NGC 5921: red-shifted HI emission 6.5.2: Difference between revisions
| (109 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
Disclaimer: Due to continuous CASA software updates, GUI images may look different on more recent versions than those shown here. | Disclaimer: Due to continuous CASA software updates, GUI images may look different on more recent versions than those shown here. | ||
This guide is meant to be used with monolithic CASA and not pip-wheel, because the GUIs are not necessarily validated. | |||
We would like to note that the CASAviewer has not been maintained for a few years anymore and will be removed from future versions of CASA. | |||
The NRAO replacement visualization tool for images and cubes is CARTA, the “Cube Analysis and Rendering Tool for Astronomy”. It is available from the [https://cartavis.org/ CARTA website]. We strongly recommend using CARTA, as it provides a much more efficient, stable, and feature rich user experience. A comparison of the CASAviewer and CARTA, as well as instructions on how to use CARTA at NRAO is provided in the [https://casadocs.readthedocs.io/en/stable/notebooks/carta.html relevant section of CASA docs]. This tutorial shows figures generated with CARTA for visualization. | |||
[[Category: VLA]] [[Category: Calibration]] [[Category: Imaging]] [[Category: Spectral Line]] | [[Category: VLA]] [[Category: Calibration]] [[Category: Imaging]] [[Category: Spectral Line]] | ||
<div style="background-color: #CCFF99;"> | |||
Last checked on CASA Version 6.5.2. | |||
</div> | |||
== Overview == | == Overview == | ||
The technique used to calibrate and image continuum datasets generally applies to spectral line observations, except that an additional calibration step is required. '''Bandpass calibration''' flattens the spectral response of the observations, ensuring that spectral channel images are properly calibrated in amplitude and phase. | The technique used to calibrate and image continuum datasets generally applies to spectral line observations, except that an additional calibration step is required. '''Bandpass calibration''' flattens the spectral response of the observations, ensuring that spectral channel images are properly calibrated in amplitude and phase. | ||
The following tutorial derives from an annotated script provided in the [http://casa.nrao.edu/ref_cookbook.shtml CASA Cookbook] | The following tutorial derives from an annotated script originally provided in the [http://casa.nrao.edu/ref_cookbook.shtml CASA Cookbook]. | ||
'''DATA''': The data are included with the CASA installation. | '''DATA''': The data are included with the CASA installation. | ||
| Line 42: | Line 45: | ||
</source> | </source> | ||
To help clean up the bookkeeping and further avoid issues of write privileges, remove any prior versions of the measurement set and calibration tables. This should be done with the [https://casadocs.readthedocs.io/en/ | To help clean up the bookkeeping and further avoid issues of write privileges, remove any prior versions of the measurement set and calibration tables. This should be done with the [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.manipulation.rmtables.html rmtables] task, either interactively or using ''rmtables('table_name')'', in preference to the operating system ''rm -rf'' command, as rmtables also clears data in the CASA cache. | ||
== Import the data == | == Import the data == | ||
The next step is to import the multisource UVFITS data to a CASA measurement set via the [https://casadocs.readthedocs.io/en/ | The next step is to import the multisource UVFITS data to a CASA measurement set via the [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.data.importuvfits.html importuvfits] task. This task is used to read in data UVFITS data from the legacy VLA or other telescope; for reading in EVLA data in ASDM format the task [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.data.importasdm.html importasdm] would be used. | ||
Note that you can set each parameter for any particular task one-by-one ('interactive CASA', or you can supply the task and input parameters with a single command ('pseudo-interactive CASA'). | Note that you can set each parameter for any particular task one-by-one ('interactive CASA', or you can supply the task and input parameters with a single command ('pseudo-interactive CASA'). | ||
| Line 78: | Line 81: | ||
A few things to note: | A few things to note: | ||
* The first [https://casadocs.readthedocs.io/en/ | * The first [https://casadocs.readthedocs.io/en/v6.5.2/api/casashell/inp.html inp] reveals three parameters with defaults fitsfile=' '; vis=' '; antnamescheme='old'. | ||
* fitsfile=fitsdata sets the value of fitsfile to the fitsdata string variable defined earlier. If you were using your own dataset, you could enter it directly as a string (e.g., fitsfile='mydata.fits') or define it as a variable before setting fitsfile (e.g. fitsdata='mydata.fits', then fitsfile=fitsdata). You should avoid using 'fitsfile' as a variable name, as this will be overwritten by 'default importuvfits' or 'tget importuvfits'. | * fitsfile=fitsdata sets the value of fitsfile to the fitsdata string variable defined earlier. If you were using your own dataset, you could enter it directly as a string (e.g., fitsfile='mydata.fits') or define it as a variable before setting fitsfile (e.g. fitsdata='mydata.fits', then fitsfile=fitsdata). You should avoid using 'fitsfile' as a variable name, as this will be overwritten by 'default importuvfits' or 'tget importuvfits'. | ||
* The second [https://casadocs.readthedocs.io/en/ | * The second [https://casadocs.readthedocs.io/en/v6.5.2/api/casashell/inp.html inp] shows that the string variable fitsdata, defined earlier, has been expanded. A valid entry shows in blue, while an invalid entry shows in red. Default entries (as for antnamescheme in the initial listing of inputs) show in black. | ||
== Inspecting the data == | == Inspecting the data == | ||
The next step is to inspect to measurement set using the [https://casadocs.readthedocs.io/en/ | The next step is to inspect to measurement set using the [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.information.listobs.html listobs] task. This provides almost all relevant observational parameters – to the extent that the exist in the dataset – such as correlator setup (frequencies, bandwidths, channel number and widths, polarization products), sources, scans, scan intents, and antenna locations. Setting verbose=True (the default, so not included in the commands below) will display all of the contents of the raw data and setting listfile='listobs.txt' will create a text file you can refer to later (if this is not set, the output will instead display in the CASA Logger). Some of the parameters (such as scan intents) are not available for legacy VLA data such as this, so are left blank in the output. | ||
<pre style="background-color: #d8bfd8;"> | <pre style="background-color: #d8bfd8;"> | ||
| Line 168: | Line 171: | ||
=== Key information from listobs === | === Key information from listobs === | ||
The output of [https://casadocs.readthedocs.io/en/ | The output of [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.information.listobs.html listobs] is dense with information, but some facts to note are (working down the file): | ||
* The observations were taken on 13 April 1995 between 09:18:45 and 10:47:15 | * The observations were taken on 13 April 1995 between 09:18:45 and 10:47:15 | ||
| Line 236: | Line 239: | ||
===Antenna positions=== | ===Antenna positions=== | ||
[[file:plotants_6.5.2.png|thumb|Plotants output. Click to enlarge.]] | |||
The antenna positions are given in the output from listobs, but a graphical display can be easier to understand and aid in selecting a good reference antenna. This can be plotted using the [https://casadocs.readthedocs.io/en/ | The antenna positions are given in the output from listobs, but a graphical display can be easier to understand and aid in selecting a good reference antenna. This can be plotted using the [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.visualization.plotants.html plotants] task. | ||
<pre style="background-color: #d8bfd8;"> | <pre style="background-color: #d8bfd8;"> | ||
| Line 263: | Line 267: | ||
== Flagging == | == Flagging == | ||
The next step is to flag the data, for which we use the task [https://casadocs.readthedocs.io/en/ | The next step is to flag the data, for which we use the task [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.flagging.flagdata.html flagdata]. Flagging is a large topic, so for more details see the [[VLA CASA Flagging]] Topical CASA Guide. | ||
=== Flag the autocorrelations === | === Flag the autocorrelations === | ||
| Line 286: | Line 290: | ||
CASA issues a warning when running this command, as it is having to make assumptions about the data due to information not being available in legacy VLA files. This can be safely ignored. | CASA issues a warning when running this command, as it is having to make assumptions about the data due to information not being available in legacy VLA files. This can be safely ignored. | ||
If you run this in interactive mode, you will see that there are a lot of options for this task. We're going to use many of them later, but for now we just use vis to set the input measurement set and autocorr=True to say that we want to flag the autocorrelations. One parameter to check is flagbackup=True (which should be set by default). This backs up the flag state ''before'' carrying out the requested operation, allowing you to roll back to an earlier state using [https://casadocs.readthedocs.io/en/ | If you run this in interactive mode, you will see that there are a lot of options for this task. We're going to use many of them later, but for now we just use vis to set the input measurement set and autocorr=True to say that we want to flag the autocorrelations. One parameter to check is flagbackup=True (which should be set by default). This backs up the flag state ''before'' carrying out the requested operation, allowing you to roll back to an earlier state using [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.flagging.flagmanager.html flagmanager]. It is good practice when flagging to note which flagbackup version is associated with which flagging commands in your data reduction log. | ||
=== Quack flagging === | === Quack flagging === | ||
| Line 314: | Line 318: | ||
=== Interactive flagging === | === Interactive flagging === | ||
We use [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casaplotms.plotms.html plotms] to inspect the spectral line data. While plotms allows flagging inside the graphical interface, this does ''not'' write flagbackup files; we can use plotms to visualize the data in conjunction with using flagdata to actually set the flags in order to have flagbackup files. | |||
We use [https://casadocs.readthedocs.io/en/ | |||
To start plotms, you can simply run | To start plotms, you can simply run | ||
| Line 357: | Line 359: | ||
Alternatively, this can be changed in the plotms GUI. | Alternatively, this can be changed in the plotms GUI. | ||
The resulting plot shows that there is good signal on channels 6 to 56, which we can specify using spw='0:6~56'. There does not appear to be any visible RFI on this field; before moving on it's worth checking the other fields for RFI as well. This can be done from the GUI or using: | [[file:amp_vs_channel_6.5.2.png|thumb|Amplitude vs channel shown in plotms. Click to enlarge.]] | ||
The resulting plot (see figure) shows that there is good signal on channels 6 to 56, which we can specify using spw='0:6~56'. There does not appear to be any visible RFI on this field; before moving on it's worth checking the other fields for RFI as well. This can be done from the GUI or using: | |||
<pre style="background-color: #d8bfd8;"> | <pre style="background-color: #d8bfd8;"> | ||
| Line 385: | Line 389: | ||
</source> | </source> | ||
This plots amplitude against time. We can see from the colored striping on the sources that different baselines (coloraxis='baseline' means the different colors represent different baselines) have slightly different amplitudes, including one baseline (pink on the example plot) that is always high. | This plots amplitude against time (see figure). We can see from the colored striping on the sources that different baselines (coloraxis='baseline' means the different colors represent different baselines) have slightly different amplitudes, including one baseline (pink on the example plot) that is always high. | ||
[[file:amp_vs_time_6.5.2.png|thumb|Amplitude vs time plotted with plotms. Click to enlarge.]] | |||
We can draw a box around some of those pink points in plotms by clicking the '''mark regions''' [[Image:MarkRegionsButton.png]] tool to select that function and then drawing a box on the screen to mark a region. This may take a few seconds to return, but eventually the box will be filled with dots showing it has been selected. The points within the marked region can then be inspected using the '''locate''' [[File:casaplotms-locate-tool.png]] tool to the right of the region buttons. This outputs information on the selected points to the CASA logger. Once you are finished with a marked region, you should clear it with the '''clear regions''' [[File:Casaplotms-clear-region.png]] tool (attempting to change the plot while a marked region is present will often crash plotms). | We can draw a box around some of those pink points in plotms by clicking the '''mark regions''' [[Image:MarkRegionsButton.png]] tool to select that function and then drawing a box on the screen to mark a region. This may take a few seconds to return, but eventually the box will be filled with dots showing it has been selected. The points within the marked region can then be inspected using the '''locate''' [[File:casaplotms-locate-tool.png]] tool to the right of the region buttons. This outputs information on the selected points to the CASA logger. Once you are finished with a marked region, you should clear it with the '''clear regions''' [[File:Casaplotms-clear-region.png]] tool (attempting to change the plot while a marked region is present will often crash plotms). | ||
| Line 401: | Line 407: | ||
It can be seen that although the data is slightly high on the LL correlation of this baseline, it isn't particularly noisy. We have not yet calibrated this data, so some variation is possible. It does not look like this is a bad baseline/correlation, but rather that this will be dealt with by calibration. | It can be seen that although the data is slightly high on the LL correlation of this baseline, it isn't particularly noisy. We have not yet calibrated this data, so some variation is possible. It does not look like this is a bad baseline/correlation, but rather that this will be dealt with by calibration. | ||
<div style="padding-left: 50px; background-color: #F8F9FA;"> | |||
'''Flagging data:''' | |||
[[file:spectrum-flagging.png|thumb|Plotms settings for flagging spectral line data. Click to enlarge.]] | |||
If we did want to flag this data, we could either do so within plotms using the '''flag''' [[File:FlagThoseData.png]] tool or we could use flagdata again. If flagging baselines within plotms, you should ensure that you flag all of the channels; to do this, under the '''Flag''' tab, specify '''Extend flags''' to '''Channel''' (obviously, if flagging RFI channels you should not to this). You can read more in the tutorial on [[data flagging with plotms]]. However, if you want flag backups to be created (which is a good idea), you need to flag using flagdata rather than plotms. This can be done using: | If we did want to flag this data, we could either do so within plotms using the '''flag''' [[File:FlagThoseData.png]] tool or we could use flagdata again. If flagging baselines within plotms, you should ensure that you flag all of the channels; to do this, under the '''Flag''' tab, specify '''Extend flags''' to '''Channel''' (obviously, if flagging RFI channels you should not to this). You can read more in the tutorial on [[data flagging with plotms]]. However, if you want flag backups to be created (which is a good idea), you need to flag using flagdata rather than plotms. This can be done using: | ||
| Line 416: | Line 427: | ||
If executed, this would flag the left-hand correlation of the baseline between antennas VA12 and VA21. Other fields displayed by 'inp' in the section under "mode = 'manual'" can be used to set other data selection parameters, such as spw and timerange, so you can select the data to be flagged. If you control plotms from the GUI, you can identify data to be flagged using the '''mark regions''' and '''locate''' tools and then enter the appropriate parameters into flagdata to flag that data; when running like this you can use 'plotms' at the command line to restart it if it crashes, but while flagdata is the active task 'go' will execute flagdata rather than starting plotms. | If executed, this would flag the left-hand correlation of the baseline between antennas VA12 and VA21. Other fields displayed by 'inp' in the section under "mode = 'manual'" can be used to set other data selection parameters, such as spw and timerange, so you can select the data to be flagged. If you control plotms from the GUI, you can identify data to be flagged using the '''mark regions''' and '''locate''' tools and then enter the appropriate parameters into flagdata to flag that data; when running like this you can use 'plotms' at the command line to restart it if it crashes, but while flagdata is the active task 'go' will execute flagdata rather than starting plotms. | ||
</div> | |||
== Calibration == | == Calibration == | ||
Calibration of spectral line data broadly follows the approach for [[Calibrating a VLA 5 GHz continuum survey | continuum data]], except that the amplitude and phase corrections are a function of frequency and so must be corrected by '''bandpass calibration.''' The basic calibration steps follow. | Calibration of spectral line data broadly follows the approach for [[Calibrating a VLA 5 GHz continuum survey | continuum data]], except that the amplitude and phase corrections are a function of frequency and so must be corrected by '''bandpass calibration.''' The basic calibration steps follow. | ||
* [[#Setting the | * [[#Setting the flux scale | Set the flux scale]] of the primary calibrator, here, 1331+305 = 3C 286. | ||
* [[#A priori calibration| ''A priori'' calibration]] | * [[#A priori calibration| ''A priori'' calibration]] | ||
* [[#Bandpass calibration | Determine bandpass corrections]] based on the primary calibrator. In the script that follows, the bandpass calibration will be stored in '' | * [[#Bandpass calibration | Determine bandpass corrections]] based on the primary calibrator. In the script that follows, the bandpass calibration will be stored in ''bandpass.cal''. | ||
* [[#Inspect the | * [[#Inspect the bandpass response curve | Inspect the bandpass calibration]] for problems and to determine viable channels for averaging and imaging. We want to toss out end channels where the response is poor. | ||
* [[#Gain | * [[#Gain calibration | Determine the gain calibrations]] on the bandpass-corrected and channel-averaged data. In this step, we effectively turn the spectral line data into a single-channel continuum data set and calibrate accordingly. The calibration is first stored in ''apcal.gcal''. In the second part of this step we correct the fluxscale of the apcal table, and store the final calibration solutions with correct fluxes in ''fluxscale.cal'' (this is the table that needs to be applied later to the data, not the apcal.gcal version). | ||
* [[#Inspect the Calibration Solutions | Inspect the gain calibration solutions]] to look for any aberrant solutions that hint at bad calibrator data. | * [[#Inspect the Calibration Solutions | Inspect the gain calibration solutions]] to look for any aberrant solutions that hint at bad calibrator data. | ||
* [[#Apply the Solutions | Apply the calibration solutions]] to the source (N5921_2). This action literally adds a new column of data to the measurement set. This new column contains the data with the gain calibration and bandpass calibration applied, but it does not overwrite the raw data in case the calibration needs revision. | * [[#Apply the Solutions | Apply the calibration solutions]] to the source (N5921_2). This action literally adds a new column of data to the measurement set. This new column contains the data with the gain calibration and bandpass calibration applied, but it does not overwrite the raw data in case the calibration needs revision. | ||
=== Setting the | === Setting the flux scale === | ||
The [https://casadocs.readthedocs.io/en/ | The [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.imaging.setjy.html setjy] task generates a source model for the primary calibrator, 1331+305 = 3C286. From CASA 5.3 onwards, the default standard is Perley-Butler 2017 and includes resolved structure of the calibrators. This is 1.4GHz D-config and 1331+305 is sufficiently unresolved that, in principle, we don't need a model image; however, here we proceed with applying the detailed model, as a good practice. Setjy also looks up the radio SED for common flux calibrators and automatically assigns the total flux density. | ||
We can first get a listing of the available models. Setjy will look within the CASA working directory for files matching *.im* and *.mod*, i.e. user-defined images or models, and in the CalModels directory of the installation. Here we will want to apply a model from CalModels but, in principle, it is possible to use a user-defined model instead. | We can first get a listing of the available models. Setjy will look within the CASA working directory for files matching *.im* and *.mod*, i.e. user-defined images or models, and in the CalModels directory of the installation. Here we will want to apply a model from CalModels but, in principle, it is possible to use a user-defined model instead. | ||
| Line 511: | Line 523: | ||
A summary of the operation is sent to the logger window. Here's a listing of the output. | A summary of the operation is sent to the logger window. Here's a listing of the output. | ||
<pre> | <pre style="background-color: #fffacd;"> | ||
2022-12-02 00:32:41 INFO setjy ########################################## | 2022-12-02 00:32:41 INFO setjy ########################################## | ||
2022-12-02 00:32:41 INFO setjy ##### Begin Task: setjy ##### | 2022-12-02 00:32:41 INFO setjy ##### Begin Task: setjy ##### | ||
| Line 537: | Line 549: | ||
===''A priori'' calibration=== | ===''A priori'' calibration=== | ||
The [https://casadocs.readthedocs.io/en/ | The [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.calibration.gencal.html gencal] task carries out general calibrations using information that is known prior to inspecting the data, and thus known as ''a priori'' calibration. We can use it to apply the antenna gain curve and (if necessary) correct the antenna positions. | ||
'''Antenna gain-elevation curve calibration''' | '''Antenna gain-elevation curve calibration''' | ||
| Line 566: | Line 578: | ||
</pre> | </pre> | ||
<div style="padding-left: 50px; background-color: #F8F9FA;"> | |||
'''Antenna position corrections''' | '''Antenna position corrections''' | ||
Unlike the EVLA, where antenna position offsets can be looked up automatically and applied by gencal using caltype='antpos', you need to look up and apply these corrections manually for the legacy VLA. If needed, and if we have the necessary information, we can correct the antenna positions using gencal with caltype='antposvla'. This takes the offsets to be applied (dBx, dBy, dBz) in meters for a specified antenna. Please see the [https://casadocs.readthedocs.io/en/ | Unlike the EVLA, where antenna position offsets can be looked up automatically and applied by gencal using caltype='antpos', you need to look up and apply these corrections manually for the legacy VLA. If needed, and if we have the necessary information, we can correct the antenna positions using gencal with caltype='antposvla'. This takes the offsets to be applied (dBx, dBy, dBz) in meters for a specified antenna. Please see the [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.calibration.gencal.html gencal] documentation for further information. | ||
</div> | |||
=== Bandpass calibration === | === Bandpass calibration === | ||
| Line 583: | Line 598: | ||
caltable='bandpass.cal' | caltable='bandpass.cal' | ||
field='0' | field='0' | ||
solint='inf' | solint='inf' | ||
combine='scan' | combine='scan' | ||
refant='VA15' | refant='VA15' | ||
bandtype='B' | bandtype='B' | ||
gaintable='antgain.cal' | gaintable='antgain.cal' # ONLY IF YOU CREATED THIS WITH GENCAL EARLIER | ||
inp | |||
go | |||
</pre> | </pre> | ||
<source lang="python"> | <source lang="python"> | ||
# Pseudo-interactive CASA | # Pseudo-interactive CASA | ||
bandpass(vis='ngc5921.demo.ms', caltable='bandpass.cal', field='0' | bandpass(vis='ngc5921.demo.ms', caltable='bandpass.cal', field='0', solint='inf', combine='scan', refant='VA15', bandtype='B', gaintable='antgain.cal') | ||
</source> | |||
<pre style="background-color:lightgrey;"> | |||
Where: | |||
vis='ngc5921.demo.ms'. # Input measurement set | |||
caltable='bandpass.cal' # Output calibration table | |||
field='0' # Use the flux calibrator 1331+305 = 3C286 (FIELD_ID 0) as bandpass calibrator | |||
bandtype='B'. # Choose bandpass solution type. Pick standard time-binned B (rather than BPOLY) [default] | |||
solint='inf' # Set solution interval arbitrarily long (to get single bandpass) [default] | |||
combine='scan' # Combine scans (but break at obs, field and spw boundaries) [default] | |||
refant = 'VA15' # Set antenna VA15 (VLA:N2) as the reference antenna | |||
gaintable='antgain.cal' # Use the antgain.cal table created by gencal (only if you did this optional step) | |||
</pre> | |||
=== Inspect the bandpass response curve === | |||
In the [[#Gain Calibration | gain calibration]] to follow, we will effectively convert the spectral line data into a continuum data set. Before proceeding, we need to inspect the bandpass calibration to make sure that it contains no bad values and also to inspect which channels to average to produce the continuum data. | |||
We will plot the [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.calibration.bandpass.html bandpass] solutions as amplitude vs channel and phase vs channel per antenna. Start by looking at the amplitudes. These should be smooth across the channels. You will see the edge channels have lower amplitudes due to the edges of the bandpass being less sensitive. | |||
<pre style="background-color: #d8bfd8;"> | |||
# Interactive CASA | |||
default plotms | |||
inp | |||
vis='bandpass.cal' | |||
xaxis='chan' | |||
yaxis='amp' | |||
coloraxis='corr' | |||
iteraxis='antenna' | |||
gridrows=3 | |||
gridcols=3 | |||
xselfscale=True | |||
yselfscale=True | |||
inp | |||
go | |||
</pre> | |||
<source lang="python"> | |||
# Psuedo-interactive CASA | |||
plotms(vis='bandpass.cal',xaxis='chan',yaxis='amp',coloraxis='corr',iteraxis='antenna',gridrows=3,gridcols=3,xselfscale=True,yselfscale=True) | |||
</source> | </source> | ||
<pre style="background-color:lightgrey;"> | <pre style="background-color:lightgrey;"> | ||
Where: | Where: | ||
vis='bandpass.cal' # the bandpass calibration table | |||
xaxis='chan' # plot channels across the x axis | |||
yaxis='amp' # plot amplitudes across the y axis | |||
coloraxis='corr' # display the two correlations (RR & LL) in different colors | |||
iteraxis='antenna' # pages the plots by antenna | |||
gridrows=3 # places 3 antennas per row | |||
gridcols=3 # places 3 antennas per column | |||
xselfscale=True # create a global xaxis scale for all antennas | |||
yselfscale=True # create a global yaxis scale for all antennas | |||
</pre> | </pre> | ||
= | [[file:gainamp_vs_channel_6.5.2.png|thumb|Amplitude vs channel plotted in a 3×3 grid in plotms. Click to enlarge.]] | ||
This creates a 3×33 grid showing the amplitude on each antenna, with 9 antennas per page. You can scroll through the antennas using the green arrows at the bottom of the plotms GUI. Note that antenna VA23 in position VLA:OUT is displayed blank (it was not in the array at the time) and antenna VA28 is shown on a fourth page. This is not a problem. Looking at the amplitudes confirms our earlier observation that channels 6 to 56 (spw='0:6~56') look good. We will use this spw setting again in the later calibration stages. | |||
You may see the phases vs channel by changing the yaxis to phase interactively, or using the commands below. The phases should be relatively flat and continuous. Note that the phases do wander a bit at the edges of the bandpass where the amplitudes are lower. Some phases (e.g. on antenna VA10) wrap around between -180 and +180 – this is not a real discontinuity. VA15 has a very small variation in phases because it was used as the reference antenna. | |||
[[file:gainphase_vs_channel_6.5.2.png|thumb|Phase vs channel plotted in a 3×3 grid in plotms. Click to enlarge.]] | |||
<pre style="background-color: #d8bfd8;"> | |||
# Interactive CASA | |||
yaxis='phase' | |||
inp | |||
go | |||
</pre> | |||
Note, this follows on from having set the other parameters in interactive CASA as in the previous box, just changing the y-axis to phase. | |||
<source lang="python"> | |||
# Psuedo-interactive CASA | |||
plotms(vis='bandpass.cal',xaxis='chan',yaxis='phase',coloraxis='corr',iteraxis='antenna',gridrows=3,gridcols=3,xselfscale=True,yselfscale=True) | |||
</source> | |||
<!-- | |||
<source lang="python"> | <source lang="python"> | ||
# Set up 2x1 panels - upper panel amp vs. channel | # Set up 2x1 panels - upper panel amp vs. channel | ||
| Line 624: | Line 699: | ||
Notice that plotms is run twice: once to display gain amplitudes as a function of channel (frequency), and again to plot gain phases as a function of channel. | Notice that plotms is run twice: once to display gain amplitudes as a function of channel (frequency), and again to plot gain phases as a function of channel. | ||
--> | |||
<!-- | <!-- | ||
If the '''scriptmode''' is set to '''False''', the plot is saved to ngc5921.demo.plotcal.png. | If the '''scriptmode''' is set to '''False''', the plot is saved to ngc5921.demo.plotcal.png. | ||
| Line 651: | Line 726: | ||
--> | --> | ||
=== Gain | === Gain calibration === | ||
From [[#Inspect the | From our [[#Inspect the bandpass response curve | inspection of the bandpass response curve]], we can average channels 6 to 56 to produce continuum-like data for the calibrators. This averaging is specified through the '''spw''' (spectral window) parameter in the form spw = '0:6~56'. This specifies the spectral window (IF) to be used as '''0''' (there is only one spectral window in this measurement set, as seen in the listobs output) and channels '''6~56''' within that spectral window. | ||
Gain calibrations are otherwise determined as for [[Calibrating a VLA 5 GHz continuum survey#Calibration | continuum data]]. | |||
* [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.calibration.gaincal.html gaincal] is run only on the calibrators, 1331+305 (flux calibrator) and 1445+099 (phase calibrator). | |||
* The default model for gain calibrations is a 1 Jy point source. The flux scale is overridden by [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.imaging.setjy.html setjy], which has been performed for the flux calibrator. We need to transfer that flux scale to the phase calibrator using [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.calibration.fluxscale.html fluxscale]. | |||
* Note that fluxscale determines the flux density of the phase calibrator and accordingly adjusts its model and calibration solutions. A report of the results are sent to the logger window. | |||
* Unless you use parameter ''incremental=True'' while executing fluxscale (the default is ''False''), the resulting fluxscale.cal table will replace the apcal.gcal table at this point. This is particularly important when it comes to applying the calibration tables to the data in the [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.calibration.applycal.html applycal] stage. | |||
'''Complex gain calibration''' | |||
First, we will calibrate the amplitude and phases of the data using the flux/bandpass and complex gain calibrators. To do this we will use [https://casadocs.readthedocs.io/en/v6.2.0/api/tt/casatasks.calibration.gaincal.html gaincal] and the previously-made calibration tables. This will give us antenna-based solutions that will show variations across time. | |||
<pre style="background-color: #d8bfd8;"> | |||
# Interactive CASA | |||
default gaincal | |||
inp | |||
vis='ngc5921.demo.ms' | |||
caltable='apcal.gcal' | |||
field='0,1' | |||
spw='0:6~56' | |||
refant='VA15' | |||
calmode='ap' | |||
solint='inf' | |||
gaintable=['antgain.cal','bandpass.cal'] | |||
inp | |||
go | |||
</pre> | |||
<source lang="python"> | <source lang="python"> | ||
spw = '0:6~56' | # Psuedo-interactive CASA | ||
gaincal(vis='TDEM0025_HI.ms',caltable='apcal.gcal',field='0,1',spw='0:6~56',refant='VA15',calmode='ap',solint='inf',gaintable=['antgain.cal','bandpass.cal']) | |||
</source> | </source> | ||
<pre style="background-color:lightgrey;"> | |||
Where: | |||
vis='ngc5921.demo.ms' # the measurement set | |||
caltable='apcal.gcal' # the name you wish to give the calibration table | |||
field='0,1' # the field ids of both calibrators | |||
refant='VA15' # the reference antenna | |||
calmode='ap' # to find both amplitude and phase solutions | |||
solint='inf' # the solution interval time | |||
gaintable=['antgain.cal','bandpass.cal'] # existing gain calibration table(s) to apply on the fly | |||
</pre> | |||
Now look at solutions. First we will look at the phase solutions. These should be at zero with no phase jumps; phase jumps are when the phases change drastically with time. When this happens we are not able to interpolate the phases surrounding that scan that has the phase jump. | |||
[[File:cg_phase_before_casa540.png|200px|thumb|right|Plot of complex gain phase vs time from plotms. Click to enlarge.]] | |||
<pre style="background-color: #d8bfd8;"> | |||
# Interactive CASA | |||
default plotms | |||
inp | |||
vis='apcal.gcal' | |||
xaxis='time' | |||
yaxis='phase' | |||
iteraxis='antenna' | |||
coloraxis='corr' | |||
gridrows=3 | |||
gridcols=3 | |||
plotrange=[0,0,-180,180] | |||
inp | |||
go | |||
</pre> | |||
<source lang="python"> | <source lang="python"> | ||
# | # Psuedo-interactive CASA | ||
plotms(vis='apcal.gcal',xaxis='time',yaxis='phase',iteraxis='antenna',gridrows=3,gridcols=3,plotrange=[0,0,-180,180]) | |||
</source> | </source> | ||
< | <pre style="background-color:lightgrey;"> | ||
Where: | |||
vis='apcal.gcal' # the complex gain calibration table | |||
xaxis='time' # plot channels across the x axis | |||
yaxis='phase' # plot amplitudes across the y axis | |||
iteraxis='antenna' # pages the plots by antenna | |||
coloraxis='corr' # colorizes the plots by correlation (RR and LL) | |||
gridrows=3 # places 3 antennas per row | |||
gridcols=3 # places 3 antennas per column | |||
plotrange=[0,0,-180,180] # sets the plot range | |||
</pre> | |||
Now look at the amplitudes to see if there are any irregularities there. | |||
<pre style="background-color: #d8bfd8;"> | |||
# Interactive CASA | |||
default plotms | |||
inp | |||
vis='apcal.gcal' | |||
xaxis='time' | |||
yaxis='amp' | |||
iteraxis='antenna' | |||
coloraxis='corr' | |||
gridrows=3 | |||
gridcols=3 | |||
xselfscale=True | |||
yselfscale=True | |||
inp | |||
go | |||
</pre> | |||
<source lang="python"> | <source lang="python"> | ||
# | # Psuedo-interactive CASA | ||
plotms(vis='apcal.gcal',xaxis='time',yaxis='amp',iteraxis='antenna',gridrows=3,gridcols=3,xselfscale=True,yselfscale=True) | |||
</source> | </source> | ||
< | <pre style="background-color:lightgrey;"> | ||
Where: | |||
vis='apcal.gcal' # the complex gain calibration table | |||
xaxis='time' # plot channels across the x axis | |||
yaxis='amp' # plot amplitudes across the y axis | |||
iteraxis='antenna' # pages the plots by antenna | |||
coloraxis='corr' # colorizes the plots by correlation (RR and LL) | |||
gridrows=3 # places 3 antennas per row | |||
gridcols=3 # places 3 antennas per column | |||
xselfscale=True # create a global xaxis scale for all antennas | |||
yselfscale=True # create a global yaxis scale for all antennas | |||
</pre> | |||
Don't worry about the shift in amplitude between the flux calibrator and the phase calibrator; this is because we haven't yet transferred the flux scale to the phase calibrator. What we're looking for here is whether there are large variations on the individual calibrators. | |||
Next we use the gain calibration derived above and the known flux of the flux calibrator to transfer the flux scale to the phase calibrator using the fluxscale task. | |||
<pre style="background-color: #d8bfd8;"> | |||
# Interactive CASA | |||
default fluxscale | |||
inp | |||
vis='ngc5921.demo.ms' | |||
caltable='apcal.gcal' | |||
reference='0' | |||
fluxtable='fluxscale.cal' | |||
incremental=False | |||
inp | |||
go | |||
</pre> | |||
<source lang="python"> | <source lang="python"> | ||
# Psuedo-interactive CASA | |||
vis = | fluxscale(vis='ngc5921.demo.ms',caltable='apcal.gcal',reference='0',fluxtable='fluxscale.cal',incremental=False) | ||
caltable | |||
</source> | </source> | ||
<pre | <pre style="background-color:lightgrey;"> | ||
Where: | |||
vis='ngc5921.demo.ms' # the measurement set | |||
caltable='apcal.gcal' # the calibration table with the amplitude and phase corrections | |||
reference='0' # the field ID of the reference field (i.e. the flux calibrator) | |||
fluxtable='fluxscale.cal' # the name of the output, flux-scaled calibration table | |||
incremental=False # create a single fluxscale table with all the gain calibration | |||
# ('False' is the default; if you use 'True' you need to supply | |||
# both the apcal.gcal and fluxscale.cal tables to applycal below) | |||
</pre> | </pre> | ||
< | The output from [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.calibration.fluxscale.html fluxscale] follows. A relatively large uncertainty for the phase calibrator is a sign that something went wrong, such as having bad solutions in [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.calibration.gaincal.html gaincal]. Here, the phase calibrator scaled to 2.521 ± 0.002 Jy, which looks reasonable. | ||
<pre style="background-color: #fffacd;"> | |||
2023-01-17 22:47:15 INFO fluxscale ########################################## | |||
2023-01-17 22:47:15 INFO fluxscale ##### Begin Task: fluxscale ##### | |||
2023-01-17 22:47:15 INFO fluxscale fluxscale( vis='ngc5921.demo.ms', caltable='apcal.gcal', fluxtable='fluxscale.cal', reference=['0'], transfer=[], listfile='', append=False, refspwmap=[-1], gainthreshold=-1.0, antenna='', timerange='', scan='', incremental=False, fitorder=1, display=False ) | |||
2023-01-17 22:47:15 INFO fluxscale ****Using NEW VI2-driven calibrater tool**** | |||
2023-01-17 22:47:15 INFO fluxscale Opening MS: ngc5921.demo.ms for calibration. | |||
2023-01-17 22:47:15 INFO fluxscale Initializing nominal selection to the whole MS. | |||
2023-01-17 22:47:15 INFO fluxscale Beginning fluxscale--(MSSelection version)------- | |||
2023-01-17 22:47:16 INFO fluxscale Assuming all non-reference fields are transfer fields. | |||
2023-01-17 22:47:16 INFO fluxscale Found reference field(s): 1331+30500002_0 | |||
2023-01-17 22:47:16 INFO fluxscale Found transfer field(s): 1445+09900002_0 | |||
2023-01-17 22:47:16 INFO fluxscale Flux density for 1445+09900002_0 in SpW=0 (freq=1.41342e+09 Hz) is: 2.52141 +/- 0.00223811 (SNR = 1126.58, N = 54) | |||
2023-01-17 22:47:16 INFO fluxscale Storing result in fluxscale.cal | |||
2023-01-17 22:47:16 INFO fluxscale Writing solutions to table: fluxscale.cal | |||
2023-01-17 22:47:16 INFO fluxscale Task fluxscale complete. Start time: 2023-01-17 15:47:15.041544 End time: 2023-01-17 15:47:16.120049 | |||
2023-01-17 22:47:16 INFO fluxscale ##### End Task: fluxscale ##### | |||
2023-01-17 22:47:16 INFO fluxscale ########################################## | |||
</pre> | </pre> | ||
Now we can look at the amplitudes again, using the transferred flux scale which should eliminate the differences between the calibrators. This is identical to the previous inspection, but using fluxscale.cal rather than apcal.gcal (so could be run in interactive CASA by retrieving the previous inputs with "tget plotms" then just changing the input table using "vis='fluxscale.cal'"). | |||
<pre style="background-color: #d8bfd8;"> | |||
# Interactive CASA | |||
default plotms | |||
inp | |||
vis='fluxscale.cal' | |||
xaxis='time' | |||
yaxis='amp' | |||
iteraxis='antenna' | |||
coloraxis='corr' | |||
gridrows=3 | |||
gridcols=3 | |||
xselfscale=True | |||
yselfscale=True | |||
inp | |||
go | |||
</pre> | |||
<source lang="python"> | |||
# Psuedo-interactive CASA | |||
plotms(vis='fluxscale.cal',xaxis='time',yaxis='amp',iteraxis='antenna',gridrows=3,gridcols=3,xselfscale=True,yselfscale=True) | |||
plotms(vis=' | |||
</source> | </source> | ||
<pre style="background-color:lightgrey;"> | |||
Where: | |||
vis='fluxscale.cal' # the complex gain calibration table with the flux scale applied | |||
xaxis='time' # plot channels across the x axis | |||
yaxis='amp' # plot amplitudes across the y axis | |||
iteraxis='antenna' # pages the plots by antenna | |||
coloraxis='corr' # colorizes the plots by correlation (RR and LL) | |||
gridrows=3 # places 3 antennas per row | |||
gridcols=3 # places 3 antennas per column | |||
xselfscale=True # create a global xaxis scale for all antennas | |||
yselfscale=True # create a global yaxis scale for all antennas | |||
</pre> | |||
=== Apply the solutions === | |||
Next, we apply the calibration solutions to the calibrators themselves, and finally transfer the calibration solutions by interpolation (or nearest-neighbor sampling) to the source. The relevant task is [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.calibration.applycal.html applycal], which fills out a new column (CORRECTED_DATA) of calibrated data in the measurement set without replacing the raw data column (DATA). The application is identical to that used for [[Calibrating a VLA 5 GHz continuum survey|continuum data]], except that the bandpass table is also included in the calibration. In order to apply multiple calibrations at once, we provide the '''gaintable''' parameter with a list of calibration tables rather than a single entry. If we had run fluxscale with "incremental=True", we would also include the apcal.gcal table here. | |||
First, apply the tables to the flux calibrator: | |||
<pre style="background-color: #d8bfd8;"> | |||
# Interactive CASA | |||
default applycal | |||
inp | |||
vis='ngc5921.demo.ms' | |||
field='0' | |||
gaintable=['antgain.cal', 'bandpass.cal', 'fluxscale.cal'] | |||
gainfield=['','0','0'] | |||
calwt=False | |||
flagbackup=True | |||
inp | |||
go | |||
</pre> | |||
<source lang="python"> | <source lang="python"> | ||
# | # Psuedo-interactive CASA | ||
# for the flux/bandpass calibrator (field '0') | |||
applycal(vis='ngc5921.demo.ms', field='0', | |||
gaintable=['antgain.cal', 'bandpass.cal', 'fluxscale.cal'], | |||
gainfield=['','0','0'], | |||
calwt=False, flagbackup=True) | |||
</source> | </source> | ||
<pre style="background-color:lightgrey;"> | |||
Where: | |||
vis='ngc5921.demo.ms' # the measurement set | |||
field='0' # field id(s) to work on - 0 = the flux calibrator | |||
gaintable=['antgain.cal''bandpass.cal','fluxscale.cal'] # gain calibration table(s) to apply | |||
gainfield=['','0','0'] # field id(s) from which each respective gaintable was derived | |||
calwt=False # determines if the weights are calibrated along with the data | |||
flagbackup=True # automatically back up the state of flags before the run | |||
</pre> | |||
Within the gainfield parameter, we do not specify a field for the ''antgain.cal'' table since this was derived independently from the calibrator fields. For the calibrators, all bandpass solutions are the same as for the flux/bandpass calibrator (id=0), and the phase and amplitude calibration as derived from their own visibility data. So when applying corrections to any of our sources, the respective field for ''bandpass.cal'' will be our bandpass calibrator (field='0'), but the field for ''fluxscale.cal'' will be '1' for both the phase calibrator itself and the target field. | |||
field | |||
Next, apply the tables to the phase calibrator: | |||
=== | <pre style="background-color: #d8bfd8;"> | ||
# Interactive CASA | |||
default applycal | |||
inp | |||
vis='ngc5921.demo.ms' | |||
field='1' | |||
gaintable=['antgain.cal', 'bandpass.cal', 'fluxscale.cal'] | |||
gainfield=['','0','1'] | |||
calwt=False | |||
flagbackup=True | |||
inp | |||
go | |||
</pre> | |||
(Note: the only changes from the application to the flux calibrator are the field to apply the calibration to and the gainfield for the fluxscale.cal table, both changed to refer to the phase calibrator.) | |||
<source lang="python"> | <source lang="python"> | ||
# Psuedo-interactive CASA | |||
# for the phase calibrator (field '1') | |||
applycal(vis='ngc5921.demo.ms', field='1', | |||
gaintable=['antgain.cal', 'bandpass.cal', 'fluxscale.cal'], | |||
gainfield=['','0','1'], | |||
calwt=False, flagbackup=True) | |||
</source> | </source> | ||
<pre style="background-color:lightgrey;"> | |||
Where: | |||
vis='ngc5921.demo.ms' # the measurement set | |||
field='1' # field id(s) to work on - 1 = the flux calibrator | |||
gaintable=['antgain.cal''bandpass.cal','fluxscale.cal'] # gain calibration table(s) to apply | |||
gainfield=['','0','1'] # field id(s) from which each respective gaintable was derived | |||
calwt=False # determines if the weights are calibrated along with the data | |||
flagbackup=True # automatically back up the state of flags before the run | |||
</pre> | |||
Finally, apply the tables to the science target: | |||
< | <pre style="background-color: #d8bfd8;"> | ||
applycal | # Interactive CASA | ||
default applycal | |||
</ | inp | ||
vis='ngc5921.demo.ms' | |||
field='2' | |||
gaintable=['antgain.cal', 'bandpass.cal', 'fluxscale.cal'] | |||
gainfield=['','0','1'] | |||
calwt=False | |||
flagbackup=True | |||
inp | |||
go | |||
</pre> | |||
(Note: the only change from the phase calibrator is the field to apply the calibration to – the gain fields are all the same.) | |||
<source lang="python"> | <source lang="python"> | ||
applycal(vis='ngc5921.demo.ms', field=' | # Psuedo-interactive CASA | ||
# for the science target (field '2') | |||
applycal(vis='ngc5921.demo.ms', field='2', | |||
gaintable=['antgain.cal', 'bandpass.cal', 'fluxscale.cal'], | |||
gainfield=['','0','1'], | |||
calwt=False, flagbackup=True) | |||
</source> | </source> | ||
< | <pre style="background-color:lightgrey;"> | ||
Where: | |||
vis='ngc5921.demo.ms' # the measurement set | |||
field='2' # field id(s) to work on - 2 = the science target | |||
gaintable=['antgain.cal''bandpass.cal','fluxscale.cal'] # gain calibration table(s) to apply | |||
gainfield=['','0','1'] # field id(s) from which each respective gaintable was derived | |||
calwt=False # determines if the weights are calibrated along with the data | |||
flagbackup=True # automatically back up the state of flags before the run | |||
</pre> | |||
== Plot the spectrum == | |||
[[File:spectrum_plot_colors_6.5.2.png|thumb|Integrated spectrum from the calibrated visibilities plotted with the two polarizations shown in different colors. This shows which channels contain an HI signal and which are good to use for continuum subtraction. Click to enlarge]] | |||
Before we attempt to image the 21 cm cube of the source, we need to subtract off the underlying continuum, which means we need to plot the integrated spectrum of the source to determine the line-free channels where we can fit the continuum. | |||
[ | We can do this in [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casaplotms.plotms.html plotms]. | ||
<pre style="background-color: #d8bfd8;"> | |||
# Interactive CASA | |||
default plotms | |||
inp | |||
vis='ngc5921.demo.ms' | |||
field='2' | |||
spw='0:6~56' | |||
averagedata=True | |||
avgtime='3600' | |||
avgscan=True | |||
avgbaseline=True | |||
xaxis='channel' | |||
yaxis='amp' | |||
coloraxis='corr' | |||
ydatacolumn='corrected' | |||
inp | |||
go | |||
</pre> | |||
<source lang="python"> | <source lang="python"> | ||
plotms(vis='ngc5921.demo.ms' | # Psuedo-interactive CASA | ||
plotms(vis='ngc5921.demo.ms', field='2', spw='0:6~56', averagedata=True, avgtime='3600', avgscan=True, avgbaseline=True, xaxis='channel', yaxis='amp', ydatacolumn='corrected') | |||
</source> | </source> | ||
<pre style="background-color:lightgrey;"> | |||
Where: | |||
vis='ngc5921.demo.ms' # the measurement set | |||
field='2' # select the science target | |||
averagedata=True # average the data (enables data-averaging options) | |||
avgtime='3600' # average over 3600s (i.e. the whole dataset) | |||
avgscan=True # average across scan boundaries | |||
avgbaseline=True # average baselines | |||
xaxis='channel' # plot channels across the x axis | |||
yaxis='amp' # plot amplitudes across the y axis | |||
coloraxis='corr' # colorizes the plots by correlation (RR and LL) | |||
ydatacolumn='corrected' # use the CORRECTED_DATA column (with the calibration applied) | |||
</pre> | |||
The resulting plot is illustrated in the figure at right. Briefly, we want to average both in time and over baselines to get the signal-to-noise necessary to reveal the 21 cm profile (see [[Averaging data in plotms]] for more details on averaging options). If the symbols appear too small, the size may be increased via the Display tab on the GUI: change the Unflagged Points Symbol to 'Custom', change the plotting symbol to 'circle', 'square' or 'diamond', and increase the number of pixels for the plotting symbol. | |||
Note that we have entered all the relevent parameters via the command line interface rather than selecting options in the GUI. If you wish to enter the values directly into the GUI, you can follow the (Tab)'''Command''' convention of the [[Data flagging with casaplotms|flagging tutorial]] with the following settings : | |||
* (Data)'''field''' = N5921* | * (Data)'''field''' = N5921* | ||
* (Data)'''Averaging → Time''' = 3600 (average over some long time window) | * (Data)'''Averaging → Time''' = 3600 (average over some long time window) | ||
* (Data)'''Averaging → Scan''' = True (checkmark; average in time across scan boundaries) | * (Data)'''Averaging → Scan''' = True (checkmark; average in time across scan boundaries) | ||
| Line 907: | Line 1,113: | ||
* (Axes)'''X Axis''' = Channel | * (Axes)'''X Axis''' = Channel | ||
* (Axes)'''Y Axis''' = Amp | * (Axes)'''Y Axis''' = Amp | ||
* (Display)'''Colorize''' = Corr | |||
From inspection of this plot, it looks like channels 4~8 and 50~59 contain line-free channels, suitable to use for continuum subtraction. | |||
== Continuum | == Continuum subtraction == | ||
The next step is to split off the NGC 5921 data from the multisource measurement set and subtract the continuum. Splitting uses the | The next step is to split off the NGC 5921 data from the multisource measurement set and subtract the continuum. Splitting uses the [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.manipulation.split.html split] command, as follows. | ||
<pre style="background-color: #d8bfd8;"> | |||
# Interactive CASA | |||
default split | |||
inp | |||
vis='ngc5921.demo.ms' | |||
field='2' | |||
outputvis='ngc5921.demo.split.ms' | |||
spw='' | |||
datacolumn='corrected' | |||
inp | |||
go | |||
</pre> | |||
<source lang="python"> | <source lang="python"> | ||
# Psuedo-interactive CASA | |||
plotms(vis='ngc5921.demo.ms', outputvis='ngc5921.demo.split.ms', field='2', spw='', datacolumn='corrected') | |||
</source> | </source> | ||
< | <pre style="background-color:lightgrey;"> | ||
Where: | |||
vis = | vis='ngc5921.demo.ms' # the input measurement set | ||
outputvis='ngc5921.demo.split.ms # the output split measurement set | |||
field='2' # select the science target | |||
field = ' | spw='' # select the entire spectral window | ||
spw = '' | datacolumn='corrected' # use the CORRECTED_DATA column (with the calibration applied) | ||
datacolumn = 'corrected' | </pre> | ||
This creates a new measurement set called ''ngc5921.demo.split.ms'' and copies the ''calibrated'' source data (datacolumn = 'corrected') from the CORRECTED_DATA column into it, writing it to the DATA column in the output measurement set. | |||
We can now subtract the continuum from this split dataset in the visibility (''uv'') plane using [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.manipulation.uvcontsub.html uvcontsub]. As we determined above, we can use channels 4~8 and 50~59 for this. Note that the uvcontsub task was substantially revised in CASA 6.5 and now has new parameter inputs that are not backwards compatible with inputs based on older CASAguides. | |||
<pre style="background-color: #d8bfd8;"> | |||
# Interactive CASA | |||
default uvcontsub | |||
ins | |||
vis='ngc5921.demo.split.ms' | |||
outputvis='ngc5921.demo.uvcontsub.ms' | |||
fitspec='0:4~8;50~59' | |||
fitorder=0 | |||
inp | |||
go | |||
</pre> | |||
<source lang="python"> | <source lang="python"> | ||
# Psuedo-interactive CASA | |||
plotms(vis='ngc5921.demo.ms', outputvis='ngc5921.demo.split.ms', fitspec='0:4~8;50~59', fitorder=0) | |||
</source> | </source> | ||
< | <pre style="background-color:lightgrey;"> | ||
Where: | |||
vis = | vis='ngc5921.demo.split ms' # the input split measurement set | ||
outputvis='ngc5921.demo.uvcontsub.ms # the output continuum-subtracted measurement set | |||
# | fitspec='0:4~8;50~59' # select the channels over which to fit the continuum | ||
fitorder=0 # fit with polynomial order 0 (constant) | |||
# | </pre> | ||
# | |||
</ | |||
(Note: this could be run directly on the original measurement set by specifying vis='ngc5921.demo.ms', field='2', and datacolumn='corrected' to select the science target and the | |||
calibrated data column – in a change from its earlier behavior, datacolumn='data' is the default rather than the fallback for uvcontsub, so this must be specified explicitly if not using split data) | |||
The fitspec parameter, if given as a simple string as done here, specifies the spw to be used for the fitting, with the fit order given using fitorder. However, fitspect can be used in more complex ways by specifying it as a python dictionary. For details, see the uvcontsub documentation. | |||
The polynomial fit order specified is used to fit the real and imaginary components separately, so the fitted model amplitude is the quadrature combination of these two fits and is thus not, in general, a polynomial. | |||
== Imaging == | == Imaging == | ||
[[File:spectrum-uvplot.png|thumb|Plot of amplitude vs | [[File:spectrum-uvplot.png|thumb|Plot of amplitude vs projected baseline length (in units of the observing wavelength) produced by plotms. The maximum baseline is just below 5 kλ. Click to enlarge.]] | ||
Now we can generate the primary science product, a | Now we can use [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.imaging.tclean.html tclean] to generate the primary science product, a cleaned data cube (ra, dec, velocity) from the continuum-subtracted (''u'', ''v'', channel) measurement set, ''ngc5921.demo.uvcontsub.ms''. Things to consider in using tclean: | ||
* To ensure channels aren't averaged prior to imaging, choose ''' | * To ensure channels aren't averaged prior to imaging and proper Doppler combination, choose '''specmode='cube''''. | ||
* The maximum baseline is just under 5 kλ (see the figure at right, showing this plotted with plotms), and so the expected synthetic beam is roughly 1.22 × 206265 / 5000 = 50 arcseconds (subject to the details of ''u'', ''v'' weighting), as expected for D configuration. Pixels should sample the beam better than 3 times, so 15 arcseconds is a good choice of pixel size ('''cell = ['15.0arcsec','15.0arcsec']'''). | |||
* The maximum baseline is just under 5 | * We only want to clean down to the noise, which can be determined by examining line-free channels in an image made without cleaning (a 'dirty' image), as detailed below. | ||
* We only want to | |||
[[file:CARTA_dirty_cube_6.5.2.png|thumb|A channel near the center of the source in the dirty cube in CARTA, compare to the same channel in the tcleaned cube [[#The integrated spectrum|below]]. Click to enlarge.]] | |||
To create a dirty image: | |||
< | <pre style="background-color: #d8bfd8;"> | ||
# | # Interactive CASA | ||
tclean | default tclean | ||
inp | |||
vis='ngc5921.demo.uvcontsub.ms' | |||
specmode=cube | |||
inp #Run inp again, as setting specmode=cube opens new options | |||
cell=['15.0arcsec','15.0arcsec'] | |||
outframe='bary' | |||
niter = 0 | |||
imagename='ngc5921.demo.dirty' | |||
datacolumn='data' | |||
imsize=[256,256] | |||
pblimit=-0.2 | |||
weighting='briggs' | |||
robust=0.5 | |||
inp | |||
go | |||
</pre> | |||
imsize = [256,256] | |||
weighting = 'briggs' | |||
robust = 0.5 | |||
<source lang="python"> | <source lang="python"> | ||
# Pseudo-interactive CASA | |||
tclean(vis='ngc5921.demo.uvcontsub.ms', imagename='ngc5921.demo.dirty', specmode='cube', niter=0, imsize=[256,256], cell=['15.0arcsec','15.0arcsec'], outframe='baby', weighting='briggs', robust=0.5, interactive=False, pblimit=-0.2) | |||
</source> | </source> | ||
< | <pre style="background-color:lightgrey;"> | ||
Where: | |||
vis='ngc5921.demo.uvcontsub.ms'. # the input measurement set | |||
imagename = | specmode=cube # sets the spectral mode to properly handle line data | ||
cell=['15.0arcsec','15.0arcsec'] # sets the cell (pixel) size | |||
outframe='bary' # sets the velocity frame to the extragalactic standard of barycentric | |||
- | niter = 0 # sets the number of clean cycles to 0 (i.e. no cleaning) | ||
imagename='ngc5921.demo.dirty' # the output image name | |||
datacolumn='data' # tells tclean to use the DATA column | |||
imsize=[256,256] # sets the image size (in pixels) | |||
pblimit=-0.2 # negative value to not blank pixels beyond a primary beam gain of 20% | |||
weighting='briggs' # use Briggs weighting... | |||
robust=0.5 # ...with a robust value of 0.5 | |||
</pre> | |||
The output, | The output ''ngc5921.demo.dirty.image'' file (note that .image gets appended – other outputs such as the .psf, .pb, etc. are also created by tclean) can then be viewed using [https://cartavis.org CARTA]. By defining a rectangular region that takes in the whole image plane and setting the Z profile to 'RMS', it is possible to see how the RMS level varies by channel. You can see how the noise falls from the edge of the bandpass to around 1.75 mJy by channel 6, then rises again from channel 9 when the source starts to contribute, then falls back to 1.75 mJy again by channel 50, rising again from channel 56. This also confirms our earlier choices of which channels looked line-free and which channels were in the best signal-to-noise section of the cube. If we thus want to clean to 4 sigma, we can set '''threshold='7.0mJy''''. | ||
The dirty image can also be used to determine the region to be cleaned by tclean. This is most easily done by defining a region that encloses the area where flux is visible on the dirty image (which can be checked by stepping through the channels) and then exporting this region as a CRTF file, here ''cleanbox.crtf'', that can be read by CASA. | |||
We are now ready to run tclean again and create the clean cube. | |||
<pre style="background-color: #d8bfd8;"> | |||
<pre | # Interactive CASA | ||
default tclean | |||
inp | |||
vis='ngc5921.demo.uvcontsub.ms' | |||
specmode=cube | |||
cell=['15.0arcsec','15.0arcsec'] | |||
outframe='bary' | |||
datacolumn='data' | |||
imsize=[256,256] | |||
pblimit=-0.2 | |||
weighting='briggs' | |||
robust=0.5 | |||
imagename='ngc5921.demo.clean' | |||
niter=6000 | |||
mask='cleanbox.crtf' | |||
threshold='7.0mJy' | |||
inp | |||
go | |||
</pre> | </pre> | ||
(Note: only the last four parameters, imagename, niter, mask and threshold have changed) | |||
< | <source lang="python"> | ||
# Pseudo-interactive CASA | |||
tclean(vis='ngc5921.demo.uvcontsub.ms', imagename='ngc5921.demo.clean', specmode='cube', niter=6000, threshold='7.0mJy', imsize=[256,256], cell=['15.0arcsec','15.0arcsec'], outframe='baby', weighting='briggs', robust=0.5, mask = 'clean box.crtf', interactive=False, pblimit=-0.2) | |||
</source> | |||
</ | |||
<pre style="background-color:lightgrey;"> | |||
Where: | |||
vis='ngc5921.demo.uvcontsub.ms'. # the input measurement set | |||
specmode=cube # sets the spectral mode to properly handle line data | |||
cell=['15.0arcsec','15.0arcsec'] # sets the cell (pixel) size | |||
outframe='bary' # sets the velocity frame to the extragalactic standard of barycentric | |||
datacolumn='data' # tells tclean to use the DATA column | |||
imsize=[256,256] # sets the image size (in pixels) | |||
pblimit=-0.2 # negative value to not blank pixels beyond a primary beam gain of 20% | |||
weighting='briggs' # use Briggs weighting... | |||
robust=0.5 # ...with a robust value of 0.5 | |||
imagename='ngc5921.demo.clean' # the output image name | |||
niter=6000 # sets the number of clean cycles to 6000 (will stop at threshold value) | |||
mask='cleanbox.crtf' # use the mask defined in the cleanbox.crtf file created by CARTA | |||
threshold='7.0mJy' # stop cleaning at this flux level | |||
</pre> | |||
== | ==Inspecting the data cube== | ||
We can use [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.information.imhead.html imhead] to examine the cube header: | |||
== | <pre style="background-color: #d8bfd8;"> | ||
# Interactive CASA | |||
default imhead | |||
inp | |||
imagename='ngc5921.demo.clean.image' | |||
inp | |||
go | |||
</pre> | |||
<source lang="python"> | <source lang="python"> | ||
# Pseudo-interactive CASA | |||
imhead(imagename='ngc5921.demo.clean.image', mode='summary') | |||
</source> | </source> | ||
The output, as follows, appears in the logger window: | |||
<pre style="background-color: #fffacd;"> | |||
2023-01-18 21:15:06 INFO imhead ########################################## | |||
2023-01-18 21:15:06 INFO imhead ##### Begin Task: imhead ##### | |||
2023-01-18 21:15:06 INFO imhead imhead( imagename='ngc5921.demo.clean.image', mode='summary', hdkey='', hdvalue='', verbose=False ) | |||
2023-01-18 21:15:08 INFO ImageMetaData | |||
2023-01-18 21:15:08 INFO ImageMetaData Image name : ngc5921.demo.clean.image | |||
2023-01-18 21:15:08 INFO ImageMetaData Object name : N5921_2 | |||
2023-01-18 21:15:08 INFO ImageMetaData Image type : PagedImage | |||
2023-01-18 21:15:08 INFO ImageMetaData Image quantity : Intensity | |||
2023-01-18 21:15:08 INFO ImageMetaData Pixel mask(s) : None | |||
2023-01-18 21:15:08 INFO ImageMetaData Region(s) : None | |||
2023-01-18 21:15:08 INFO ImageMetaData Image units : Jy/beam | |||
2023-01-18 21:15:08 INFO ImageMetaData Restoring Beams | |||
2023-01-18 21:15:08 INFO ImageMetaData Pol Type Chan Freq Vel | |||
2023-01-18 21:15:08 INFO ImageMetaData I Max 0 1.41273e+09 1619.82 53.0827 arcsec x 47.0911 arcsec pa= 7.9378 deg | |||
2023-01-18 21:15:08 INFO ImageMetaData I Min 48 1.41390e+09 1372.47 53.0292 arcsec x 46.9723 arcsec pa= 9.2164 deg | |||
2023-01-18 21:15:08 INFO ImageMetaData I Median 31 1.41349e+09 1460.08 53.1100 arcsec x 47.0231 arcsec pa= 8.0658 deg | |||
2023-01-18 21:15:08 INFO ImageMetaData | |||
2023-01-18 21:15:08 INFO ImageMetaData Direction reference : J2000 | |||
2023-01-18 21:15:08 INFO ImageMetaData Spectral reference : BARY | |||
2023-01-18 21:15:08 INFO ImageMetaData Velocity type : RADIO | |||
2023-01-18 21:15:08 INFO ImageMetaData Rest frequency : 1.42041e+09 Hz | |||
2023-01-18 21:15:08 INFO ImageMetaData Pointing center : 15:22:00.000000 +05.04.00.000000 | |||
2023-01-18 21:15:08 INFO ImageMetaData Telescope : VLA | |||
2023-01-18 21:15:08 INFO ImageMetaData Observer : TEST | |||
2023-01-18 21:15:08 INFO ImageMetaData Date observation : 1995/04/13/09:33:00.000687 | |||
2023-01-18 21:15:08 INFO ImageMetaData Telescope position: [-1.60119e+06m, -5.04198e+06m, 3.55488e+06m] (ITRF) | |||
2023-01-18 21:15:08 INFO ImageMetaData | |||
2023-01-18 21:15:08 INFO ImageMetaData Axis Coord Type Name Proj Shape Tile Coord value at pixel Coord incr Units | |||
2023-01-18 21:15:08 INFO ImageMetaData ------------------------------------------------------------------------------------------------ | |||
2023-01-18 21:15:08 INFO ImageMetaData 0 0 Direction Right Ascension SIN 256 64 15:22:00.000 128.00 -1.500000e+01 arcsec | |||
2023-01-18 21:15:08 INFO ImageMetaData 1 0 Direction Declination SIN 256 64 +05.04.00.000 128.00 1.500000e+01 arcsec | |||
2023-01-18 21:15:08 INFO ImageMetaData 2 1 Stokes Stokes 1 1 I | |||
2023-01-18 21:15:08 INFO ImageMetaData 3 2 Spectral Frequency 63 9 1.41273e+09 0.00 2.4415203e+04 Hz | |||
2023-01-18 21:15:08 INFO ImageMetaData Velocity 1619.82 0.00 -5.153101e+00 km/s | |||
2023-01-18 21:15:08 INFO imhead Task imhead complete. Start time: 2023-01-18 14:15:06.134860 End time: 2023-01-18 14:15:07.997282 | |||
2023-01-18 21:15:08 INFO imhead ##### End Task: imhead ##### | |||
2023-01-18 21:15:08 INFO imhead ########################################## | |||
</pre> | </pre> | ||
=== Cube statistics === | |||
We can use [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.information.imstat.html imstat] to display statistics of the cube cubes. The following example displays the statistics for an empty region of the cube (as identified visually in CARTA). | |||
< | <pre style="background-color: #d8bfd8;"> | ||
#Interactive CASA | |||
</ | default instate | ||
inp | |||
imagename = 'ngc5921.demo.clean.image' | |||
box = '10,10,100,100' | |||
inp | |||
go | |||
</pre> | |||
As with imhead, the output goes to the logger window: | |||
<pre> | <pre style="background-color: #fffacd;"> | ||
2023-01-18 21:42:36 INFO imstat ########################################## | |||
2023-01-18 21:42:36 INFO imstat ##### Begin Task: imstat ##### | |||
2023-01-18 21:42:36 INFO imstat imstat( imagename='ngc5921.demo.clean.image', axes=[], region='', box='10,10,100,100', chans='', stokes='', listit=True, verbose=True, mask='', stretch=False, logfile='', append=True, algorithm='classic', fence=-1.0, center='mean', lside=True, zscore=-1.0, maxiter=-1, clmethod='auto', niter=3 ) | |||
2023-01-18 21:42:36 INFO imstat Using specified box(es) 10,10,100,100 | |||
2023-01-18 21:42:36 INFO imstat Determining stats for image ngc5921.demo.clean.image | |||
2023-01-18 21:42:36 INFO imstat Selected bounding box : | |||
2023-01-18 21:42:36 INFO imstat [10, 10, 0, 0] to [100, 100, 0, 62] (15:23:58.379, +04.34.29.305, I, 1.41273e+09Hz to 15:22:28.105, +04.56.59.962, I, 1.41424e+09Hz) | |||
2023-01-18 21:42:36 INFO imstat Statistics calculated using Classic algorithm | |||
2023-01-18 21:42:36 INFO imstat Regions --- | |||
2023-01-18 21:42:36 INFO imstat -- bottom-left corner (pixel) [blc]: [10, 10, 0, 0] | |||
2023-01-18 21:42:36 INFO imstat -- top-right corner (pixel) [trc]: [100, 100, 0, 62] | |||
2023-01-18 21:42:36 INFO imstat -- bottom-left corner (world) [blcf]: 15:23:58.379, +04.34.29.305, I, 1.41273e+09Hz | |||
2023-01-18 21:42:36 INFO imstat -- top-right corner (world) [trcf]: 15:22:28.105, +04.56.59.962, I, 1.41424e+09Hz | |||
2023-01-18 21:42:36 INFO imstat Values --- | |||
2023-01-18 21:42:36 INFO imstat -- flux [flux]: 1.41948 Jy.km/s | |||
2023-01-18 21:42:36 INFO imstat -- number of points [npts]: 521703 | |||
2023-01-18 21:42:36 INFO imstat -- maximum value [max]: 0.0230701 Jy/beam | |||
2023-01-18 21:42:36 INFO imstat -- minimum value [min]: -0.0222025 Jy/beam | |||
2023-01-18 21:42:36 INFO imstat -- position of max value (pixel) [maxpos]: [20, 44, 0, 61] | |||
2023-01-18 21:42:36 INFO imstat -- position of min value (pixel) [minpos]: [35, 70, 0, 61] | |||
2023-01-18 21:42:36 INFO imstat -- position of max value (world) [maxposf]: 15:23:48.368, +04.42.59.428, I, 1.41422e+09Hz | |||
2023-01-18 21:42:36 INFO imstat -- position of min value (world) [minposf]: 15:23:33.331, +04.49.29.579, I, 1.41422e+09Hz | |||
2023-01-18 21:42:36 INFO imstat -- Sum of pixel values [sum]: 3.46393 Jy/beam | |||
2023-01-18 21:42:36 INFO imstat -- Sum of squared pixel values [sumsq]: 2.32674 Jy/beam.Jy/beam | |||
2023-01-18 21:42:36 INFO imstat Statistics --- | |||
2023-01-18 21:42:36 INFO imstat -- Mean of the pixel values [mean]: 6.63965e-06 Jy/beam | |||
2023-01-18 21:42:36 INFO imstat -- Variance of the pixel values : 4.45985e-06 Jy/beam | |||
2023-01-18 21:42:36 INFO imstat -- Standard deviation of the Mean [sigma]: 0.00211184 Jy/beam | |||
2023-01-18 21:42:36 INFO imstat -- Root mean square [rms]: 0.00211184 Jy/beam | |||
2023-01-18 21:42:36 INFO imstat -- Median of the pixel values [median]: 0 Jy/beam | |||
2023-01-18 21:42:36 INFO imstat -- Median of the deviations [medabsdevmed]: 0.00126323 Jy/beam | |||
2023-01-18 21:42:36 INFO imstat -- IQR [quartile]: 0.00252672 Jy/beam | |||
2023-01-18 21:42:36 INFO imstat -- First quartile [q1]: -0.00125785 Jy/beam | |||
2023-01-18 21:42:36 INFO imstat -- Third quartile [q3]: 0.00126887 Jy/beam | |||
2023-01-18 21:42:36 INFO imstat Sum column unit = Jy/beam | |||
2023-01-18 21:42:36 INFO imstat Mean column unit = Jy/beam | |||
2023-01-18 21:42:36 INFO imstat Std_dev column unit = Jy/beam | |||
2023-01-18 21:42:36 INFO imstat Minimum column unit = Jy/beam | |||
2023-01-18 21:42:36 INFO imstat Maximum column unit = Jy/beam | |||
2023-01-18 21:42:36 INFO imstat Npts Sum Mean Rms Std_dev Minimum Maximum | |||
2023-01-18 21:42:36 INFO imstat 5.217030e+05 3.463928e+00 6.639655e-06 2.111845e-03 2.111836e-03 -2.220254e-02 2.307010e-02 | |||
2023-01-18 21:42:36 INFO imstat Task imstat complete. Start time: 2023-01-18 14:42:36.025027 End time: 2023-01-18 14:42:36.481128 | |||
2023-01-18 21:42:36 INFO imstat ##### End Task: imstat ##### | |||
2023-01-18 21:42:36 INFO imstat ########################################## | |||
</pre> | </pre> | ||
Alternatively, we can use the cleanbox.crtf file created earlier to examine the stats in the region containing the source: | |||
=== | <pre style="background-color: #d8bfd8;"> | ||
#Interactive CASA | |||
default instate | |||
inp | |||
imagename = 'ngc5921.demo.clean.image' | |||
box = '' | |||
region='cleanbox.crtf' | |||
inp | |||
go | |||
</pre> | |||
Which gives the output: | |||
<pre style="background-color: #fffacd;"> | |||
2023-01-18 21:44:58 INFO imstat ########################################## | |||
2023-01-18 21:44:58 INFO imstat ##### Begin Task: imstat ##### | |||
2023-01-18 21:44:58 INFO imstat imstat( imagename='ngc5921.demo.clean.image', axes=[], region='cleanbox.crtf', box='', chans='', stokes='', listit=True, verbose=True, mask='', stretch=False, logfile='', append=True, algorithm='classic', fence=-1.0, center='mean', lside=True, zscore=-1.0, maxiter=-1, clmethod='auto', niter=3 ) | |||
2023-01-18 21:44:59 INFO imstat RegionTextParser::_determineVersion: Found spec version 0 | |||
2023-01-18 21:44:59 INFO imstat Combined 1 image regions (which excludes any annotation regions) | |||
2023-01-18 21:44:59 INFO imstat The specified region will select all pixels that are included in the region. Full pixels will be included even when they are only partially covered by the region(s). | |||
2023-01-18 21:44:59 INFO imstat Region read from CRTF file cleanbox.crtf | |||
2023-01-18 21:44:59 INFO imstat Determining stats for image ngc5921.demo.clean.image | |||
2023-01-18 21:44:59 INFO imstat Selected bounding box : | |||
2023-01-18 21:44:59 INFO imstat [111, 107, 0, 0] to [150, 151, 0, 62] (15:22:17.064, +04.58.44.986, I, 1.41273e+09Hz to 15:21:37.910, +05.09.44.977, I, 1.41424e+09Hz) | |||
2023-01-18 21:44:59 INFO imstat Statistics calculated using Classic algorithm | |||
2023-01-18 21:44:59 INFO imstat Regions --- | |||
2023-01-18 21:44:59 INFO imstat -- bottom-left corner (pixel) [blc]: [111, 107, 0, 0] | |||
2023-01-18 21:44:59 INFO imstat -- top-right corner (pixel) [trc]: [150, 151, 0, 62] | |||
2023-01-18 21:44:59 INFO imstat -- bottom-left corner (world) [blcf]: 15:22:17.064, +04.58.44.986, I, 1.41273e+09Hz | |||
2023-01-18 21:44:59 INFO imstat -- top-right corner (world) [trcf]: 15:21:37.910, +05.09.44.977, I, 1.41424e+09Hz | |||
2023-01-18 21:44:59 INFO imstat Values --- | |||
2023-01-18 21:44:59 INFO imstat -- flux [flux]: 28.2043 Jy.km/s | |||
2023-01-18 21:44:59 INFO imstat -- number of points [npts]: 113400 | |||
2023-01-18 21:44:59 INFO imstat -- maximum value [max]: 0.0559169 Jy/beam | |||
2023-01-18 21:44:59 INFO imstat -- minimum value [min]: -0.0221281 Jy/beam | |||
2023-01-18 21:44:59 INFO imstat -- position of max value (pixel) [maxpos]: [134, 134, 0, 43] | |||
2023-01-18 21:44:59 INFO imstat -- position of min value (pixel) [minpos]: [145, 138, 0, 61] | |||
2023-01-18 21:44:59 INFO imstat -- position of max value (world) [maxposf]: 15:21:53.976, +05.05.29.998, I, 1.41378e+09Hz | |||
2023-01-18 21:44:59 INFO imstat -- position of min value (world) [minposf]: 15:21:42.932, +05.06.29.986, I, 1.41422e+09Hz | |||
2023-01-18 21:44:59 INFO imstat -- Sum of pixel values [sum]: 68.8153 Jy/beam | |||
2023-01-18 21:44:59 INFO imstat -- Sum of squared pixel values [sumsq]: 2.27782 Jy/beam.Jy/beam | |||
2023-01-18 21:44:59 INFO imstat Statistics --- | |||
2023-01-18 21:44:59 INFO imstat -- Mean of the pixel values [mean]: 0.000606837 Jy/beam | |||
2023-01-18 21:44:59 INFO imstat -- Variance of the pixel values : 1.97185e-05 Jy/beam | |||
2023-01-18 21:44:59 INFO imstat -- Standard deviation of the Mean [sigma]: 0.00444055 Jy/beam | |||
2023-01-18 21:44:59 INFO imstat -- Root mean square [rms]: 0.0044818 Jy/beam | |||
2023-01-18 21:44:59 INFO imstat -- Median of the pixel values [median]: 0 Jy/beam | |||
2023-01-18 21:44:59 INFO imstat -- Median of the deviations [medabsdevmed]: 0.00138569 Jy/beam | |||
2023-01-18 21:44:59 INFO imstat -- IQR [quartile]: 0.00277213 Jy/beam | |||
2023-01-18 21:44:59 INFO imstat -- First quartile [q1]: -0.001358 Jy/beam | |||
2023-01-18 21:44:59 INFO imstat -- Third quartile [q3]: 0.00141413 Jy/beam | |||
2023-01-18 21:44:59 INFO imstat Sum column unit = Jy/beam | |||
2023-01-18 21:44:59 INFO imstat Mean column unit = Jy/beam | |||
2023-01-18 21:44:59 INFO imstat Std_dev column unit = Jy/beam | |||
2023-01-18 21:44:59 INFO imstat Minimum column unit = Jy/beam | |||
2023-01-18 21:44:59 INFO imstat Maximum column unit = Jy/beam | |||
2023-01-18 21:44:59 INFO imstat Npts Sum Mean Rms Std_dev Minimum Maximum | |||
2023-01-18 21:44:59 INFO imstat 1.134000e+05 6.881528e+01 6.068366e-04 4.481805e-03 4.440551e-03 -2.212812e-02 5.591685e-02 | |||
2023-01-18 21:44:59 INFO imstat Task imstat complete. Start time: 2023-01-18 14:44:58.195518 End time: 2023-01-18 14:44:59.067297 | |||
2023-01-18 21:44:59 INFO imstat ##### End Task: imstat ##### | |||
2023-01-18 21:44:59 INFO imstat ########################################## | |||
</pre> | |||
=== The integrated spectrum === | |||
[[File:spectrum-integrated_6.5.2.png|thumb|Example of the CARTA rectangle selection tool on one channel of the NGC 5921 21 cm data cube. The spectral profile window is shown at right. This is the same channel shown in the dirty cube [[#Imaging|above]]. Click to enlarge.]] | |||
CARTA can be used to produce an integrated spectrum from the spectral line cube. Once you have defined a region (you can use the region used earlier to define the region to be cleaned), you can set the Z profile statistic to 'Sum' to see the spectrum of the pixels summed in this region in each channel, 'Mean' to see the mean spectrum, or 'FluxDensity' to see the beam-corrected flux density spectrum (which is probably what you want). Different plotting and smoothing options are available, and you can export an image of the spectrum or export the data (as a tab-separated variables file, which you can then read into your favorite plotting program, iPython notebook, etc.). | |||
=== Cube moments === | |||
[[File:mom0_mom1_casa6_5_2.png|thumb|The moment 0 (integrated intensity) and moment 1 (velocity field) 21 cm images of NGC 5921, produced using [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.analysis.immoments.html immoments] and displayed with CARTA. Click to enlarge.]] | |||
Cube moments are maps of weighted sums along the velocity axis. In CASA, they are generated by the task [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.analysis.immoments.html immoments]. The zeroth moment (moments = 0) is a sum of intensities along the velocity axis (the integrated intensity map); the first moment (moment = 1) is the sum of velocities weighted by intensity (the ''velocity field''); the second moment (moment = 2) is a map of the velocity dispersion; see the immoments documentation for additional options. | |||
The following example produces maps of the zeroth and first moments, or the integrated intensity and velocity field. To form these maps, we will mask out pixels without signal (below 3 sigma) using hard global limits. We will also collapse along the spectral (channel) axis (the default) and include those channels that show flux in the integrated spectrum, i.e. 9~48. | |||
<pre style="background-color: #d8bfd8;"> | |||
#Interactive CASA | |||
default immoments | |||
inp | |||
imagename='ngc5921.demo.clean.image' | |||
moments=[0,1] | |||
chans='9~48' | |||
excludepix=[-100,0.00525] | |||
outfile='ngc5921.demo.moment' | |||
inp | |||
go | |||
</pre> | |||
<source lang="python"> | <source lang="python"> | ||
immoments(imagename='ngc5921.demo. | #Pseudo-interactive CASA | ||
immoments(imagename='ngc5921.demo.clean.image', moments=[0,1], excludepix=[-100, 0.00525], axis='spectral', chans='9~48', outfile='ngc5921.demo.moment') | |||
</source> | </source> | ||
Where: | |||
< | <pre style="background-color:lightgrey;"> | ||
imagename='ngc5921.demo.clean.image' # input image cube | |||
moments = [0,1] # do zeroth and first moments | |||
# | excludepix = [-100,0.00525] # use only pixels > 3 sigma | ||
moments = [0,1] | outfile='ngc5921.demo.moment' # the output filename root (or filename if only doing one moment) | ||
# | </pre> | ||
excludepix = [-100,0. | |||
# | |||
outfile = | |||
You will get a lot of warnings in the logger and the terminal about the convolving kernel having a size less than the pixel diagonal length. This is because immoments has to convolve all of the channels to have the same beam size prior to forming the moment map, but as they all have very similar beam sizes already the convolving kernel needed for this is fairly small. These warnings can be ignored. | |||
Two files are created by this: ''ngc5921.demo.moment.integrated'' (the zeroth moment) and ''ngc5921.demo.moment.weighted_coord'' (the first moment). These can be opened and manipulated in CARTA. | |||
== Export the Data == | == Export the Data == | ||
To export the | To export the uv data and the image cube as FITS files, use [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.data.exportuvfits.html exportuvfits] and [https://casadocs.readthedocs.io/en/v6.5.2/api/tt/casatasks.data.exportfits.html exportfits] respectively. | ||
For example, to export the continuum-subtracted uv data (the measurement set we used as input to tclean): | |||
<pre style="background-color: #d8bfd8;"> | |||
#Interactive python | |||
default exportuvfits | |||
inp | |||
vis='ngc5921.demo.uvcontsub.ms' | |||
fitsfile='ngc5921.demo.uvcontsub.uvfits' | |||
datacolumn='data' | |||
inp | |||
go | |||
</pre> | |||
<source lang="python"> | <source lang="python"> | ||
exportuvfits(vis='ngc5921.demo. | #Pseudo-interactive python | ||
exportuvfits(vis='ngc5921.demo.uvcontsub.ms', fitsfile='ngc5921.demo.uvcontsub.uvfits', datacolumn='data') | |||
</source> | </source> | ||
Note: the default is to export the CORRECTED_DATA column. If we wanted to export the full data-set after applying the calibration this would be appropriate, but uvcontsub writes out to the DATA column into a new measurement set. | |||
default | |||
And | And for the cleaned cube: | ||
< | <pre style="background-color: #d8bfd8;"> | ||
default | #Interactive python | ||
default exportfits | |||
inp | |||
imagename='ngc5921.demo.clean.image' | |||
fitsimage='ngc5921.demo.clean.fits' | |||
velocity=True | |||
optical=True | |||
inp | |||
go | |||
</pre> | |||
In this example, we have chosen to write the FITS cube with velocity rather than frequency on the spectral axis and to use the optical (''cz'') convention for velocity. Both of these are, of course, optional. | |||
<source lang="python"> | |||
#Pseudo-interactive python | |||
exportfits(imagename='ngc5921.demo.clean.image', fitsimage='ngc5921.demo.clean.fits') | |||
</source> | |||
The moment maps (or any other CASA images) can be similarly exported using exportfits. | |||
== Appendix: Python Notes == | == Appendix: Python Notes == | ||
| Line 1,542: | Line 1,600: | ||
[[Pre-upgrade VLA Tutorials | ↵ '''Pre-upgrade VLA Tutorials''']] | [[Pre-upgrade VLA Tutorials | ↵ '''Pre-upgrade VLA Tutorials''']] | ||
<div style="background-color: #CCFF99;"> | |||
Last checked on CASA Version 6.5.2. | |||
</div> | |||
Latest revision as of 00:51, 1 February 2023
Disclaimer: Due to continuous CASA software updates, GUI images may look different on more recent versions than those shown here.
This guide is meant to be used with monolithic CASA and not pip-wheel, because the GUIs are not necessarily validated.
We would like to note that the CASAviewer has not been maintained for a few years anymore and will be removed from future versions of CASA. The NRAO replacement visualization tool for images and cubes is CARTA, the “Cube Analysis and Rendering Tool for Astronomy”. It is available from the CARTA website. We strongly recommend using CARTA, as it provides a much more efficient, stable, and feature rich user experience. A comparison of the CASAviewer and CARTA, as well as instructions on how to use CARTA at NRAO is provided in the relevant section of CASA docs. This tutorial shows figures generated with CARTA for visualization.
Last checked on CASA Version 6.5.2.
Overview
The technique used to calibrate and image continuum datasets generally applies to spectral line observations, except that an additional calibration step is required. Bandpass calibration flattens the spectral response of the observations, ensuring that spectral channel images are properly calibrated in amplitude and phase.
The following tutorial derives from an annotated script originally provided in the CASA Cookbook.
DATA: The data are included with the CASA installation.
Setting up the CASA environment
Start up CASA in the directory you want to use.
# in bash
mkdir NGC5921
cd NGC5921
casa
We'll use a python os command to get the appropriate CASA path for your installation in order to import the data. The use of os.environ.get is explained in the Appendix. This is not part of the data reduction, just an initial step to find where the example dataset has been stored when CASA was installed – you will not normally need to do this if you are working on your own data.
import os
pathname=os.environ.get('CASAPATH').split()[0]
fitsdata=pathname+'/data/demo/NGC5921.fits'
You can see the value of the variable fitsdata, which is the path to the fits file, using the python print function:
print(fitsdata)
To help clean up the bookkeeping and further avoid issues of write privileges, remove any prior versions of the measurement set and calibration tables. This should be done with the rmtables task, either interactively or using rmtables('table_name'), in preference to the operating system rm -rf command, as rmtables also clears data in the CASA cache.
Import the data
The next step is to import the multisource UVFITS data to a CASA measurement set via the importuvfits task. This task is used to read in data UVFITS data from the legacy VLA or other telescope; for reading in EVLA data in ASDM format the task importasdm would be used.
Note that you can set each parameter for any particular task one-by-one ('interactive CASA', or you can supply the task and input parameters with a single command ('pseudo-interactive CASA').
# Interactive CASA
default importuvfits
inp
fitsfile = fitsdata
vis = 'ngc5921.demo.ms'
antnamescheme = 'new'
inp
saveinputs('importuvfits', 'ngc5921.demo.importuvfits.saved') #optional
go
# Pseudo-interactive CASA
importuvfits(fitsfile=fitsdata,vis="ngc5921.demo.ms")
Where:
default importuvfits # loads importuvfits with the default parameters
inp # lists the inputs available for this task
fitsfile='filename.fits' # sets the filename of the fits file to use
antnamescheme='new' # uses antenna names of the form VANN rather than NN
vis='filename.ms' # sets the name of the output measurement set created (.ms)
go # executes the task with the given inputs
A few things to note:
- The first inp reveals three parameters with defaults fitsfile=' '; vis=' '; antnamescheme='old'.
- fitsfile=fitsdata sets the value of fitsfile to the fitsdata string variable defined earlier. If you were using your own dataset, you could enter it directly as a string (e.g., fitsfile='mydata.fits') or define it as a variable before setting fitsfile (e.g. fitsdata='mydata.fits', then fitsfile=fitsdata). You should avoid using 'fitsfile' as a variable name, as this will be overwritten by 'default importuvfits' or 'tget importuvfits'.
- The second inp shows that the string variable fitsdata, defined earlier, has been expanded. A valid entry shows in blue, while an invalid entry shows in red. Default entries (as for antnamescheme in the initial listing of inputs) show in black.
Inspecting the data
The next step is to inspect to measurement set using the listobs task. This provides almost all relevant observational parameters – to the extent that the exist in the dataset – such as correlator setup (frequencies, bandwidths, channel number and widths, polarization products), sources, scans, scan intents, and antenna locations. Setting verbose=True (the default, so not included in the commands below) will display all of the contents of the raw data and setting listfile='listobs.txt' will create a text file you can refer to later (if this is not set, the output will instead display in the CASA Logger). Some of the parameters (such as scan intents) are not available for legacy VLA data such as this, so are left blank in the output.
# Interactive CASA default listobs inp vis='ngc5921.demo.ms' listfile='listobs.txt' inp go
listobs(vis='ngc5921.demo.ms', listfile='listobs.txt')
You will get a summary, not well formatted for human readability, in the CASA terminal. It is best to ignore this and look at the output text file, which has more information and is better formatted. This looks like:
================================================================================
MeasurementSet Name: <path to directory>/ngc5921.demo.ms MS Version 2
================================================================================
Observer: TEST Project:
Observation: VLA
Data records: 22653 Total elapsed time = 5310 seconds
Observed from 13-Apr-1995/09:18:45.0 to 13-Apr-1995/10:47:15.0 (TAI)
ObservationID = 0 ArrayID = 0
Date Timerange (TAI) Scan FldId FieldName nRows SpwIds Average Interval(s) ScanIntent
13-Apr-1995/09:18:45.0 - 09:24:45.0 1 0 1331+30500002_0 4509 [0] [30]
09:27:15.0 - 09:29:45.0 2 1 1445+09900002_0 1890 [0] [30]
09:32:45.0 - 09:48:15.0 3 2 N5921_2 6048 [0] [30]
09:50:15.0 - 09:51:15.0 4 1 1445+09900002_0 756 [0] [30]
10:21:45.0 - 10:23:15.0 5 1 1445+09900002_0 1134 [0] [30]
10:25:45.0 - 10:43:15.0 6 2 N5921_2 6804 [0] [30]
10:45:15.0 - 10:47:15.0 7 1 1445+09900002_0 1512 [0] [30]
(nRows = Total number of rows per scan)
Fields: 3
ID Code Name RA Decl Epoch SrcId nRows
0 1331+30500002_0 13:31:08.287300 +30.30.32.95900 J2000 0 4509
1 1445+09900002_0 14:45:16.465600 +09.58.36.07300 J2000 1 5292
2 N5921_2 15:22:00.000000 +05.04.00.00000 J2000 2 12852
Spectral Windows: (1 unique spectral windows and 1 unique polarization setups)
SpwID Name #Chans Frame Ch0(MHz) ChanWid(kHz) TotBW(kHz) CtrFreq(MHz) Corrs
0 none 63 LSRK 1412.665 24.414 1550.2 1413.4219 RR LL
Sources: 3
ID Name SpwId RestFreq(MHz) SysVel(km/s)
0 1331+30500002_0 0 1420.405752 0
1 1445+09900002_0 0 1420.405752 0
2 N5921_2 0 1420.405752 0
Antennas: 27:
ID Name Station Diam. Long. Lat. Offset from array center (m) ITRF Geocentric coordinates (m)
East North Elevation x y z
0 VA01 VLA:N7 25.0 m -107.37.07.2 +33.54.12.9 -30.2623 345.7477 -0.8872 -1601155.613187 -5041783.882304 3555162.343090
1 VA02 VLA:W1 25.0 m -107.37.05.9 +33.54.00.5 3.5004 -39.7725 0.9883 -1601188.991307 -5042000.530918 3554843.409670
2 VA03 VLA:W2 25.0 m -107.37.07.4 +33.54.00.9 -37.1358 -25.0262 1.0383 -1601225.244615 -5041980.431775 3554855.677111
3 VA04 VLA:E1 25.0 m -107.37.05.7 +33.53.59.2 6.9833 -79.6414 1.1565 -1601192.444530 -5042022.911771 3554810.411780
4 VA05 VLA:E3 25.0 m -107.37.02.8 +33.54.00.5 81.5188 -37.9632 1.0246 -1601114.335629 -5042023.211477 3554844.931655
5 VA06 VLA:E9 25.0 m -107.36.45.1 +33.53.53.6 536.8977 -250.3175 0.1183 -1600715.915813 -5042273.186780 3554668.167811
6 VA07 VLA:E6 25.0 m -107.36.55.6 +33.53.57.7 267.7566 -124.8145 1.2815 -1600951.554888 -5042125.947753 3554772.987072
7 VA08 VLA:W8 25.0 m -107.37.21.6 +33.53.53.0 -401.2640 -270.6305 2.2293 -1601614.059494 -5042001.699973 3554652.484758
8 VA09 VLA:N5 25.0 m -107.37.06.7 +33.54.08.0 -16.9948 194.1215 -0.1368 -1601168.756077 -5041869.099542 3555036.914367
9 VA10 VLA:W3 25.0 m -107.37.08.9 +33.54.00.1 -74.4964 -50.1921 1.1608 -1601265.132224 -5041982.597979 3554834.857504
10 VA11 VLA:N4 25.0 m -107.37.06.5 +33.54.06.1 -11.7487 134.3686 0.1774 -1601173.922897 -5041902.701204 3554987.495105
11 VA12 VLA:W5 25.0 m -107.37.13.0 +33.53.57.8 -179.2554 -120.8635 1.4872 -1601376.990711 -5041988.712764 3554776.381187
12 VA13 VLA:N3 25.0 m -107.37.06.3 +33.54.04.8 -8.2438 94.5297 0.3947 -1601177.362708 -5041925.112425 3554954.550128
13 VA14 VLA:N1 25.0 m -107.37.06.0 +33.54.01.8 -0.0030 0.0445 0.8773 -1601185.580779 -5041978.216463 3554876.396287
14 VA15 VLA:N2 25.0 m -107.37.06.2 +33.54.03.5 -4.7904 54.7090 0.5774 -1601180.839839 -5041947.470902 3554921.600805
15 VA16 VLA:E7 25.0 m -107.36.52.4 +33.53.56.5 348.8969 -162.6653 1.0336 -1600880.544215 -5042170.427468 3554741.431900
16 VA17 VLA:E8 25.0 m -107.36.48.9 +33.53.55.1 438.6654 -204.5038 0.5027 -1600801.910482 -5042219.412805 3554706.408864
17 VA18 VLA:W4 25.0 m -107.37.10.8 +33.53.59.1 -122.0163 -82.2819 1.2624 -1601315.866196 -5041985.352573 3554808.279150
18 VA19 VLA:E5 25.0 m -107.36.58.4 +33.53.58.8 195.8349 -91.2758 1.2155 -1601014.427180 -5042086.300814 3554800.787928
19 VA20 VLA:W9 25.0 m -107.37.25.1 +33.53.51.0 -491.1000 -331.2429 2.5539 -1601709.998072 -5042006.975455 3554602.355417
20 VA21 VLA:W6 25.0 m -107.37.15.6 +33.53.56.4 -244.9704 -165.2178 1.6861 -1601447.161927 -5041992.554228 3554739.677219
21 VA22 VLA:E4 25.0 m -107.37.00.8 +33.53.59.7 133.6478 -62.2829 1.0919 -1601068.773396 -5042051.970054 3554824.783566
23 VA24 VLA:E2 25.0 m -107.37.04.4 +33.54.01.1 40.6649 -18.9151 0.9550 -1601150.040469 -5042000.665669 3554860.702914
24 VA25 VLA:N6 25.0 m -107.37.06.9 +33.54.10.3 -23.2197 265.3902 -0.4819 -1601162.569974 -5041829.054708 3555095.873969
25 VA26 VLA:N9 25.0 m -107.37.07.8 +33.54.19.0 -46.5533 532.1581 -1.8550 -1601139.422904 -5041679.082136 3555316.518142
26 VA27 VLA:N8 25.0 m -107.37.07.5 +33.54.15.8 -38.0437 434.7201 -1.3387 -1601147.894127 -5041733.868915 3555235.935926
27 VA28 VLA:W7 25.0 m -107.37.18.4 +33.53.54.8 -319.1171 -215.2368 1.9407 -1601526.340031 -5041996.897001 3554698.302182
Key information from listobs
The output of listobs is dense with information, but some facts to note are (working down the file):
- The observations were taken on 13 April 1995 between 09:18:45 and 10:47:15
- In the list of scans we can see that:
- There is a gap between 09:51:15 and 10:21:45 when the array was making other observations that are not included in this demo dataset
- The averaging interval for all of the scans is 30s
- There are three fields listed under "Fields": 1331+305*, 1445+099* and N5921_2.
- As this is legacy VLA data, no scan intents are given. However, we can identify 1331+305 (field=0_ as 3C286, the flux and bandpass calibrator, 1445+099 (field=1) as the phase calibrator, and N5921_2 (field=2) as the science target.
- A single spectral window (IF) is listed under "Spectral Windows", with SpwID = 0.
- This is divided into 63 channels of 24.414 kHz width, centered on 1413.4219 MHz.
- The correlations are RR and LL (cross-pols are absent).
- The three sources listed under "Sources" correspond to the three fields under "Fields", but different information is given linking them to the spectral window(s).
- All sources are associated with SpwID = 0 (if there were multiple spectral windows, there would be a line for each spectral window, repeating sources observed in multiple windows)
- The rest frequency is set to the HI frequency and the systemic velocity to 0 km/s – although the center frequency above is clearly not at 0 km/s, the systemic velocity field is not correctly populated in this legacy dataset.
Log files
EVLA and VLA logs from October 2013 can be accessed via the operator logs] tool. For earlier observations, such as these, the older VLA operator logs page gives logs by month from January 1989. This is a single, large text file for each month and it is necessary to search through to find the relevant observation. For this observation, we find:
************************************************************************ Program: TEST/VANMOORSEL Observer: G. van Moorsel Date: Thursday, Apr 13 1995 User Number: 1102 Source File: 528MTST Subarray: #1 Observe Mode: Line Operator: T. Perreault ** Weather Information ** Time Dew Point Temp. Wind Bar. Pressure Remarks: 95Apr13 08:49:59 -13.3°C 1.1°C W @ 1 m/s 794.6mbars The sky is clear. 95Apr13 10:30:39 -10.9°C 3.3°C NNW @ 1 m/s 794.1mbars The sky is clear. ** Visibility Tape Information ** Tape # File # Time of Final Record: N6078 1 ** Monitor Tape Information ** Tape # File # Time of Final Record: N6080 3 Antenna(s) Used: 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 24 25 26 27 28 ** Operator Comments ** Start Time End Time Form # Ant # Downtime 95Apr13 08:46:40 (in Minutes) Start of observe file. The band(s) used is(are) L band(s). Start Time End Time Form # Ant # Downtime 95Apr13 08:50:00 On source 1331+305 with all available antennas. Start Time End Time Form # Ant # Downtime 95Apr13 09:50:02 The test e-mail message from the computer Zia was received. Observe mail should work fine. End of program TEST/VANMOORSEL. Total Downtime = 0.0 minutes (0.0% of total time). AsciiFileOut Version: 1.00 -- Version Date: March 31, 1995 ******************************************
From this we can see that there are no obvious problems – antenna faults or the like – affecting these observations.
Antenna positions
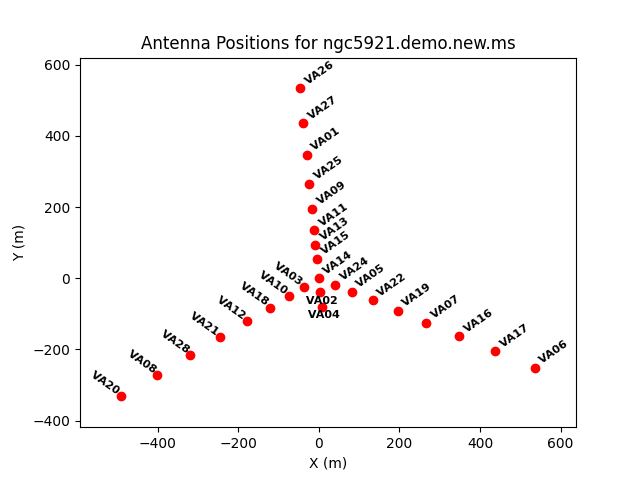
The antenna positions are given in the output from listobs, but a graphical display can be easier to understand and aid in selecting a good reference antenna. This can be plotted using the plotants task.
# Interactive CASA default plotants inp vis='ngc5921.demo.ms' inp go
# Psuedo-interactive CASA
plotants(vis='ngc5921.demo.ms')
Where:
vis='filename.ms'. # sets the name of the input measurement set
Optional parameters:
logpos=True # plots the distance from the center of the array logarithmically
figfile='antlayout.png' # saves the plot to the specified file
Flagging
The next step is to flag the data, for which we use the task flagdata. Flagging is a large topic, so for more details see the VLA CASA Flagging Topical CASA Guide.
Flag the autocorrelations
First, we can flag the autocorrelation data, which are not needed for our reduction:
# Interactive CASA default flagdata inp vis='ngc5921.demo.ms' autocorr=True inp go
# Psuedo-interactive CASA
flagdata(vis="ngc5921.demo.ms", autocorr=True)
CASA issues a warning when running this command, as it is having to make assumptions about the data due to information not being available in legacy VLA files. This can be safely ignored.
If you run this in interactive mode, you will see that there are a lot of options for this task. We're going to use many of them later, but for now we just use vis to set the input measurement set and autocorr=True to say that we want to flag the autocorrelations. One parameter to check is flagbackup=True (which should be set by default). This backs up the flag state before carrying out the requested operation, allowing you to roll back to an earlier state using flagmanager. It is good practice when flagging to note which flagbackup version is associated with which flagging commands in your data reduction log.
Quack flagging
It is common for the array to require a small amount of time to settle down at the start of a scan. Consequently, it has become standard practice to flag the initial samples from the start of each scan. This is known as 'quack' flagging. This has been implemented in flagdata.
As we've already used flagdata, we could start where we were before using 'tget flagdata' rather than 'default flagdata' to restore the last parameters used. This would avoid having to re-enter the vis= command, but at the cost of having to check parameters used before and reset them to the defaults, such as setting autocorr=False. It is safer to return to the defaults by using 'default flagdata', which is what we do here.
# Interactive CASA default flagdata inp vis='ngc5921.demo.ms' mode='quack' quackinterval = 10.0 inp go
# Psuedo-interactive CASA
flagdata(vis="ngc5921.demo.ms", mode='quack', quackinterval='10.0')
If you type 'inp' after setting mode='quack', you will see that additional inputs for this mode have appeared. These allow you to set the quackinterval, here set to 10s (any time less than the averaging interval of the data – 30s as we noted from the listobs output – will select just the first sample); we leave quackmode='beg' at its default value to flag the beginning of each scan. Taken together, these parameters tell flagdata to flag the first sample at the beginning of each scan.
Interactive flagging
We use plotms to inspect the spectral line data. While plotms allows flagging inside the graphical interface, this does not write flagbackup files; we can use plotms to visualize the data in conjunction with using flagdata to actually set the flags in order to have flagbackup files.
To start plotms, you can simply run
plotms
at the CASA terminal prompt, and set the parameters interactively in the graphical interface. This will take any relevant parameters from the current parameter set, which can lead to some unexpected behavior (e.g., if started after flagging a data selection in flagdata, plotms will start using the same data selection and will report that all the data is flagged – if this happens, simply edit the data selection parameters in the graphical interface and reload the data). Alternatively, you can load it as a CASA task and set the parameters to be used on initial load using interactive of pseudo-interactive CASA.
To start, look at the bandpass of 3C286 to see whether there is any narrow-band RFI and to see which channels look useful. We use 3C286 for this as it has higher flux, and therefore higher signal to noise, than the other sources:
# Interactive CASA default plotms inp vis='ngc5921.demo.ms' xaxis='channel' yaxis='amp' field='0' correlation='RR,LL' coloraxis='baseline' datacolumn='data' inp go
#Pseudo-interactive CASA
plotms(vis='ngc5921.demo.ms', xaxis='channel', yaxis='amp', field=0, correlation='RR,LL', coloraxis='baseline', datacolumn='data')
If we want to increase the signal-to-noise, or look for fainter RFI signals, we can use avgtime='360' to average across all of the samples on 3C286. If using CASA interactively, you can change this averaging using:
avgtime='360' go
Alternatively, this can be changed in the plotms GUI.
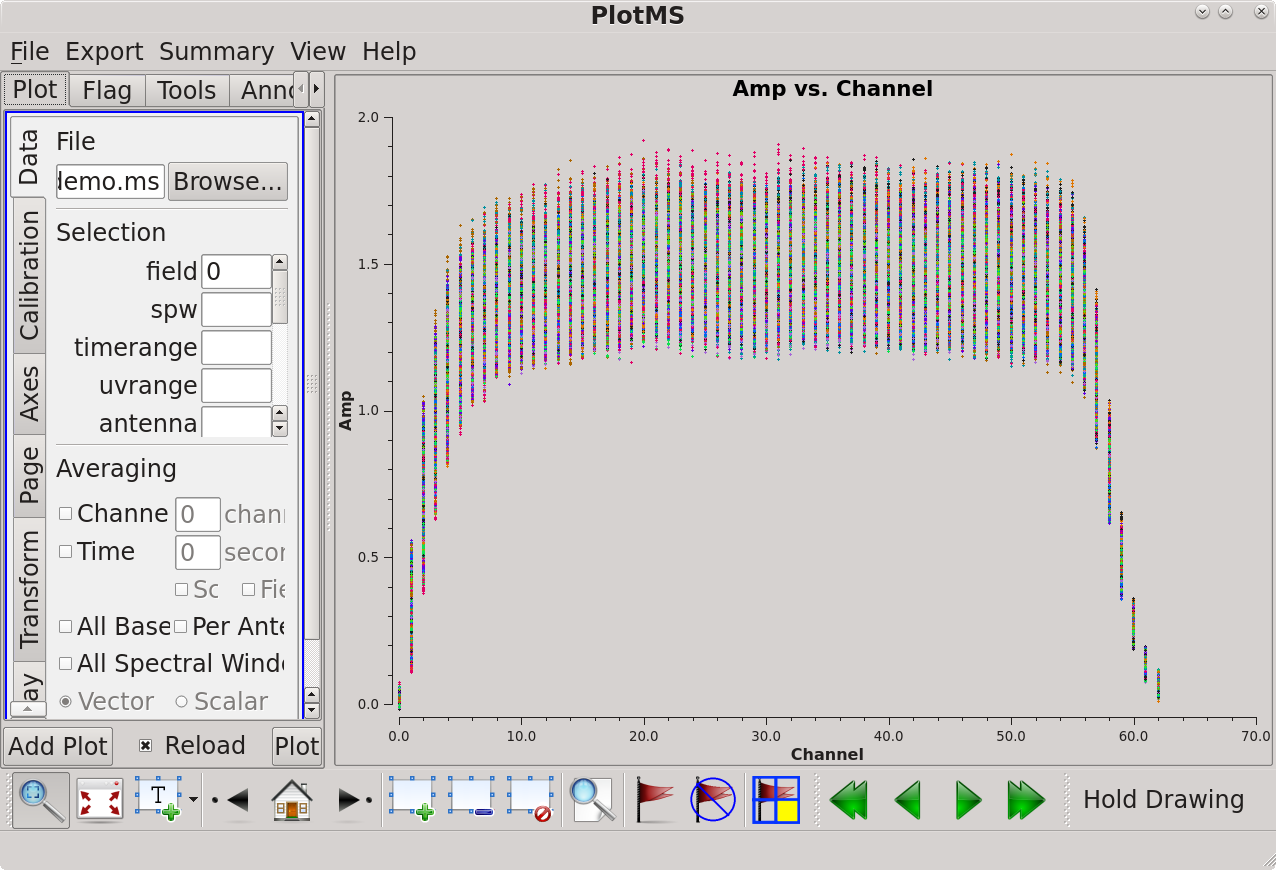
The resulting plot (see figure) shows that there is good signal on channels 6 to 56, which we can specify using spw='0:6~56'. There does not appear to be any visible RFI on this field; before moving on it's worth checking the other fields for RFI as well. This can be done from the GUI or using:
field='N' go
(Where 'N' is either '1' or '2' for the phase calibrator and the target fields respectively.)
To check for antenna-based effects, we next want to look at the summed amplitude across the 51 channels identified above. We can do this on the graphical interface or using:
avgtime='' spw='0:6~56' avgchannel='51' field='' xaxis='time' inp go
Note that here we use avgtime=' ' to unset the time averaging we set previously and field=' ' to unset the field specifier and show all fields. In pseudo-interactive CASA, we don't need to unset the time averaging but we do need to specify all non-defaults every time we run:
#Pseudo-interactive CASA
plotms(vis='ngc5921.demo.ms', xaxis='time', yaxis='amp', spw='0:6~56', avgchannel='51', correlation='RR,LL', coloraxis='baseline', datacolumn='data')
This plots amplitude against time (see figure). We can see from the colored striping on the sources that different baselines (coloraxis='baseline' means the different colors represent different baselines) have slightly different amplitudes, including one baseline (pink on the example plot) that is always high.
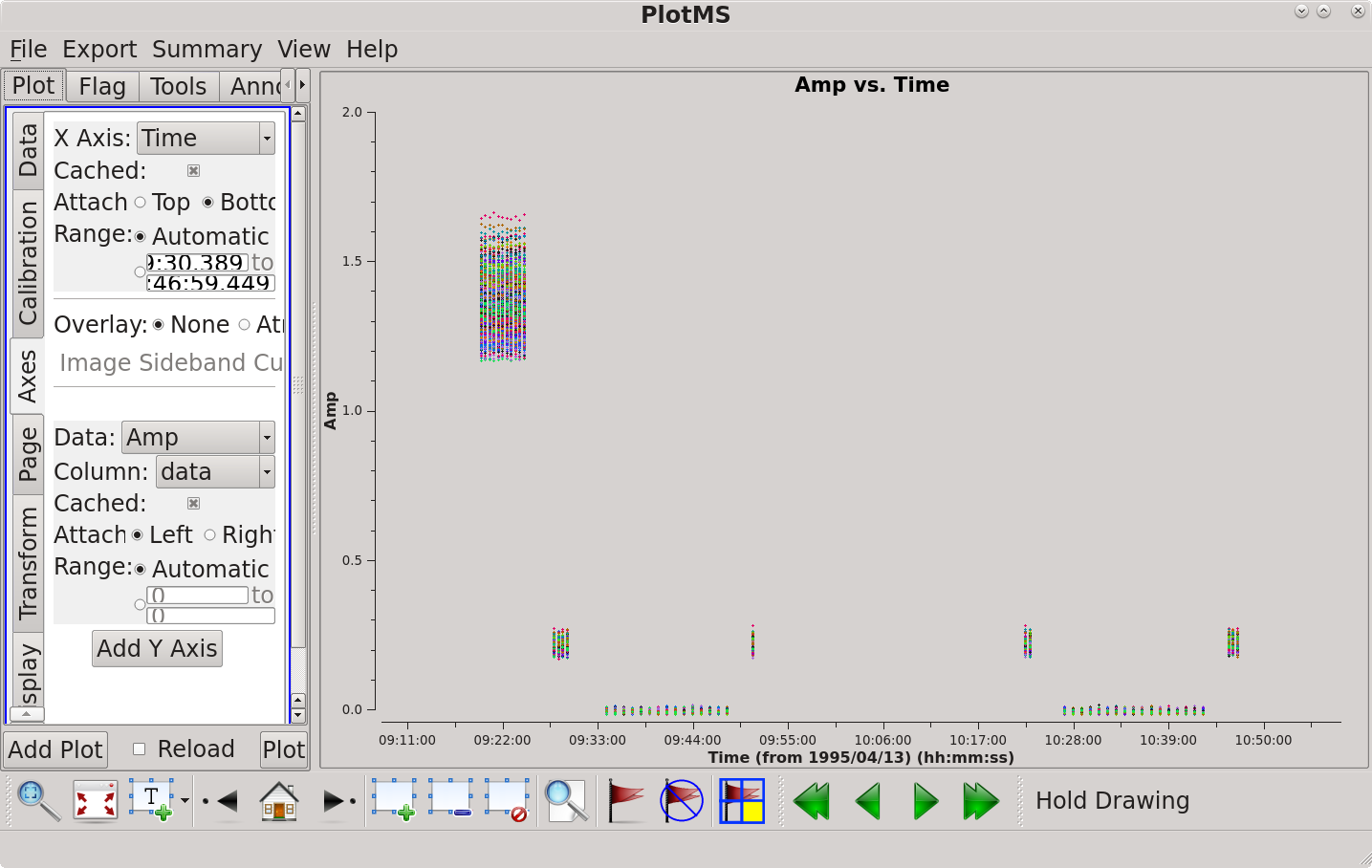
We can draw a box around some of those pink points in plotms by clicking the mark regions ![]() tool to select that function and then drawing a box on the screen to mark a region. This may take a few seconds to return, but eventually the box will be filled with dots showing it has been selected. The points within the marked region can then be inspected using the locate
tool to select that function and then drawing a box on the screen to mark a region. This may take a few seconds to return, but eventually the box will be filled with dots showing it has been selected. The points within the marked region can then be inspected using the locate ![]() tool to the right of the region buttons. This outputs information on the selected points to the CASA logger. Once you are finished with a marked region, you should clear it with the clear regions
tool to the right of the region buttons. This outputs information on the selected points to the CASA logger. Once you are finished with a marked region, you should clear it with the clear regions ![]() tool (attempting to change the plot while a marked region is present will often crash plotms).
tool (attempting to change the plot while a marked region is present will often crash plotms).
On the logger output, the baseline with the high values is identified with "BL=VA12@VLA:W5 & VA21@VLA:W6 [11&20]". This tells us that it is the baselines between antennas VA12 and VA21 at stations VLA:W5 and VLA:W6 with IDs 11 and 20. We can also see that all of the points have "Corr=LL", telling us that it is just the left-hand correlation.
We can plot just this baseline by setting antenna to "VA12&VA21" (or "VLA:W5&VLA:W6") in the GUI or, in interactive CASA:
antenna='VA12&VA21' go
(Note that 'VA12,VA21' would select all baselines from antennas VA12 and VA21, not just the baseline between the two antennas – this is particularly important when flagging using flagdata!)
It can be seen that although the data is slightly high on the LL correlation of this baseline, it isn't particularly noisy. We have not yet calibrated this data, so some variation is possible. It does not look like this is a bad baseline/correlation, but rather that this will be dealt with by calibration.
Flagging data:
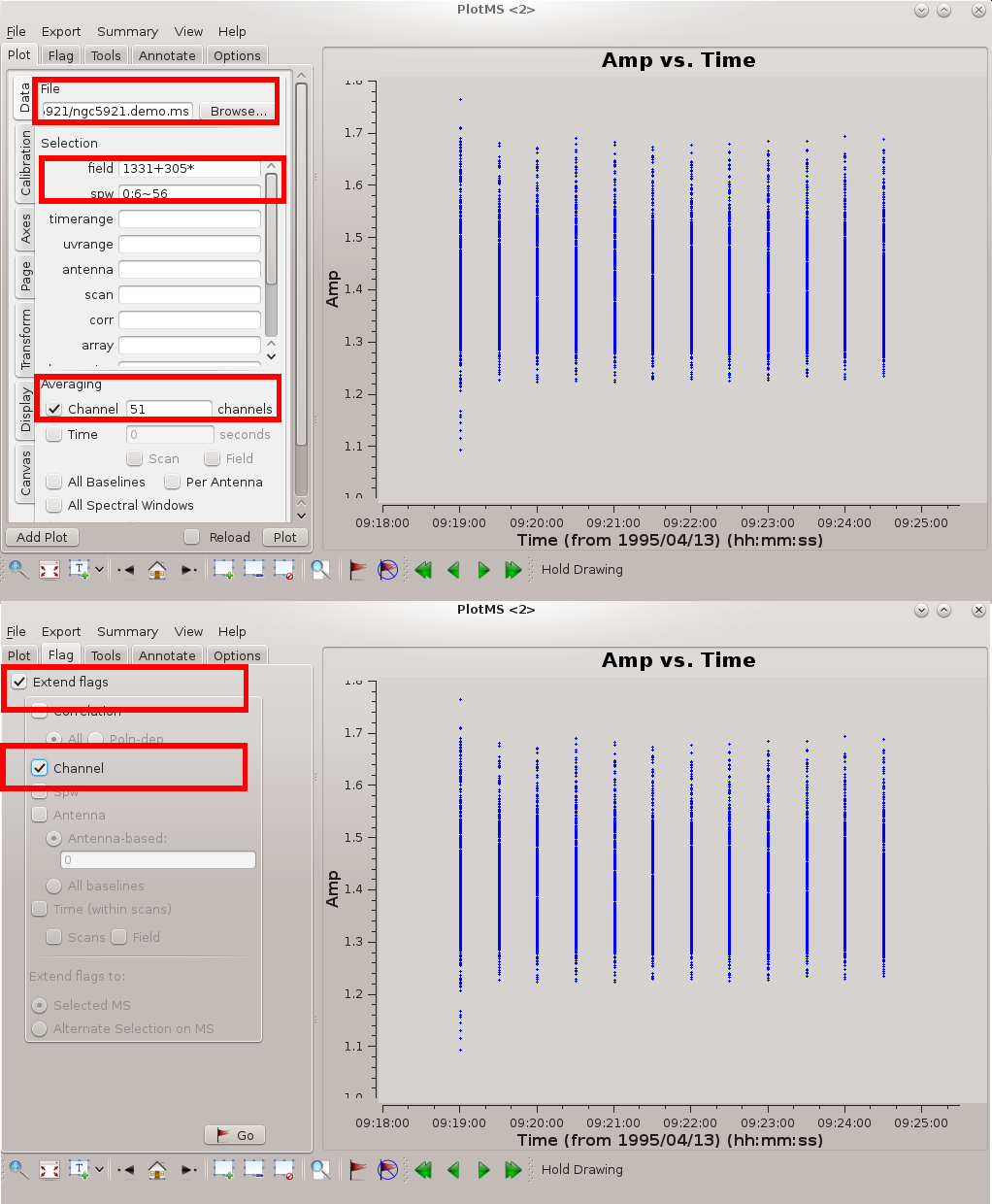
If we did want to flag this data, we could either do so within plotms using the flag ![]() tool or we could use flagdata again. If flagging baselines within plotms, you should ensure that you flag all of the channels; to do this, under the Flag tab, specify Extend flags to Channel (obviously, if flagging RFI channels you should not to this). You can read more in the tutorial on data flagging with plotms. However, if you want flag backups to be created (which is a good idea), you need to flag using flagdata rather than plotms. This can be done using:
tool or we could use flagdata again. If flagging baselines within plotms, you should ensure that you flag all of the channels; to do this, under the Flag tab, specify Extend flags to Channel (obviously, if flagging RFI channels you should not to this). You can read more in the tutorial on data flagging with plotms. However, if you want flag backups to be created (which is a good idea), you need to flag using flagdata rather than plotms. This can be done using:
Do not type 'go' at the end of this script! 'go' has not been included as we're not going to run this as we don't want to actually flag the data.
# Interactive CASA default flagdata inp vis='ngc5921.demo.ms' antenna = 'VA12&VA21' correlation = 'LL' inp
If executed, this would flag the left-hand correlation of the baseline between antennas VA12 and VA21. Other fields displayed by 'inp' in the section under "mode = 'manual'" can be used to set other data selection parameters, such as spw and timerange, so you can select the data to be flagged. If you control plotms from the GUI, you can identify data to be flagged using the mark regions and locate tools and then enter the appropriate parameters into flagdata to flag that data; when running like this you can use 'plotms' at the command line to restart it if it crashes, but while flagdata is the active task 'go' will execute flagdata rather than starting plotms.
Calibration
Calibration of spectral line data broadly follows the approach for continuum data, except that the amplitude and phase corrections are a function of frequency and so must be corrected by bandpass calibration. The basic calibration steps follow.
- Set the flux scale of the primary calibrator, here, 1331+305 = 3C 286.
- A priori calibration
- Determine bandpass corrections based on the primary calibrator. In the script that follows, the bandpass calibration will be stored in bandpass.cal.
- Inspect the bandpass calibration for problems and to determine viable channels for averaging and imaging. We want to toss out end channels where the response is poor.
- Determine the gain calibrations on the bandpass-corrected and channel-averaged data. In this step, we effectively turn the spectral line data into a single-channel continuum data set and calibrate accordingly. The calibration is first stored in apcal.gcal. In the second part of this step we correct the fluxscale of the apcal table, and store the final calibration solutions with correct fluxes in fluxscale.cal (this is the table that needs to be applied later to the data, not the apcal.gcal version).
- Inspect the gain calibration solutions to look for any aberrant solutions that hint at bad calibrator data.
- Apply the calibration solutions to the source (N5921_2). This action literally adds a new column of data to the measurement set. This new column contains the data with the gain calibration and bandpass calibration applied, but it does not overwrite the raw data in case the calibration needs revision.
Setting the flux scale
The setjy task generates a source model for the primary calibrator, 1331+305 = 3C286. From CASA 5.3 onwards, the default standard is Perley-Butler 2017 and includes resolved structure of the calibrators. This is 1.4GHz D-config and 1331+305 is sufficiently unresolved that, in principle, we don't need a model image; however, here we proceed with applying the detailed model, as a good practice. Setjy also looks up the radio SED for common flux calibrators and automatically assigns the total flux density.
We can first get a listing of the available models. Setjy will look within the CASA working directory for files matching *.im* and *.mod*, i.e. user-defined images or models, and in the CalModels directory of the installation. Here we will want to apply a model from CalModels but, in principle, it is possible to use a user-defined model instead.
# Interactive CASA default setjy inp vis='ngc5921.demo.ms' listmodels=True inp go
# Psuedo-interactive CASA
setjy(vis='ngc5921.demo.ms',listmodels=True)
This gives output in both the terminal and the logger for both the working directory and the CalModels directory.
#Logger output: Candidate models (*.im* *.mod*) in .: #Terminal output: ls: cannot access *.mod*: No such file or directory ngc5921.demo.importuvfits.saved
This is the output for the current directory. As the listmodels setting uses the linux command 'ls' to look for files matching *.im* and *.mod*, it has found the ngc5921.demo.importuvfits.saved file created earlier. If you didn't create this file, you would instead get just in the logger window:
#Logger output: No candidate models matching '*.im* *.mod*' found in . # The single period here refers to the current working directory.
With no associated output in the terminal window. The output for the CalModels directory follows immediately after this:
#Logger output: Candidate models (*) in /home/casa/data/distro/nrao/VLA/CalModels: #terminal output: 3C123_P.im 3C138_S.im 3C147_P.im 3C286_C.im 3C286_X.im 3C48_P.im 3C138_A.im 3C138_U.im 3C147_Q.im 3C286_K.im 3C295_P.im 3C48_Q.im 3C138_C.im 3C138_X.im 3C147_S.im 3C286_L.im 3C380_P.im 3C48_S.im 3C138_K.im 3C147_A.im 3C147_U.im 3C286_P.im 3C48_A.im 3C48_U.im 3C138_L.im 3C147_C.im 3C147_X.im 3C286_Q.im 3C48_C.im 3C48_X.im 3C138_P.im 3C147_K.im 3C196_P.im 3C286_S.im 3C48_K.im README 3C138_Q.im 3C147_L.im 3C286_A.im 3C286_U.im 3C48_L.im
From this, we can use the name of our flux calibrator source (3C286) and band (L) and identify the model we want as 3C286_L.im. Note that short codes are used for some bands – A for Ka and U for Ku – and that models are available for additional flux calibrators at P band. We use field='0' to identify the field in our dataset corresponding to 3C286; alternatively we could use the source name and set field='1331+305*' (the wildcard being necessary as there's a string of zeroes after 1331+305 in the source name).
# Interactive CASA default setjy inp vis='ngc5921.demo.ms' field='0' model='3C286_L.im' usescratch=True inp go
# Pseudo-interactive CASA
setjy(vis='ngc5921.demo.ms', field='0', model='3C286_L.im', usescratch=True)
Where:
vis='ngc5921.demo.ms' # defines the name the of dataset
field='0' # defines the field to be used; field 0 = 3C286
model='3C286_L.im' # defines the model to be used, here 3C286 and L-band
usescratch=True # tells CASA to use the MODEL_DATA column
A summary of the operation is sent to the logger window. Here's a listing of the output.
2022-12-02 00:32:41 INFO setjy ##########################################
2022-12-02 00:32:41 INFO setjy ##### Begin Task: setjy #####
2022-12-02 00:32:41 INFO setjy setjy( vis='ngc5921.demo.ms', field='0', spw='', selectdata=False, timerange='', scan='', intent='', observation='', scalebychan=True, standard='Perley-Butler 2017', model='3C286_L.im', listmodels=False, fluxdensity=-1, spix=0.0, reffreq='1GHz', polindex=[], polangle=[], rotmeas=0.0, fluxdict={}, useephemdir=False, interpolation='nearest', usescratch=False, ismms=False )
2022-12-02 00:32:44 INFO setjy {'field': '0'}
2022-12-02 00:32:45 INFO Imager Opening MeasurementSet <path to directory>/ngc5921.demo.ms
2022-12-02 00:32:45 INFO setjy Using /home/casa/data/distro/nrao/VLA/CalModels/3C286_L.im for model.
2022-12-02 00:32:45 INFO imager Using channel dependent flux densities
2022-12-02 00:32:45 INFO imager Selected 4509 out of 22653 rows.
2022-12-02 00:32:45 INFO imager 1331+30500002_0 (fld ind 0) spw 0 [I=15.016, Q=0, U=0, V=0] Jy @ 1.4127e+09Hz, (Perley-Butler 2017)
2022-12-02 00:32:46 INFO imager Using model image /home/casa/data/distro/nrao/VLA/CalModels/3C286_L.im
2022-12-02 00:32:46 INFO imager Scaling spw(s) [0]'s model image by channel to I = 15.0159, 15.0118, 15.0077 Jy @(1.41265e+09, 1.41343e+09, 1.41419e+09)Hz (LSRK) for visibility prediction (a few representative values are shown).
2022-12-02 00:32:46 INFO imager The model image's reference pixel is 0.00904522 arcsec from 1331+30500002_0's phase center.
2022-12-02 00:32:46 INFO imager Will clear any existing model with matching field=1331+30500002_0 and spw=*
2022-12-02 00:32:46 INFO Clearing model records in MS header for selected fields.
2022-12-02 00:32:46 INFO 1331+30500002_0 (id = 0) not found.
2022-12-02 00:32:46 INFO imager Selected 4509 out of 22653 rows.
2022-12-02 00:32:46 INFO setjy Task setjy complete. Start time: 2022-12-01 17:32:41.433462 End time: 2022-12-01 17:32:46.302627
2022-12-02 00:32:46 INFO setjy ##### End Task: setjy #####
2022-12-02 00:32:46 INFO setjy ##########################################
An important piece of information from this is that the flux of 3C286 was set to a bit over 15 Jy, with some slight variation across the frequency range of the data cube, and the data cube was scaled by channel to match this.
A priori calibration
The gencal task carries out general calibrations using information that is known prior to inspecting the data, and thus known as a priori calibration. We can use it to apply the antenna gain curve and (if necessary) correct the antenna positions.
Antenna gain-elevation curve calibration
With elevation, the effective collecting area and surface accuracy of antennas varies as gravity tugs at the surface of the non-rigid antenna. This calibration is only strictly necessary at high frequencies (> 15 GHz), so can be skipped if desired. Using gencal, we will write the gain curve solutions to a calibration table.
# Interactive CASA default gencal inp vis='ngc5921.demo.ms' caltable='antgain.cal' caltype='gc' inp go
# Psuedo-interactive CASA
gencal(vis='ngc5921.demo.ms',caltable='antgain.cal',caltype='gc')
Where:
vis='ngc5921.demo.ms' # the measurement set
caltable='antgain.cal' # name of output calibration table where values are stored
caltype='gc' # calculate the gain curve
Antenna position corrections
Unlike the EVLA, where antenna position offsets can be looked up automatically and applied by gencal using caltype='antpos', you need to look up and apply these corrections manually for the legacy VLA. If needed, and if we have the necessary information, we can correct the antenna positions using gencal with caltype='antposvla'. This takes the offsets to be applied (dBx, dBy, dBz) in meters for a specified antenna. Please see the gencal documentation for further information.
Bandpass calibration
The flux calibrator 1331+305 = 3C 286 now has a model assigned to it from the setjy step. Since the bandwidth of our observations is only 1.55 MHz, the model doesn't change over this narrow range of frequencies, so we can use it to determine amplitude and phase (gain) corrections for each channel independently. The result is the bandpass calibration.
As for any antenna-based calibration scheme, we have to pick an antenna to act as the reference point for the calibration. Any antenna will do, but it's better to pick one near the center of the array, avoiding any that suffered from shadowing or had to be flagged for any other reason. For the remainder of the calibration, we will use refant = 'VA15' (the antenna in position N2). We can first do the bandpass on the single 5min scan on 1331+305. At 1.4GHz phase stablility should be sufficient to do this without a first (rough) gain calibration. This will give us the relative antenna gain as a function of frequency.
# Interactive CASA default bandpass inp vis='ngc5921.demo.ms' caltable='bandpass.cal' field='0' solint='inf' combine='scan' refant='VA15' bandtype='B' gaintable='antgain.cal' # ONLY IF YOU CREATED THIS WITH GENCAL EARLIER inp go
# Pseudo-interactive CASA
bandpass(vis='ngc5921.demo.ms', caltable='bandpass.cal', field='0', solint='inf', combine='scan', refant='VA15', bandtype='B', gaintable='antgain.cal')
Where:
vis='ngc5921.demo.ms'. # Input measurement set
caltable='bandpass.cal' # Output calibration table
field='0' # Use the flux calibrator 1331+305 = 3C286 (FIELD_ID 0) as bandpass calibrator
bandtype='B'. # Choose bandpass solution type. Pick standard time-binned B (rather than BPOLY) [default]
solint='inf' # Set solution interval arbitrarily long (to get single bandpass) [default]
combine='scan' # Combine scans (but break at obs, field and spw boundaries) [default]
refant = 'VA15' # Set antenna VA15 (VLA:N2) as the reference antenna
gaintable='antgain.cal' # Use the antgain.cal table created by gencal (only if you did this optional step)
Inspect the bandpass response curve
In the gain calibration to follow, we will effectively convert the spectral line data into a continuum data set. Before proceeding, we need to inspect the bandpass calibration to make sure that it contains no bad values and also to inspect which channels to average to produce the continuum data.
We will plot the bandpass solutions as amplitude vs channel and phase vs channel per antenna. Start by looking at the amplitudes. These should be smooth across the channels. You will see the edge channels have lower amplitudes due to the edges of the bandpass being less sensitive.
# Interactive CASA default plotms inp vis='bandpass.cal' xaxis='chan' yaxis='amp' coloraxis='corr' iteraxis='antenna' gridrows=3 gridcols=3 xselfscale=True yselfscale=True inp go
# Psuedo-interactive CASA
plotms(vis='bandpass.cal',xaxis='chan',yaxis='amp',coloraxis='corr',iteraxis='antenna',gridrows=3,gridcols=3,xselfscale=True,yselfscale=True)
Where:
vis='bandpass.cal' # the bandpass calibration table
xaxis='chan' # plot channels across the x axis
yaxis='amp' # plot amplitudes across the y axis
coloraxis='corr' # display the two correlations (RR & LL) in different colors
iteraxis='antenna' # pages the plots by antenna
gridrows=3 # places 3 antennas per row
gridcols=3 # places 3 antennas per column
xselfscale=True # create a global xaxis scale for all antennas
yselfscale=True # create a global yaxis scale for all antennas
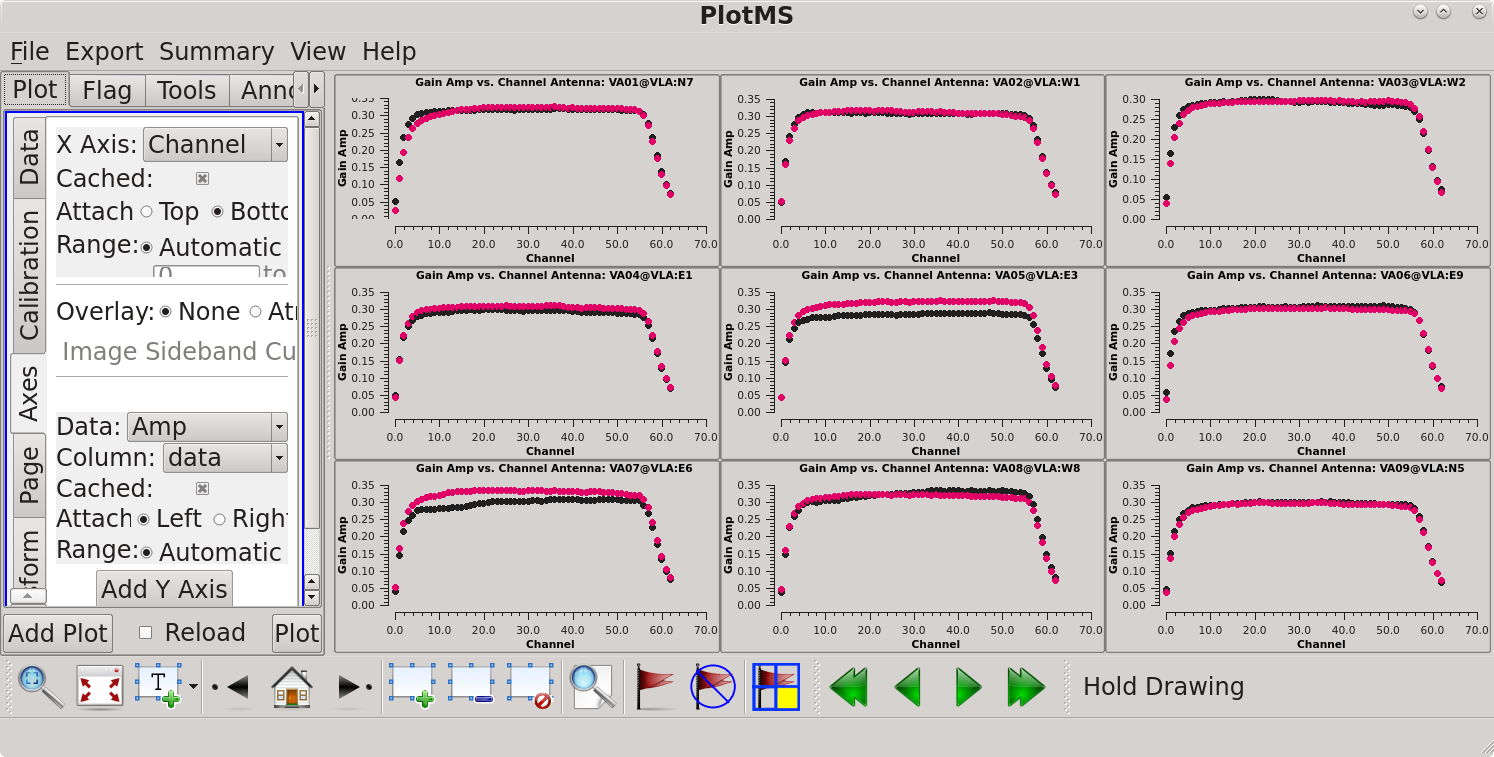
This creates a 3×33 grid showing the amplitude on each antenna, with 9 antennas per page. You can scroll through the antennas using the green arrows at the bottom of the plotms GUI. Note that antenna VA23 in position VLA:OUT is displayed blank (it was not in the array at the time) and antenna VA28 is shown on a fourth page. This is not a problem. Looking at the amplitudes confirms our earlier observation that channels 6 to 56 (spw='0:6~56') look good. We will use this spw setting again in the later calibration stages.
You may see the phases vs channel by changing the yaxis to phase interactively, or using the commands below. The phases should be relatively flat and continuous. Note that the phases do wander a bit at the edges of the bandpass where the amplitudes are lower. Some phases (e.g. on antenna VA10) wrap around between -180 and +180 – this is not a real discontinuity. VA15 has a very small variation in phases because it was used as the reference antenna.
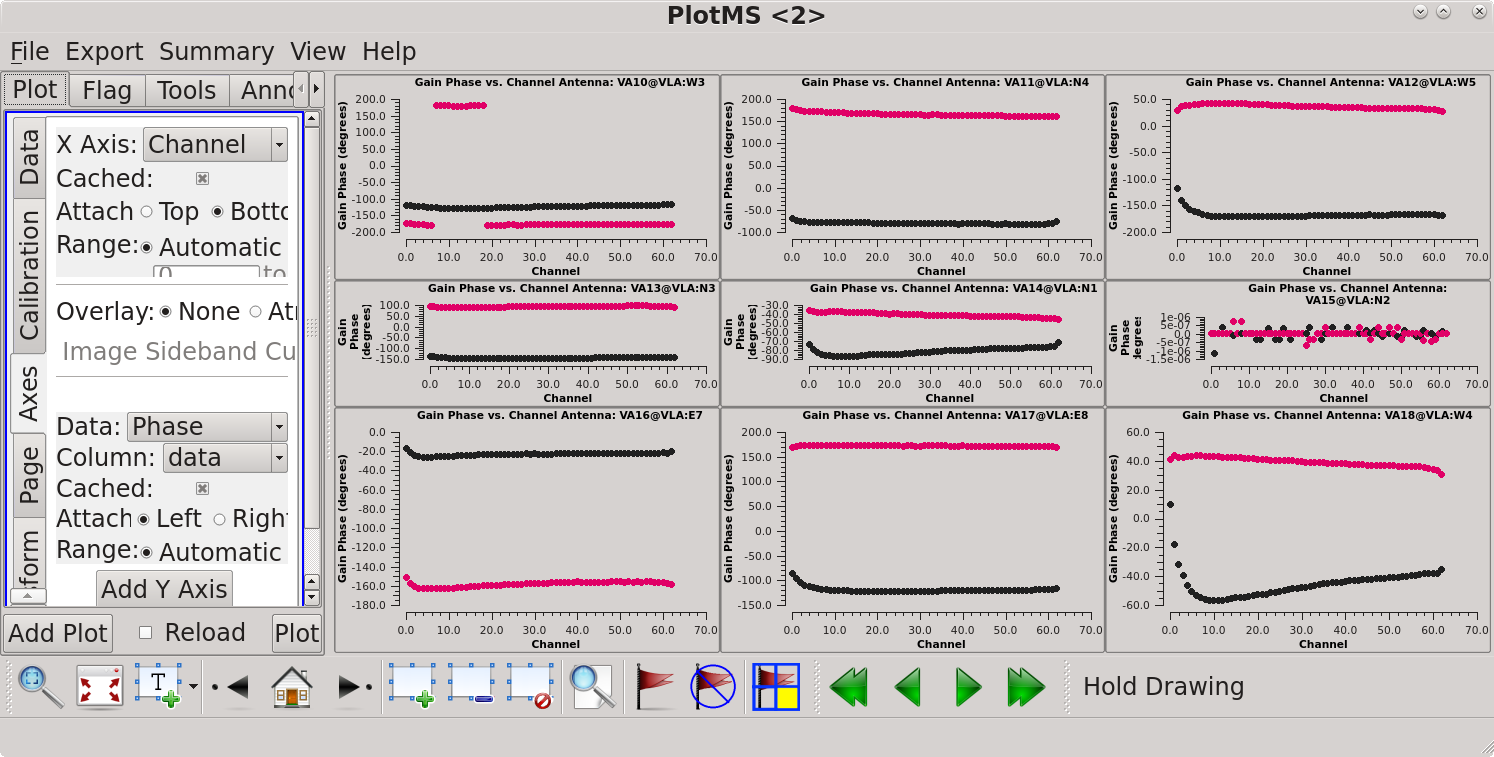
# Interactive CASA yaxis='phase' inp go
Note, this follows on from having set the other parameters in interactive CASA as in the previous box, just changing the y-axis to phase.
# Psuedo-interactive CASA
plotms(vis='bandpass.cal',xaxis='chan',yaxis='phase',coloraxis='corr',iteraxis='antenna',gridrows=3,gridcols=3,xselfscale=True,yselfscale=True)
Gain calibration
From our inspection of the bandpass response curve, we can average channels 6 to 56 to produce continuum-like data for the calibrators. This averaging is specified through the spw (spectral window) parameter in the form spw = '0:6~56'. This specifies the spectral window (IF) to be used as 0 (there is only one spectral window in this measurement set, as seen in the listobs output) and channels 6~56 within that spectral window.
Gain calibrations are otherwise determined as for continuum data.
- gaincal is run only on the calibrators, 1331+305 (flux calibrator) and 1445+099 (phase calibrator).
- The default model for gain calibrations is a 1 Jy point source. The flux scale is overridden by setjy, which has been performed for the flux calibrator. We need to transfer that flux scale to the phase calibrator using fluxscale.
- Note that fluxscale determines the flux density of the phase calibrator and accordingly adjusts its model and calibration solutions. A report of the results are sent to the logger window.
- Unless you use parameter incremental=True while executing fluxscale (the default is False), the resulting fluxscale.cal table will replace the apcal.gcal table at this point. This is particularly important when it comes to applying the calibration tables to the data in the applycal stage.
Complex gain calibration
First, we will calibrate the amplitude and phases of the data using the flux/bandpass and complex gain calibrators. To do this we will use gaincal and the previously-made calibration tables. This will give us antenna-based solutions that will show variations across time.
# Interactive CASA default gaincal inp vis='ngc5921.demo.ms' caltable='apcal.gcal' field='0,1' spw='0:6~56' refant='VA15' calmode='ap' solint='inf' gaintable=['antgain.cal','bandpass.cal'] inp go
# Psuedo-interactive CASA
gaincal(vis='TDEM0025_HI.ms',caltable='apcal.gcal',field='0,1',spw='0:6~56',refant='VA15',calmode='ap',solint='inf',gaintable=['antgain.cal','bandpass.cal'])
Where:
vis='ngc5921.demo.ms' # the measurement set
caltable='apcal.gcal' # the name you wish to give the calibration table
field='0,1' # the field ids of both calibrators
refant='VA15' # the reference antenna
calmode='ap' # to find both amplitude and phase solutions
solint='inf' # the solution interval time
gaintable=['antgain.cal','bandpass.cal'] # existing gain calibration table(s) to apply on the fly
Now look at solutions. First we will look at the phase solutions. These should be at zero with no phase jumps; phase jumps are when the phases change drastically with time. When this happens we are not able to interpolate the phases surrounding that scan that has the phase jump.
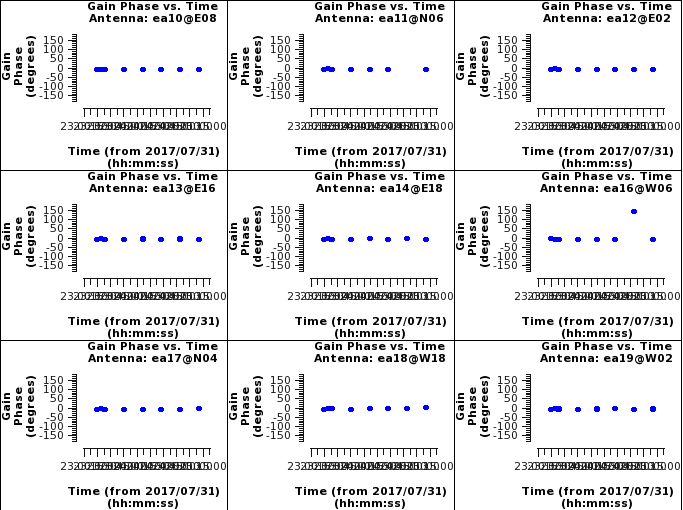
# Interactive CASA default plotms inp vis='apcal.gcal' xaxis='time' yaxis='phase' iteraxis='antenna' coloraxis='corr' gridrows=3 gridcols=3 plotrange=[0,0,-180,180] inp go
# Psuedo-interactive CASA
plotms(vis='apcal.gcal',xaxis='time',yaxis='phase',iteraxis='antenna',gridrows=3,gridcols=3,plotrange=[0,0,-180,180])
Where:
vis='apcal.gcal' # the complex gain calibration table
xaxis='time' # plot channels across the x axis
yaxis='phase' # plot amplitudes across the y axis
iteraxis='antenna' # pages the plots by antenna
coloraxis='corr' # colorizes the plots by correlation (RR and LL)
gridrows=3 # places 3 antennas per row
gridcols=3 # places 3 antennas per column
plotrange=[0,0,-180,180] # sets the plot range
Now look at the amplitudes to see if there are any irregularities there.
# Interactive CASA default plotms inp vis='apcal.gcal' xaxis='time' yaxis='amp' iteraxis='antenna' coloraxis='corr' gridrows=3 gridcols=3 xselfscale=True yselfscale=True inp go
# Psuedo-interactive CASA
plotms(vis='apcal.gcal',xaxis='time',yaxis='amp',iteraxis='antenna',gridrows=3,gridcols=3,xselfscale=True,yselfscale=True)
Where:
vis='apcal.gcal' # the complex gain calibration table
xaxis='time' # plot channels across the x axis
yaxis='amp' # plot amplitudes across the y axis
iteraxis='antenna' # pages the plots by antenna
coloraxis='corr' # colorizes the plots by correlation (RR and LL)
gridrows=3 # places 3 antennas per row
gridcols=3 # places 3 antennas per column
xselfscale=True # create a global xaxis scale for all antennas
yselfscale=True # create a global yaxis scale for all antennas
Don't worry about the shift in amplitude between the flux calibrator and the phase calibrator; this is because we haven't yet transferred the flux scale to the phase calibrator. What we're looking for here is whether there are large variations on the individual calibrators.
Next we use the gain calibration derived above and the known flux of the flux calibrator to transfer the flux scale to the phase calibrator using the fluxscale task.
# Interactive CASA default fluxscale inp vis='ngc5921.demo.ms' caltable='apcal.gcal' reference='0' fluxtable='fluxscale.cal' incremental=False inp go
# Psuedo-interactive CASA
fluxscale(vis='ngc5921.demo.ms',caltable='apcal.gcal',reference='0',fluxtable='fluxscale.cal',incremental=False)
Where:
vis='ngc5921.demo.ms' # the measurement set
caltable='apcal.gcal' # the calibration table with the amplitude and phase corrections
reference='0' # the field ID of the reference field (i.e. the flux calibrator)
fluxtable='fluxscale.cal' # the name of the output, flux-scaled calibration table
incremental=False # create a single fluxscale table with all the gain calibration
# ('False' is the default; if you use 'True' you need to supply
# both the apcal.gcal and fluxscale.cal tables to applycal below)
The output from fluxscale follows. A relatively large uncertainty for the phase calibrator is a sign that something went wrong, such as having bad solutions in gaincal. Here, the phase calibrator scaled to 2.521 ± 0.002 Jy, which looks reasonable.
2023-01-17 22:47:15 INFO fluxscale ########################################## 2023-01-17 22:47:15 INFO fluxscale ##### Begin Task: fluxscale ##### 2023-01-17 22:47:15 INFO fluxscale fluxscale( vis='ngc5921.demo.ms', caltable='apcal.gcal', fluxtable='fluxscale.cal', reference=['0'], transfer=[], listfile='', append=False, refspwmap=[-1], gainthreshold=-1.0, antenna='', timerange='', scan='', incremental=False, fitorder=1, display=False ) 2023-01-17 22:47:15 INFO fluxscale ****Using NEW VI2-driven calibrater tool**** 2023-01-17 22:47:15 INFO fluxscale Opening MS: ngc5921.demo.ms for calibration. 2023-01-17 22:47:15 INFO fluxscale Initializing nominal selection to the whole MS. 2023-01-17 22:47:15 INFO fluxscale Beginning fluxscale--(MSSelection version)------- 2023-01-17 22:47:16 INFO fluxscale Assuming all non-reference fields are transfer fields. 2023-01-17 22:47:16 INFO fluxscale Found reference field(s): 1331+30500002_0 2023-01-17 22:47:16 INFO fluxscale Found transfer field(s): 1445+09900002_0 2023-01-17 22:47:16 INFO fluxscale Flux density for 1445+09900002_0 in SpW=0 (freq=1.41342e+09 Hz) is: 2.52141 +/- 0.00223811 (SNR = 1126.58, N = 54) 2023-01-17 22:47:16 INFO fluxscale Storing result in fluxscale.cal 2023-01-17 22:47:16 INFO fluxscale Writing solutions to table: fluxscale.cal 2023-01-17 22:47:16 INFO fluxscale Task fluxscale complete. Start time: 2023-01-17 15:47:15.041544 End time: 2023-01-17 15:47:16.120049 2023-01-17 22:47:16 INFO fluxscale ##### End Task: fluxscale ##### 2023-01-17 22:47:16 INFO fluxscale ##########################################
Now we can look at the amplitudes again, using the transferred flux scale which should eliminate the differences between the calibrators. This is identical to the previous inspection, but using fluxscale.cal rather than apcal.gcal (so could be run in interactive CASA by retrieving the previous inputs with "tget plotms" then just changing the input table using "vis='fluxscale.cal'").
# Interactive CASA default plotms inp vis='fluxscale.cal' xaxis='time' yaxis='amp' iteraxis='antenna' coloraxis='corr' gridrows=3 gridcols=3 xselfscale=True yselfscale=True inp go
# Psuedo-interactive CASA
plotms(vis='fluxscale.cal',xaxis='time',yaxis='amp',iteraxis='antenna',gridrows=3,gridcols=3,xselfscale=True,yselfscale=True)
Where:
vis='fluxscale.cal' # the complex gain calibration table with the flux scale applied
xaxis='time' # plot channels across the x axis
yaxis='amp' # plot amplitudes across the y axis
iteraxis='antenna' # pages the plots by antenna
coloraxis='corr' # colorizes the plots by correlation (RR and LL)
gridrows=3 # places 3 antennas per row
gridcols=3 # places 3 antennas per column
xselfscale=True # create a global xaxis scale for all antennas
yselfscale=True # create a global yaxis scale for all antennas
Apply the solutions
Next, we apply the calibration solutions to the calibrators themselves, and finally transfer the calibration solutions by interpolation (or nearest-neighbor sampling) to the source. The relevant task is applycal, which fills out a new column (CORRECTED_DATA) of calibrated data in the measurement set without replacing the raw data column (DATA). The application is identical to that used for continuum data, except that the bandpass table is also included in the calibration. In order to apply multiple calibrations at once, we provide the gaintable parameter with a list of calibration tables rather than a single entry. If we had run fluxscale with "incremental=True", we would also include the apcal.gcal table here.
First, apply the tables to the flux calibrator:
# Interactive CASA default applycal inp vis='ngc5921.demo.ms' field='0' gaintable=['antgain.cal', 'bandpass.cal', 'fluxscale.cal'] gainfield=['','0','0'] calwt=False flagbackup=True inp go
# Psuedo-interactive CASA
# for the flux/bandpass calibrator (field '0')
applycal(vis='ngc5921.demo.ms', field='0',
gaintable=['antgain.cal', 'bandpass.cal', 'fluxscale.cal'],
gainfield=['','0','0'],
calwt=False, flagbackup=True)
Where:
vis='ngc5921.demo.ms' # the measurement set
field='0' # field id(s) to work on - 0 = the flux calibrator
gaintable=['antgain.cal''bandpass.cal','fluxscale.cal'] # gain calibration table(s) to apply
gainfield=['','0','0'] # field id(s) from which each respective gaintable was derived
calwt=False # determines if the weights are calibrated along with the data
flagbackup=True # automatically back up the state of flags before the run
Within the gainfield parameter, we do not specify a field for the antgain.cal table since this was derived independently from the calibrator fields. For the calibrators, all bandpass solutions are the same as for the flux/bandpass calibrator (id=0), and the phase and amplitude calibration as derived from their own visibility data. So when applying corrections to any of our sources, the respective field for bandpass.cal will be our bandpass calibrator (field='0'), but the field for fluxscale.cal will be '1' for both the phase calibrator itself and the target field.
Next, apply the tables to the phase calibrator:
# Interactive CASA default applycal inp vis='ngc5921.demo.ms' field='1' gaintable=['antgain.cal', 'bandpass.cal', 'fluxscale.cal'] gainfield=['','0','1'] calwt=False flagbackup=True inp go
(Note: the only changes from the application to the flux calibrator are the field to apply the calibration to and the gainfield for the fluxscale.cal table, both changed to refer to the phase calibrator.)
# Psuedo-interactive CASA
# for the phase calibrator (field '1')
applycal(vis='ngc5921.demo.ms', field='1',
gaintable=['antgain.cal', 'bandpass.cal', 'fluxscale.cal'],
gainfield=['','0','1'],
calwt=False, flagbackup=True)
Where:
vis='ngc5921.demo.ms' # the measurement set
field='1' # field id(s) to work on - 1 = the flux calibrator
gaintable=['antgain.cal''bandpass.cal','fluxscale.cal'] # gain calibration table(s) to apply
gainfield=['','0','1'] # field id(s) from which each respective gaintable was derived
calwt=False # determines if the weights are calibrated along with the data
flagbackup=True # automatically back up the state of flags before the run
Finally, apply the tables to the science target:
# Interactive CASA default applycal inp vis='ngc5921.demo.ms' field='2' gaintable=['antgain.cal', 'bandpass.cal', 'fluxscale.cal'] gainfield=['','0','1'] calwt=False flagbackup=True inp go
(Note: the only change from the phase calibrator is the field to apply the calibration to – the gain fields are all the same.)
# Psuedo-interactive CASA
# for the science target (field '2')
applycal(vis='ngc5921.demo.ms', field='2',
gaintable=['antgain.cal', 'bandpass.cal', 'fluxscale.cal'],
gainfield=['','0','1'],
calwt=False, flagbackup=True)
Where:
vis='ngc5921.demo.ms' # the measurement set
field='2' # field id(s) to work on - 2 = the science target
gaintable=['antgain.cal''bandpass.cal','fluxscale.cal'] # gain calibration table(s) to apply
gainfield=['','0','1'] # field id(s) from which each respective gaintable was derived
calwt=False # determines if the weights are calibrated along with the data
flagbackup=True # automatically back up the state of flags before the run
Plot the spectrum
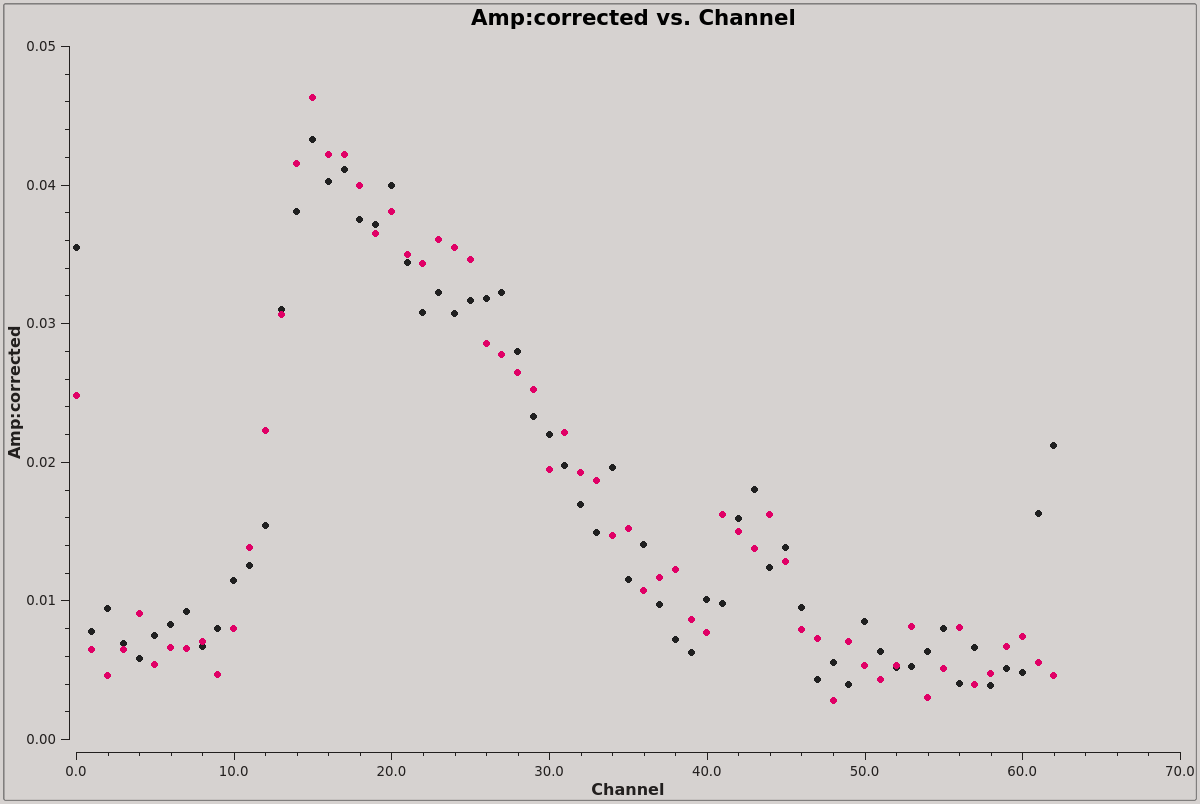
Before we attempt to image the 21 cm cube of the source, we need to subtract off the underlying continuum, which means we need to plot the integrated spectrum of the source to determine the line-free channels where we can fit the continuum.
We can do this in plotms.
# Interactive CASA default plotms inp vis='ngc5921.demo.ms' field='2' spw='0:6~56' averagedata=True avgtime='3600' avgscan=True avgbaseline=True xaxis='channel' yaxis='amp' coloraxis='corr' ydatacolumn='corrected' inp go
# Psuedo-interactive CASA
plotms(vis='ngc5921.demo.ms', field='2', spw='0:6~56', averagedata=True, avgtime='3600', avgscan=True, avgbaseline=True, xaxis='channel', yaxis='amp', ydatacolumn='corrected')
Where:
vis='ngc5921.demo.ms' # the measurement set
field='2' # select the science target
averagedata=True # average the data (enables data-averaging options)
avgtime='3600' # average over 3600s (i.e. the whole dataset)
avgscan=True # average across scan boundaries
avgbaseline=True # average baselines
xaxis='channel' # plot channels across the x axis
yaxis='amp' # plot amplitudes across the y axis
coloraxis='corr' # colorizes the plots by correlation (RR and LL)
ydatacolumn='corrected' # use the CORRECTED_DATA column (with the calibration applied)
The resulting plot is illustrated in the figure at right. Briefly, we want to average both in time and over baselines to get the signal-to-noise necessary to reveal the 21 cm profile (see Averaging data in plotms for more details on averaging options). If the symbols appear too small, the size may be increased via the Display tab on the GUI: change the Unflagged Points Symbol to 'Custom', change the plotting symbol to 'circle', 'square' or 'diamond', and increase the number of pixels for the plotting symbol.
Note that we have entered all the relevent parameters via the command line interface rather than selecting options in the GUI. If you wish to enter the values directly into the GUI, you can follow the (Tab)Command convention of the flagging tutorial with the following settings :
- (Data)field = N5921*
- (Data)Averaging → Time = 3600 (average over some long time window)
- (Data)Averaging → Scan = True (checkmark; average in time across scan boundaries)
- (Data)Averaging → All Baselines = True (checkmark)
- (Axes)X Axis = Channel
- (Axes)Y Axis = Amp
- (Display)Colorize = Corr
From inspection of this plot, it looks like channels 4~8 and 50~59 contain line-free channels, suitable to use for continuum subtraction.
Continuum subtraction
The next step is to split off the NGC 5921 data from the multisource measurement set and subtract the continuum. Splitting uses the split command, as follows.
# Interactive CASA default split inp vis='ngc5921.demo.ms' field='2' outputvis='ngc5921.demo.split.ms' spw='' datacolumn='corrected' inp go
# Psuedo-interactive CASA
plotms(vis='ngc5921.demo.ms', outputvis='ngc5921.demo.split.ms', field='2', spw='', datacolumn='corrected')
Where:
vis='ngc5921.demo.ms' # the input measurement set
outputvis='ngc5921.demo.split.ms # the output split measurement set
field='2' # select the science target
spw='' # select the entire spectral window
datacolumn='corrected' # use the CORRECTED_DATA column (with the calibration applied)
This creates a new measurement set called ngc5921.demo.split.ms and copies the calibrated source data (datacolumn = 'corrected') from the CORRECTED_DATA column into it, writing it to the DATA column in the output measurement set.
We can now subtract the continuum from this split dataset in the visibility (uv) plane using uvcontsub. As we determined above, we can use channels 4~8 and 50~59 for this. Note that the uvcontsub task was substantially revised in CASA 6.5 and now has new parameter inputs that are not backwards compatible with inputs based on older CASAguides.
# Interactive CASA default uvcontsub ins vis='ngc5921.demo.split.ms' outputvis='ngc5921.demo.uvcontsub.ms' fitspec='0:4~8;50~59' fitorder=0 inp go
# Psuedo-interactive CASA
plotms(vis='ngc5921.demo.ms', outputvis='ngc5921.demo.split.ms', fitspec='0:4~8;50~59', fitorder=0)
Where:
vis='ngc5921.demo.split ms' # the input split measurement set
outputvis='ngc5921.demo.uvcontsub.ms # the output continuum-subtracted measurement set
fitspec='0:4~8;50~59' # select the channels over which to fit the continuum
fitorder=0 # fit with polynomial order 0 (constant)
(Note: this could be run directly on the original measurement set by specifying vis='ngc5921.demo.ms', field='2', and datacolumn='corrected' to select the science target and the calibrated data column – in a change from its earlier behavior, datacolumn='data' is the default rather than the fallback for uvcontsub, so this must be specified explicitly if not using split data)
The fitspec parameter, if given as a simple string as done here, specifies the spw to be used for the fitting, with the fit order given using fitorder. However, fitspect can be used in more complex ways by specifying it as a python dictionary. For details, see the uvcontsub documentation.
The polynomial fit order specified is used to fit the real and imaginary components separately, so the fitted model amplitude is the quadrature combination of these two fits and is thus not, in general, a polynomial.
Imaging
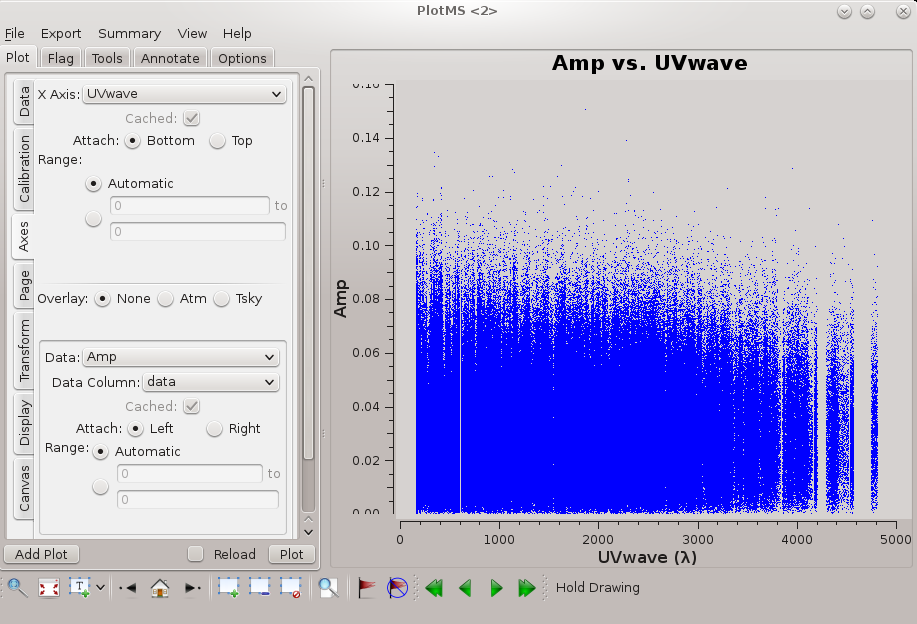
Now we can use tclean to generate the primary science product, a cleaned data cube (ra, dec, velocity) from the continuum-subtracted (u, v, channel) measurement set, ngc5921.demo.uvcontsub.ms. Things to consider in using tclean:
- To ensure channels aren't averaged prior to imaging and proper Doppler combination, choose specmode='cube'.
- The maximum baseline is just under 5 kλ (see the figure at right, showing this plotted with plotms), and so the expected synthetic beam is roughly 1.22 × 206265 / 5000 = 50 arcseconds (subject to the details of u, v weighting), as expected for D configuration. Pixels should sample the beam better than 3 times, so 15 arcseconds is a good choice of pixel size (cell = ['15.0arcsec','15.0arcsec']).
- We only want to clean down to the noise, which can be determined by examining line-free channels in an image made without cleaning (a 'dirty' image), as detailed below.
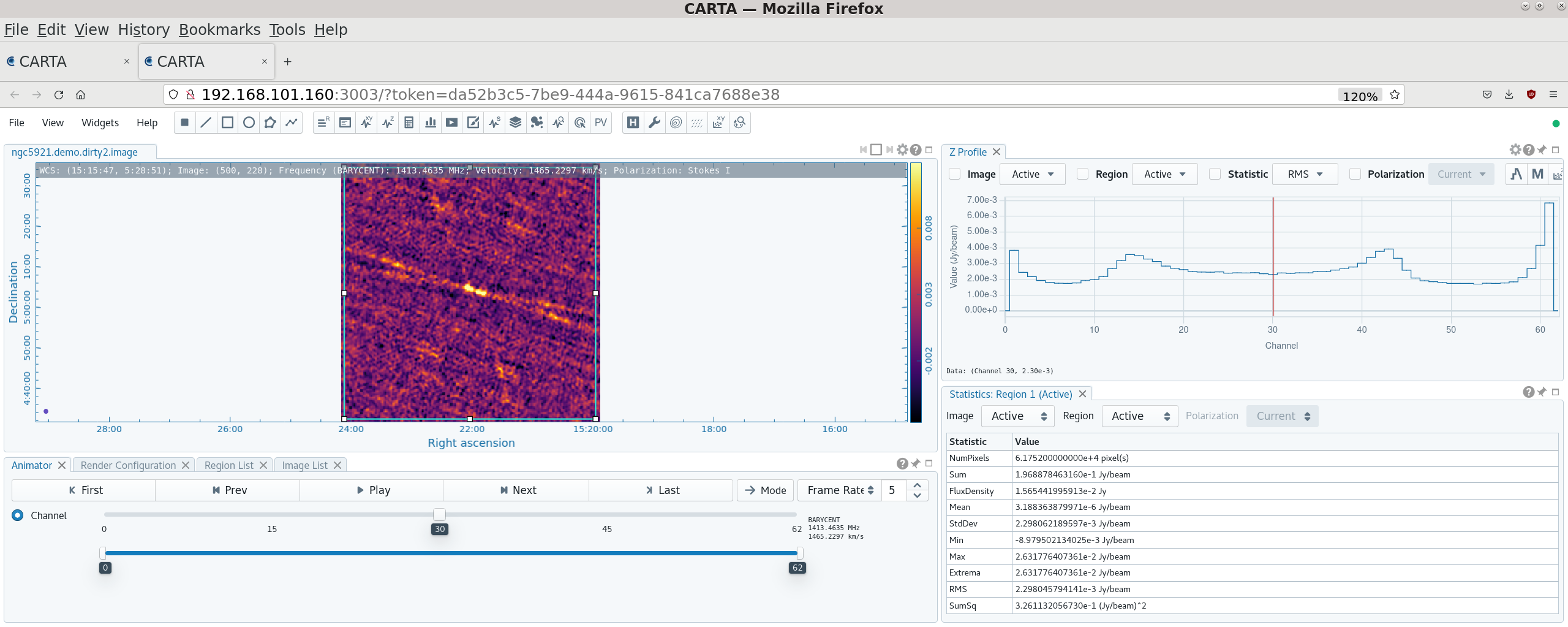
To create a dirty image:
# Interactive CASA default tclean inp vis='ngc5921.demo.uvcontsub.ms' specmode=cube inp #Run inp again, as setting specmode=cube opens new options cell=['15.0arcsec','15.0arcsec'] outframe='bary' niter = 0 imagename='ngc5921.demo.dirty' datacolumn='data' imsize=[256,256] pblimit=-0.2 weighting='briggs' robust=0.5 inp go
# Pseudo-interactive CASA
tclean(vis='ngc5921.demo.uvcontsub.ms', imagename='ngc5921.demo.dirty', specmode='cube', niter=0, imsize=[256,256], cell=['15.0arcsec','15.0arcsec'], outframe='baby', weighting='briggs', robust=0.5, interactive=False, pblimit=-0.2)
Where: vis='ngc5921.demo.uvcontsub.ms'. # the input measurement set specmode=cube # sets the spectral mode to properly handle line data cell=['15.0arcsec','15.0arcsec'] # sets the cell (pixel) size outframe='bary' # sets the velocity frame to the extragalactic standard of barycentric niter = 0 # sets the number of clean cycles to 0 (i.e. no cleaning) imagename='ngc5921.demo.dirty' # the output image name datacolumn='data' # tells tclean to use the DATA column imsize=[256,256] # sets the image size (in pixels) pblimit=-0.2 # negative value to not blank pixels beyond a primary beam gain of 20% weighting='briggs' # use Briggs weighting... robust=0.5 # ...with a robust value of 0.5
The output ngc5921.demo.dirty.image file (note that .image gets appended – other outputs such as the .psf, .pb, etc. are also created by tclean) can then be viewed using CARTA. By defining a rectangular region that takes in the whole image plane and setting the Z profile to 'RMS', it is possible to see how the RMS level varies by channel. You can see how the noise falls from the edge of the bandpass to around 1.75 mJy by channel 6, then rises again from channel 9 when the source starts to contribute, then falls back to 1.75 mJy again by channel 50, rising again from channel 56. This also confirms our earlier choices of which channels looked line-free and which channels were in the best signal-to-noise section of the cube. If we thus want to clean to 4 sigma, we can set threshold='7.0mJy'.
The dirty image can also be used to determine the region to be cleaned by tclean. This is most easily done by defining a region that encloses the area where flux is visible on the dirty image (which can be checked by stepping through the channels) and then exporting this region as a CRTF file, here cleanbox.crtf, that can be read by CASA.
We are now ready to run tclean again and create the clean cube.
# Interactive CASA default tclean inp vis='ngc5921.demo.uvcontsub.ms' specmode=cube cell=['15.0arcsec','15.0arcsec'] outframe='bary' datacolumn='data' imsize=[256,256] pblimit=-0.2 weighting='briggs' robust=0.5 imagename='ngc5921.demo.clean' niter=6000 mask='cleanbox.crtf' threshold='7.0mJy' inp go
(Note: only the last four parameters, imagename, niter, mask and threshold have changed)
# Pseudo-interactive CASA
tclean(vis='ngc5921.demo.uvcontsub.ms', imagename='ngc5921.demo.clean', specmode='cube', niter=6000, threshold='7.0mJy', imsize=[256,256], cell=['15.0arcsec','15.0arcsec'], outframe='baby', weighting='briggs', robust=0.5, mask = 'clean box.crtf', interactive=False, pblimit=-0.2)
Where: vis='ngc5921.demo.uvcontsub.ms'. # the input measurement set specmode=cube # sets the spectral mode to properly handle line data cell=['15.0arcsec','15.0arcsec'] # sets the cell (pixel) size outframe='bary' # sets the velocity frame to the extragalactic standard of barycentric datacolumn='data' # tells tclean to use the DATA column imsize=[256,256] # sets the image size (in pixels) pblimit=-0.2 # negative value to not blank pixels beyond a primary beam gain of 20% weighting='briggs' # use Briggs weighting... robust=0.5 # ...with a robust value of 0.5 imagename='ngc5921.demo.clean' # the output image name niter=6000 # sets the number of clean cycles to 6000 (will stop at threshold value) mask='cleanbox.crtf' # use the mask defined in the cleanbox.crtf file created by CARTA threshold='7.0mJy' # stop cleaning at this flux level
Inspecting the data cube
We can use imhead to examine the cube header:
# Interactive CASA default imhead inp imagename='ngc5921.demo.clean.image' inp go
# Pseudo-interactive CASA
imhead(imagename='ngc5921.demo.clean.image', mode='summary')
The output, as follows, appears in the logger window:
2023-01-18 21:15:06 INFO imhead ########################################## 2023-01-18 21:15:06 INFO imhead ##### Begin Task: imhead ##### 2023-01-18 21:15:06 INFO imhead imhead( imagename='ngc5921.demo.clean.image', mode='summary', hdkey='', hdvalue='', verbose=False ) 2023-01-18 21:15:08 INFO ImageMetaData 2023-01-18 21:15:08 INFO ImageMetaData Image name : ngc5921.demo.clean.image 2023-01-18 21:15:08 INFO ImageMetaData Object name : N5921_2 2023-01-18 21:15:08 INFO ImageMetaData Image type : PagedImage 2023-01-18 21:15:08 INFO ImageMetaData Image quantity : Intensity 2023-01-18 21:15:08 INFO ImageMetaData Pixel mask(s) : None 2023-01-18 21:15:08 INFO ImageMetaData Region(s) : None 2023-01-18 21:15:08 INFO ImageMetaData Image units : Jy/beam 2023-01-18 21:15:08 INFO ImageMetaData Restoring Beams 2023-01-18 21:15:08 INFO ImageMetaData Pol Type Chan Freq Vel 2023-01-18 21:15:08 INFO ImageMetaData I Max 0 1.41273e+09 1619.82 53.0827 arcsec x 47.0911 arcsec pa= 7.9378 deg 2023-01-18 21:15:08 INFO ImageMetaData I Min 48 1.41390e+09 1372.47 53.0292 arcsec x 46.9723 arcsec pa= 9.2164 deg 2023-01-18 21:15:08 INFO ImageMetaData I Median 31 1.41349e+09 1460.08 53.1100 arcsec x 47.0231 arcsec pa= 8.0658 deg 2023-01-18 21:15:08 INFO ImageMetaData 2023-01-18 21:15:08 INFO ImageMetaData Direction reference : J2000 2023-01-18 21:15:08 INFO ImageMetaData Spectral reference : BARY 2023-01-18 21:15:08 INFO ImageMetaData Velocity type : RADIO 2023-01-18 21:15:08 INFO ImageMetaData Rest frequency : 1.42041e+09 Hz 2023-01-18 21:15:08 INFO ImageMetaData Pointing center : 15:22:00.000000 +05.04.00.000000 2023-01-18 21:15:08 INFO ImageMetaData Telescope : VLA 2023-01-18 21:15:08 INFO ImageMetaData Observer : TEST 2023-01-18 21:15:08 INFO ImageMetaData Date observation : 1995/04/13/09:33:00.000687 2023-01-18 21:15:08 INFO ImageMetaData Telescope position: [-1.60119e+06m, -5.04198e+06m, 3.55488e+06m] (ITRF) 2023-01-18 21:15:08 INFO ImageMetaData 2023-01-18 21:15:08 INFO ImageMetaData Axis Coord Type Name Proj Shape Tile Coord value at pixel Coord incr Units 2023-01-18 21:15:08 INFO ImageMetaData ------------------------------------------------------------------------------------------------ 2023-01-18 21:15:08 INFO ImageMetaData 0 0 Direction Right Ascension SIN 256 64 15:22:00.000 128.00 -1.500000e+01 arcsec 2023-01-18 21:15:08 INFO ImageMetaData 1 0 Direction Declination SIN 256 64 +05.04.00.000 128.00 1.500000e+01 arcsec 2023-01-18 21:15:08 INFO ImageMetaData 2 1 Stokes Stokes 1 1 I 2023-01-18 21:15:08 INFO ImageMetaData 3 2 Spectral Frequency 63 9 1.41273e+09 0.00 2.4415203e+04 Hz 2023-01-18 21:15:08 INFO ImageMetaData Velocity 1619.82 0.00 -5.153101e+00 km/s 2023-01-18 21:15:08 INFO imhead Task imhead complete. Start time: 2023-01-18 14:15:06.134860 End time: 2023-01-18 14:15:07.997282 2023-01-18 21:15:08 INFO imhead ##### End Task: imhead ##### 2023-01-18 21:15:08 INFO imhead ##########################################
Cube statistics
We can use imstat to display statistics of the cube cubes. The following example displays the statistics for an empty region of the cube (as identified visually in CARTA).
#Interactive CASA default instate inp imagename = 'ngc5921.demo.clean.image' box = '10,10,100,100' inp go
As with imhead, the output goes to the logger window:
2023-01-18 21:42:36 INFO imstat ########################################## 2023-01-18 21:42:36 INFO imstat ##### Begin Task: imstat ##### 2023-01-18 21:42:36 INFO imstat imstat( imagename='ngc5921.demo.clean.image', axes=[], region='', box='10,10,100,100', chans='', stokes='', listit=True, verbose=True, mask='', stretch=False, logfile='', append=True, algorithm='classic', fence=-1.0, center='mean', lside=True, zscore=-1.0, maxiter=-1, clmethod='auto', niter=3 ) 2023-01-18 21:42:36 INFO imstat Using specified box(es) 10,10,100,100 2023-01-18 21:42:36 INFO imstat Determining stats for image ngc5921.demo.clean.image 2023-01-18 21:42:36 INFO imstat Selected bounding box : 2023-01-18 21:42:36 INFO imstat [10, 10, 0, 0] to [100, 100, 0, 62] (15:23:58.379, +04.34.29.305, I, 1.41273e+09Hz to 15:22:28.105, +04.56.59.962, I, 1.41424e+09Hz) 2023-01-18 21:42:36 INFO imstat Statistics calculated using Classic algorithm 2023-01-18 21:42:36 INFO imstat Regions --- 2023-01-18 21:42:36 INFO imstat -- bottom-left corner (pixel) [blc]: [10, 10, 0, 0] 2023-01-18 21:42:36 INFO imstat -- top-right corner (pixel) [trc]: [100, 100, 0, 62] 2023-01-18 21:42:36 INFO imstat -- bottom-left corner (world) [blcf]: 15:23:58.379, +04.34.29.305, I, 1.41273e+09Hz 2023-01-18 21:42:36 INFO imstat -- top-right corner (world) [trcf]: 15:22:28.105, +04.56.59.962, I, 1.41424e+09Hz 2023-01-18 21:42:36 INFO imstat Values --- 2023-01-18 21:42:36 INFO imstat -- flux [flux]: 1.41948 Jy.km/s 2023-01-18 21:42:36 INFO imstat -- number of points [npts]: 521703 2023-01-18 21:42:36 INFO imstat -- maximum value [max]: 0.0230701 Jy/beam 2023-01-18 21:42:36 INFO imstat -- minimum value [min]: -0.0222025 Jy/beam 2023-01-18 21:42:36 INFO imstat -- position of max value (pixel) [maxpos]: [20, 44, 0, 61] 2023-01-18 21:42:36 INFO imstat -- position of min value (pixel) [minpos]: [35, 70, 0, 61] 2023-01-18 21:42:36 INFO imstat -- position of max value (world) [maxposf]: 15:23:48.368, +04.42.59.428, I, 1.41422e+09Hz 2023-01-18 21:42:36 INFO imstat -- position of min value (world) [minposf]: 15:23:33.331, +04.49.29.579, I, 1.41422e+09Hz 2023-01-18 21:42:36 INFO imstat -- Sum of pixel values [sum]: 3.46393 Jy/beam 2023-01-18 21:42:36 INFO imstat -- Sum of squared pixel values [sumsq]: 2.32674 Jy/beam.Jy/beam 2023-01-18 21:42:36 INFO imstat Statistics --- 2023-01-18 21:42:36 INFO imstat -- Mean of the pixel values [mean]: 6.63965e-06 Jy/beam 2023-01-18 21:42:36 INFO imstat -- Variance of the pixel values : 4.45985e-06 Jy/beam 2023-01-18 21:42:36 INFO imstat -- Standard deviation of the Mean [sigma]: 0.00211184 Jy/beam 2023-01-18 21:42:36 INFO imstat -- Root mean square [rms]: 0.00211184 Jy/beam 2023-01-18 21:42:36 INFO imstat -- Median of the pixel values [median]: 0 Jy/beam 2023-01-18 21:42:36 INFO imstat -- Median of the deviations [medabsdevmed]: 0.00126323 Jy/beam 2023-01-18 21:42:36 INFO imstat -- IQR [quartile]: 0.00252672 Jy/beam 2023-01-18 21:42:36 INFO imstat -- First quartile [q1]: -0.00125785 Jy/beam 2023-01-18 21:42:36 INFO imstat -- Third quartile [q3]: 0.00126887 Jy/beam 2023-01-18 21:42:36 INFO imstat Sum column unit = Jy/beam 2023-01-18 21:42:36 INFO imstat Mean column unit = Jy/beam 2023-01-18 21:42:36 INFO imstat Std_dev column unit = Jy/beam 2023-01-18 21:42:36 INFO imstat Minimum column unit = Jy/beam 2023-01-18 21:42:36 INFO imstat Maximum column unit = Jy/beam 2023-01-18 21:42:36 INFO imstat Npts Sum Mean Rms Std_dev Minimum Maximum 2023-01-18 21:42:36 INFO imstat 5.217030e+05 3.463928e+00 6.639655e-06 2.111845e-03 2.111836e-03 -2.220254e-02 2.307010e-02 2023-01-18 21:42:36 INFO imstat Task imstat complete. Start time: 2023-01-18 14:42:36.025027 End time: 2023-01-18 14:42:36.481128 2023-01-18 21:42:36 INFO imstat ##### End Task: imstat ##### 2023-01-18 21:42:36 INFO imstat ##########################################
Alternatively, we can use the cleanbox.crtf file created earlier to examine the stats in the region containing the source:
#Interactive CASA default instate inp imagename = 'ngc5921.demo.clean.image' box = '' region='cleanbox.crtf' inp go
Which gives the output:
2023-01-18 21:44:58 INFO imstat ########################################## 2023-01-18 21:44:58 INFO imstat ##### Begin Task: imstat ##### 2023-01-18 21:44:58 INFO imstat imstat( imagename='ngc5921.demo.clean.image', axes=[], region='cleanbox.crtf', box='', chans='', stokes='', listit=True, verbose=True, mask='', stretch=False, logfile='', append=True, algorithm='classic', fence=-1.0, center='mean', lside=True, zscore=-1.0, maxiter=-1, clmethod='auto', niter=3 ) 2023-01-18 21:44:59 INFO imstat RegionTextParser::_determineVersion: Found spec version 0 2023-01-18 21:44:59 INFO imstat Combined 1 image regions (which excludes any annotation regions) 2023-01-18 21:44:59 INFO imstat The specified region will select all pixels that are included in the region. Full pixels will be included even when they are only partially covered by the region(s). 2023-01-18 21:44:59 INFO imstat Region read from CRTF file cleanbox.crtf 2023-01-18 21:44:59 INFO imstat Determining stats for image ngc5921.demo.clean.image 2023-01-18 21:44:59 INFO imstat Selected bounding box : 2023-01-18 21:44:59 INFO imstat [111, 107, 0, 0] to [150, 151, 0, 62] (15:22:17.064, +04.58.44.986, I, 1.41273e+09Hz to 15:21:37.910, +05.09.44.977, I, 1.41424e+09Hz) 2023-01-18 21:44:59 INFO imstat Statistics calculated using Classic algorithm 2023-01-18 21:44:59 INFO imstat Regions --- 2023-01-18 21:44:59 INFO imstat -- bottom-left corner (pixel) [blc]: [111, 107, 0, 0] 2023-01-18 21:44:59 INFO imstat -- top-right corner (pixel) [trc]: [150, 151, 0, 62] 2023-01-18 21:44:59 INFO imstat -- bottom-left corner (world) [blcf]: 15:22:17.064, +04.58.44.986, I, 1.41273e+09Hz 2023-01-18 21:44:59 INFO imstat -- top-right corner (world) [trcf]: 15:21:37.910, +05.09.44.977, I, 1.41424e+09Hz 2023-01-18 21:44:59 INFO imstat Values --- 2023-01-18 21:44:59 INFO imstat -- flux [flux]: 28.2043 Jy.km/s 2023-01-18 21:44:59 INFO imstat -- number of points [npts]: 113400 2023-01-18 21:44:59 INFO imstat -- maximum value [max]: 0.0559169 Jy/beam 2023-01-18 21:44:59 INFO imstat -- minimum value [min]: -0.0221281 Jy/beam 2023-01-18 21:44:59 INFO imstat -- position of max value (pixel) [maxpos]: [134, 134, 0, 43] 2023-01-18 21:44:59 INFO imstat -- position of min value (pixel) [minpos]: [145, 138, 0, 61] 2023-01-18 21:44:59 INFO imstat -- position of max value (world) [maxposf]: 15:21:53.976, +05.05.29.998, I, 1.41378e+09Hz 2023-01-18 21:44:59 INFO imstat -- position of min value (world) [minposf]: 15:21:42.932, +05.06.29.986, I, 1.41422e+09Hz 2023-01-18 21:44:59 INFO imstat -- Sum of pixel values [sum]: 68.8153 Jy/beam 2023-01-18 21:44:59 INFO imstat -- Sum of squared pixel values [sumsq]: 2.27782 Jy/beam.Jy/beam 2023-01-18 21:44:59 INFO imstat Statistics --- 2023-01-18 21:44:59 INFO imstat -- Mean of the pixel values [mean]: 0.000606837 Jy/beam 2023-01-18 21:44:59 INFO imstat -- Variance of the pixel values : 1.97185e-05 Jy/beam 2023-01-18 21:44:59 INFO imstat -- Standard deviation of the Mean [sigma]: 0.00444055 Jy/beam 2023-01-18 21:44:59 INFO imstat -- Root mean square [rms]: 0.0044818 Jy/beam 2023-01-18 21:44:59 INFO imstat -- Median of the pixel values [median]: 0 Jy/beam 2023-01-18 21:44:59 INFO imstat -- Median of the deviations [medabsdevmed]: 0.00138569 Jy/beam 2023-01-18 21:44:59 INFO imstat -- IQR [quartile]: 0.00277213 Jy/beam 2023-01-18 21:44:59 INFO imstat -- First quartile [q1]: -0.001358 Jy/beam 2023-01-18 21:44:59 INFO imstat -- Third quartile [q3]: 0.00141413 Jy/beam 2023-01-18 21:44:59 INFO imstat Sum column unit = Jy/beam 2023-01-18 21:44:59 INFO imstat Mean column unit = Jy/beam 2023-01-18 21:44:59 INFO imstat Std_dev column unit = Jy/beam 2023-01-18 21:44:59 INFO imstat Minimum column unit = Jy/beam 2023-01-18 21:44:59 INFO imstat Maximum column unit = Jy/beam 2023-01-18 21:44:59 INFO imstat Npts Sum Mean Rms Std_dev Minimum Maximum 2023-01-18 21:44:59 INFO imstat 1.134000e+05 6.881528e+01 6.068366e-04 4.481805e-03 4.440551e-03 -2.212812e-02 5.591685e-02 2023-01-18 21:44:59 INFO imstat Task imstat complete. Start time: 2023-01-18 14:44:58.195518 End time: 2023-01-18 14:44:59.067297 2023-01-18 21:44:59 INFO imstat ##### End Task: imstat ##### 2023-01-18 21:44:59 INFO imstat ##########################################
The integrated spectrum
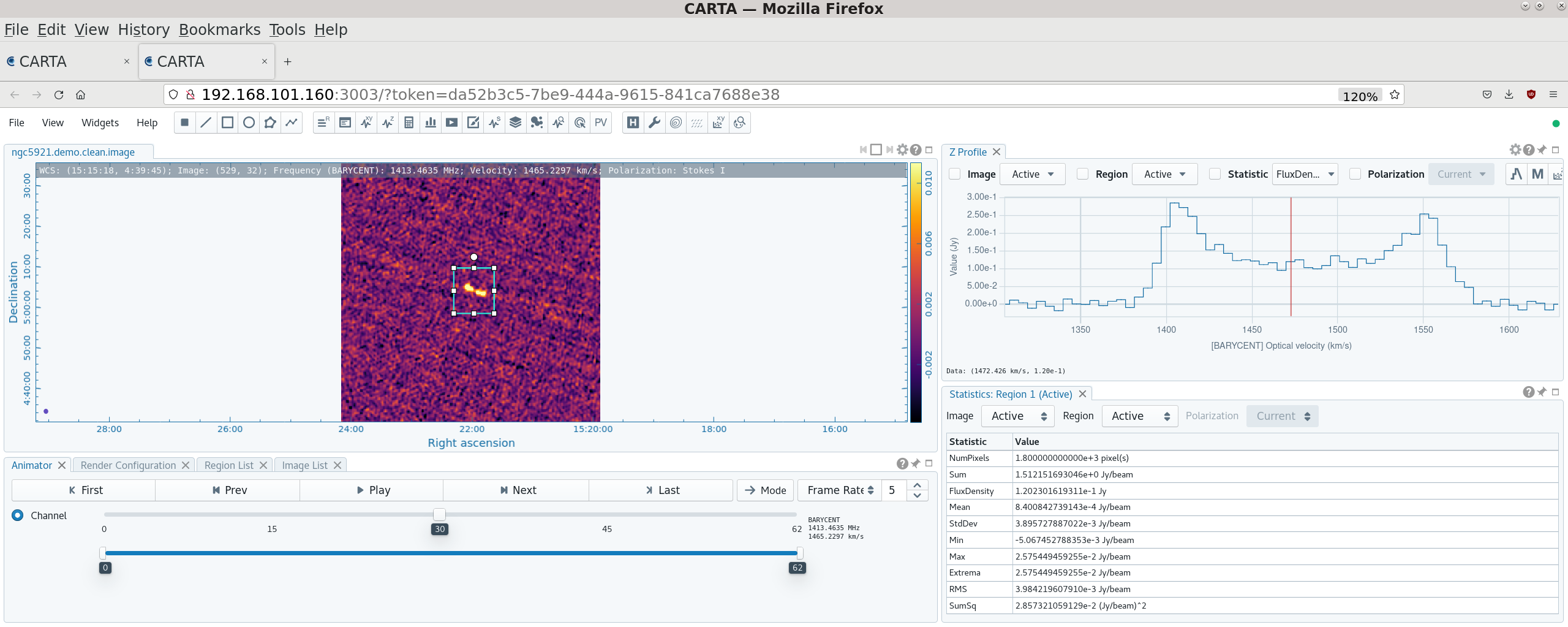
CARTA can be used to produce an integrated spectrum from the spectral line cube. Once you have defined a region (you can use the region used earlier to define the region to be cleaned), you can set the Z profile statistic to 'Sum' to see the spectrum of the pixels summed in this region in each channel, 'Mean' to see the mean spectrum, or 'FluxDensity' to see the beam-corrected flux density spectrum (which is probably what you want). Different plotting and smoothing options are available, and you can export an image of the spectrum or export the data (as a tab-separated variables file, which you can then read into your favorite plotting program, iPython notebook, etc.).
Cube moments
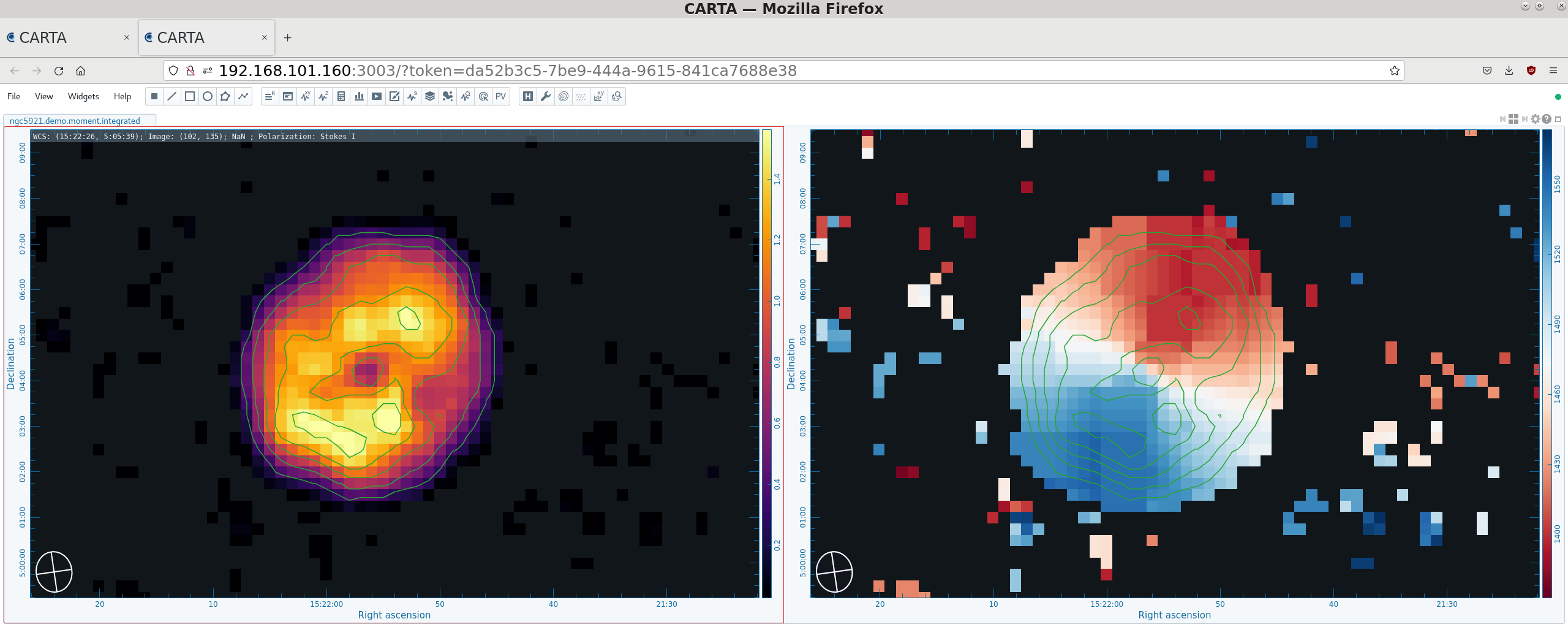
Cube moments are maps of weighted sums along the velocity axis. In CASA, they are generated by the task immoments. The zeroth moment (moments = 0) is a sum of intensities along the velocity axis (the integrated intensity map); the first moment (moment = 1) is the sum of velocities weighted by intensity (the velocity field); the second moment (moment = 2) is a map of the velocity dispersion; see the immoments documentation for additional options.
The following example produces maps of the zeroth and first moments, or the integrated intensity and velocity field. To form these maps, we will mask out pixels without signal (below 3 sigma) using hard global limits. We will also collapse along the spectral (channel) axis (the default) and include those channels that show flux in the integrated spectrum, i.e. 9~48.
#Interactive CASA default immoments inp imagename='ngc5921.demo.clean.image' moments=[0,1] chans='9~48' excludepix=[-100,0.00525] outfile='ngc5921.demo.moment' inp go
#Pseudo-interactive CASA
immoments(imagename='ngc5921.demo.clean.image', moments=[0,1], excludepix=[-100, 0.00525], axis='spectral', chans='9~48', outfile='ngc5921.demo.moment')
Where:
imagename='ngc5921.demo.clean.image' # input image cube moments = [0,1] # do zeroth and first moments excludepix = [-100,0.00525] # use only pixels > 3 sigma outfile='ngc5921.demo.moment' # the output filename root (or filename if only doing one moment)
You will get a lot of warnings in the logger and the terminal about the convolving kernel having a size less than the pixel diagonal length. This is because immoments has to convolve all of the channels to have the same beam size prior to forming the moment map, but as they all have very similar beam sizes already the convolving kernel needed for this is fairly small. These warnings can be ignored.
Two files are created by this: ngc5921.demo.moment.integrated (the zeroth moment) and ngc5921.demo.moment.weighted_coord (the first moment). These can be opened and manipulated in CARTA.
Export the Data
To export the uv data and the image cube as FITS files, use exportuvfits and exportfits respectively.
For example, to export the continuum-subtracted uv data (the measurement set we used as input to tclean):
#Interactive python default exportuvfits inp vis='ngc5921.demo.uvcontsub.ms' fitsfile='ngc5921.demo.uvcontsub.uvfits' datacolumn='data' inp go
#Pseudo-interactive python
exportuvfits(vis='ngc5921.demo.uvcontsub.ms', fitsfile='ngc5921.demo.uvcontsub.uvfits', datacolumn='data')
Note: the default is to export the CORRECTED_DATA column. If we wanted to export the full data-set after applying the calibration this would be appropriate, but uvcontsub writes out to the DATA column into a new measurement set.
And for the cleaned cube:
#Interactive python default exportfits inp imagename='ngc5921.demo.clean.image' fitsimage='ngc5921.demo.clean.fits' velocity=True optical=True inp go
In this example, we have chosen to write the FITS cube with velocity rather than frequency on the spectral axis and to use the optical (cz) convention for velocity. Both of these are, of course, optional.
#Pseudo-interactive python
exportfits(imagename='ngc5921.demo.clean.image', fitsimage='ngc5921.demo.clean.fits')
The moment maps (or any other CASA images) can be similarly exported using exportfits.
Appendix: Python Notes
os.system
os.system allows you to run shell commands from within python / CASA. For example:
import os
os.system('ls -sF')
will give an OS-level listing of the current directory's contents.
os.environ.get
It's worth having a look at the output of the os.environ.get command to understand the python syntax (alternative: os.getenv). You can think of os.environ as a list of operating system environment variables, and get is a method that extracts information about the requested environment variable, here, CASAPATH. Get returns a string of whitespace separated information, and .split() turns that string into a list. The array index [0] extracts the first element of that list, which contains the path.
To illustrate, here is some example python I/O in CASA.
CASA <12>: print os.environ.get('CASAPATH')
/usr/lib64/casapy/30.0.9709test-001 linux local el5bld64b
CASA <13>: print os.environ.get('CASAPATH').split()
['/usr/lib64/casapy/30.0.9709test-001', 'linux', 'local', 'el5bld64b']
CASA <14>: print os.environ.get('CASAPATH').split()[0]
/usr/lib64/casapy/30.0.9709test-001
Last checked on CASA Version 6.5.2.