Set the flux scale of the primary calibrator, here, 1331+305 = 3C 286.
- Determine bandpass corrections based on the primary calibrator. In the script that follows, the bandpass calibration is stored in ngc5921.demo.bcal, which is itself referenced by the python variable btable.
- Inspect the bandpass correction to determine viable channels for averaging and imaging. The bandpass response will look like a filter response curve, and we want to toss out channels where the response is poor.
- Determine the gain calibrations on the bandpass-corrected and channel-averaged data. In this step, we effectively turn the spectral line data into a single-channel continuum data set and calibrate accordingly. The calibration is stored in ngc5921.demo.gcal, which is itself referenced by the python variable gtable.
Setting the Flux Scale
print '--Setjy--'
default('setjy')
vis = msfile
#
# 1331+305 = 3C286 is our primary calibrator
# Use the wildcard on the end of the source name
# since the field names in the MS have inherited the
# AIPS qualifiers
field = '1331+305*'
# This is 1.4GHz D-config and 1331+305 is sufficiently unresolved
# that we dont need a model image. For higher frequencies
# (particularly in A and B config) you would want to use one.
modimage = ''
# Setjy knows about this source so we dont need anything more
saveinputs('setjy',prefix+'.setjy.saved')
# Pause script if you are running in scriptmode
if scriptmode:
inp()
user_check=raw_input('Return to continue script\n')
setjy()
Bandpass Calibration
#=====================================================================
#
# Bandpass calibration
#
print '--Bandpass--'
default('bandpass')
# We can first do the bandpass on the single 5min scan on 1331+305
# At 1.4GHz phase stablility should be sufficient to do this without
# a first (rough) gain calibration. This will give us the relative
# antenna gain as a function of frequency.
vis = msfile
# set the name for the output bandpass caltable
btable = prefix + '.bcal'
caltable = btable
# No gain tables yet
gaintable = ''
gainfield = ''
interp = ''
# Use flux calibrator 1331+305 = 3C286 (FIELD_ID 0) as bandpass calibrator
field = '0'
# all channels
spw = ''
# No other selection
selectdata = False
# In this band we do not need a-priori corrections for
# antenna gain-elevation curve or atmospheric opacity
# (at 8GHz and above you would want these)
gaincurve = False
opacity = 0.0
# Choose bandpass solution type
# Pick standard time-binned B (rather than BPOLY)
bandtype = 'B'
# set solution interval arbitrarily long (get single bpass)
solint = 'inf'
combine = 'scan'
# reference antenna Name 15 (15=VLA:N2) (Id 14)
refant = '15'
saveinputs('bandpass',prefix+'.bandpass.saved')
# Pause script if you are running in scriptmode
if scriptmode:
inp()
user_check=raw_input('Return to continue script\n')
bandpass()
Inspect the Bandpass Response Curve
#
#=====================================================================
#
# Use plotcal to examine the bandpass solutions
#
print '--Plotcal (bandpass)--'
default('plotcal')
caltable = btable
field = '0'
# Set up 2x1 panels - upper panel amp vs. channel
subplot = 211
yaxis = 'amp'
# No output file yet (wait to plot next panel)
saveinputs('plotcal',prefix+'.plotcal.b.amp.saved')
if scriptmode:
showgui = True
else:
showgui = False
plotcal()
#
# Set up 2x1 panels - lower panel phase vs. channel
subplot = 212
yaxis = 'phase'
saveinputs('plotcal',prefix+'.plotcal.b.phase.saved')
#
# Note the rolloff in the start and end channels. Looks like
# channels 6-56 (out of 0-62) are the best
# Pause script if you are running in scriptmode
if scriptmode:
# If you want to do this interactively and iterate over antenna, set
# iteration = 'antenna'
showgui = True
plotcal()
user_check=raw_input('Return to continue script\n')
else:
# No GUI for this script
showgui = False
# Now send final plot to file in PNG format (via .png suffix)
figfile = caltable + '.plotcal.png'
plotcal()
Gain Calibration
#=====================================================================
#
# Gain calibration
#
print '--Gaincal--'
default('gaincal')
# Armed with the bandpass, we now solve for the
# time-dependent antenna gains
vis = msfile
# set the name for the output gain caltable
gtable = prefix + '.gcal'
caltable = gtable
# Use our previously determined bandpass
# Note this will automatically be applied to all sources
# not just the one used to determine the bandpass
gaintable = btable
gainfield = ''
# Use nearest (there is only one bandpass entry)
interp = 'nearest'
# Gain calibrators are 1331+305 and 1445+099 (FIELD_ID 0 and 1)
field = '0,1'
# We have only a single spectral window (SPW 0)
# Choose 51 channels 6-56 out of the 63
# to avoid end effects.
# Channel selection is done inside spw
spw = '0:6~56'
# No other selection
selectdata = False
# In this band we do not need a-priori corrections for
# antenna gain-elevation curve or atmospheric opacity
# (at 8GHz and above you would want these)
gaincurve = False
opacity = 0.0
# scan-based G solutions for both amplitude and phase
gaintype = 'G'
solint = 'inf'
combine = ''
calmode = 'ap'
# minimum SNR allowed
minsnr = 1.0
# reference antenna 15 (15=VLA:N2)
refant = '15'
saveinputs('gaincal',prefix+'.gaincal.saved')
if scriptmode:
inp()
user_check=raw_input('Return to continue script\n')
gaincal()
#
#=====================================================================
#
# Bootstrap flux scale
#
print '--Fluxscale--'
default('fluxscale')
vis = msfile
# set the name for the output rescaled caltable
ftable = prefix + '.fluxscale'
fluxtable = ftable
# point to our first gain cal table
caltable = gtable
# we will be using 1331+305 (the source we did setjy on) as
# our flux standard reference - note its extended name as in
# the FIELD table summary above (it has a VLA seq number appended)
reference = '1331*'
# we want to transfer the flux to our other gain cal source 1445+099
transfer = '1445*'
saveinputs('fluxscale',prefix+'.fluxscale.saved')
# Pause script if you are running in scriptmode
if scriptmode:
inp()
user_check=raw_input('Return to continue script\n')
fluxscale()
# In the logger you should see something like:
# Flux density for 1445+09900002_0 in SpW=0 is:
# 2.48576 +/- 0.00123122 (SNR = 2018.94, nAnt= 27)
# If you run plotcal() on the tablein = 'ngc5921.demo.fluxscale'
# you will see now it has brought the amplitudes in line between
# the first scan on 1331+305 and the others on 1445+099
Inspect the Calibration Solutions
#=====================================================================
#
# Now use plotcal to examine the gain solutions
#
print '--Plotcal (fluxscaled gains)--'
default('plotcal')
caltable = ftable
field = '0,1'
# Set up 2x1 panels - upper panel amp vs. time
subplot = 211
yaxis = 'amp'
# No output file yet (wait to plot next panel)
saveinputs('plotcal',prefix+'.plotcal.gscaled.amp.saved')
if scriptmode:
showgui = True
else:
showgui = False
plotcal()
#
# Set up 2x1 panels - lower panel phase vs. time
subplot = 212
yaxis = 'phase'
saveinputs('plotcal',prefix+'.plotcal.gscaled.phase.saved')
#
# The amp and phase coherence looks good
# Pause script if you are running in scriptmode
if scriptmode:
# If you want to do this interactively and iterate over antenna, set
#iteration = 'antenna'
showgui = True
plotcal()
user_check=raw_input('Return to continue script\n')
else:
# No GUI for this script
showgui = False
# Now send final plot to file in PNG format (via .png suffix)
figfile = caltable + '.plotcal.png'
plotcal()
Apply the Solutions
#=====================================================================
#
# Apply our calibration solutions to the data
# (This will put calibrated data into the CORRECTED_DATA column)
#
print '--ApplyCal--'
default('applycal')
vis = msfile
# We want to correct the calibrators using themselves
# and transfer from 1445+099 to itself and the target N5921
# Start with the fluxscale/gain and bandpass tables
gaintable = [ftable,btable]
# pick the 1445+099 out of the gain table for transfer
# use all of the bandpass table
gainfield = ['1','*']
# interpolation using linear for gain, nearest for bandpass
interp = ['linear','nearest']
# only one spw, do not need mapping
spwmap = []
# all channels
spw = ''
selectdata = False
# as before
gaincurve = False
opacity = 0.0
# select the fields for 1445+099 and N5921
field = '1,2'
applycal()
# Now for completeness apply 1331+305 to itself
field = '0'
gainfield = ['0','*']
# The CORRECTED_DATA column now contains the calibrated visibilities
saveinputs('applycal',prefix+'.applycal.saved')
# Pause script if you are running in scriptmode
if scriptmode:
inp()
user_check=raw_input('Return to continue script\n')
applycal()
Appendix: Python Notes
os.system
os.system allows you to run shell commands from within python / CASA. For example:
import os
os.system('ls -sF')
will give an OS-level listing of the current directory's contents.
os.environ.get
It's worth having a look at the output of the os.environ.get command to understand the python syntax (alternative: os.getenv). You can think of os.environ as a list of operating system environment variables, and get is a method that extracts information about the requested environment variable, here, CASAPATH. Get returns a string of whitespace separated information, and .split() turns that string into a list. The array index [0] extracts the first element of that list, which contains the path.
To illustrate, here is some example python I/O in CASA.
CASA <12>: print os.environ.get('CASAPATH')
/usr/lib64/casapy/30.0.9709test-001 linux local el5bld64b
CASA <13>: print os.environ.get('CASAPATH').split()
['/usr/lib64/casapy/30.0.9709test-001', 'linux', 'local', 'el5bld64b']
CASA <14>: print os.environ.get('CASAPATH').split()[0]
/usr/lib64/casapy/30.0.9709test-001