CASA Guides:ATCA Advanced Continuum Polarization Tutorial NGC612-CASA4.7
This CASA Guide is for Version 4.7 of CASA. If you are using a later version of CASA and this is the most recent available guide, then you should be able to use most, if not all, of this casaguide, as we try to limit script breaking changes in CASA development.
Overview
This CASA guide describes the loading, flagging, calibration and imaging of a two-pointing continuum data set taken with the Australia Telescope Compact Array (ATCA) of the nearby radio galaxy NGC612. It has been adapted from the 3C391 Continuum Tutorial (CASA 4.6) for the VLA . The data were taken with 2048 MHz of bandwidth in 1 MHz channels, centered at 2.1 GHz, and recorded all 4 polarizations (XX,XY,YX,YY). For ATCA data reduction there are two paths you can follow:
- If you are already familiar with the Miriad package, you can do the loading and optionally the flagging and calibration of the data in Miriad and then import the data into a CASA MeasurementSet with importmiriad.
- Alternatively, you can use importatca to load the ATCA archive files directly into CASA.
If this is your first attempt at using CASA for Compact Array data you are encouraged to start with a flagged and calibrated Miriad dataset and then try imaging the data following [this guide]. If you feel you are ready to tackle the full reduction job in CASA keep reading.
Obtaining the Data
To download the data needed for this tutorial you will need to use the [Australia Telescope Online Archive]. If you don't have an account already, you can request one [here]. Once logged in enter the project code C2728, click search and select the files 2012-10-25_0707.C2728, 2012-10-25_0903.C2728, 2012-10-25_1304.C2728 and 2012-10-25_1705.C2728 from the results. You can then use the download button to retrieve a tar file with the data, or get a file with the links to the data to download from another machine. (If that all seems like too much trouble, a ready made MS with the data is available for download [here](3.5GB download), you can extract that instead and continue in the section #The Observation.)
Once the download is complete, extract the files (within a working directory in which you will then run CASA):
# In a Terminal:
tar xvf datafiles.tar
How to Use This CASA Guide
Here are a number of possible ways to run CASA, described in more detail in Getting Started in CASA. In brief, there are at least three different ways to run CASA:
- Interactively examining task inputs. In this mode, one types taskname to load the task, inp to examine the inputs, and go once those inputs have been set to your satisfaction. Allowed inputs are shown in blue and bad inputs are colored red. The input parameters themselves are changed one by one, e.g., selectdata=T. Screenshots of the inputs to various tasks used in the data reduction below are provided, to illustrate which parameters need to be set. More detailed help can be obtained on any task by typing help taskname. Once a task is run, the set of inputs are stored and can be retrieved via tget taskname; subsequent runs will overwrite the previous tget file.
- Pseudo-interactively via task function calls. In this case, all of the desired inputs to a task are provided at once on the CASA command line. This tutorial is made up of such calls, which were developed by looking at the inputs for each task and deciding what needed to be changed from default values. For task function calls, only parameters that you want to be different from their defaults need to be set.
- Non-interactively via a script. A series of task function calls can be combined together into a script, and run from within CASA via execfile('scriptname.py'). This and other CASA Tutorial Guides have been designed to be extracted into a script via the script extractor by using the method described at the Extracting_scripts_from_these_tutorials page. Should you use the script generated by the script extractor for this CASA Guide, be aware that it will require some small amount of interaction related to the plotting, occasionally suggesting that you close the graphics window and hitting return in the terminal to proceed. It is in fact unnecessary to close the graphics windows (it is suggested that you do so purely to keep your desktop uncluttered).
If you are a relative novice or just new to CASA, it is strongly recommended to work through this tutorial by cutting and pasting the task function calls provided below after you have read all the associated explanations. Work at your own pace, look at the inputs to the tasks to see what other options exist, and read the help files. Later, when you are more comfortable, you might try to extract the script, modify it for your purposes, and begin to reduce other data.
Importing the ATCA archive data
Before beginning our data reduction, we must start CASA. If you have not used CASA before, some helpful tips are available on the Getting Started in CASA page.
Once you have CASA up and running in the directory containing the data, then start your data reduction by getting the data into a MeasurementSet. The task to do this, importatca, lets us choose which of the simultaneously recorded frequencies to load. In this case we just want the continuum data, which we can select with spw=0. There are two other frequency settings in this data, a 33 channel continuum band and a 2048 channel spectral zoom band centered on the HI emission from NGC612. We might use the latter for a future spectral line tutorial. We specify the options 'birdie' to remove known bad channels and 'noac' to discard the autocorrelations. While importatca supports wildcards, we've noticed the order in which the files load is not consistent across platforms, so we've listed them in full here.
# In CASA
importatca(vis='ngc612.ms.0',files=['2012-10-25_0707.C2728', '2012-10-25_0903.C2728', '2012-10-25_1304.C2728', '2012-10-25_1705.C2728'],spw=0,options='birdie,noac')
As the data loads a listing appears in the logger of the data encountered. The first part is reproduced here:
##########################################
##### Begin Task: importatca #####
importatca(files=['2012-10-25_0707.C2728', '2012-10-25_0903.C2728', '2012-10-25_1304.C2728', '2012-10-25_1705.C2728'],vis="ngc612.ms",options="birdie,noac",spw=0,nscans=[0, 0],
lowfreq="0.1GHz",highfreq="999GHz",fields=[''],edge=8)
Expanded file names are :
/data/ngc612/2012-10-25_0707.C2728
/data/ngc612/2012-10-25_0903.C2728
/data/ngc612/2012-10-25_1304.C2728
/data/ngc612/2012-10-25_1705.C2728
Checking header of file /data/ngc612/2012-10-25_0707.C2728
CABB data detected
Reading file /data/ngc612/2012-10-25_0707.C2728
Scan # : 1
Object : 1934-638
Date : 2012/10/25/07:07:45
First data/header has NAnt=6, NChan=2049, NPol=4
Antenna : CA01 position:[-4.75168e+06, 2.79161e+06, -3.20048e+06]
Antenna : CA02 position:[-4.75159e+06, 2.79176e+06, -3.20048e+06]
Antenna : CA03 position:[-4.75156e+06, 2.79181e+06, -3.20048e+06]
Antenna : CA04 position:[-4.75135e+06, 2.79217e+06, -3.20048e+06]
Antenna : CA05 position:[-4.75129e+06, 2.79227e+06, -3.20048e+06]
Antenna : CA06 position:[-4.74939e+06, 2.79549e+06, -3.20048e+06]
IF 1 : Frequency = 2100 MHz, bandwidth = 2048MHz, #channels = 2049
: Polarizations XX - 9 , XY - 10 , YX - 11 , YY - 12
Scan 1 stored
Scan # : 2
Object : secndry_cal
Date : 2012/10/25/07:27:15
Scan 2 stored
Scan # : 3
Object : east_lobe
Date : 2012/10/25/07:31:15
Scan 3 stored
Scan # : 4
Object : secndry_cal
Date : 2012/10/25/07:32:25
Scan 4 stored
Scan # : 5
Object : east_lobe
Date : 2012/10/25/07:35:05
Scan 5 stored
Scan # : 6
Object : west_lobe
Date : 2012/10/25/07:40:45
Scan 6 stored
Scan # : 7
Object : east_lobe
Date : 2012/10/25/07:46:05
Scan 7 stored
Scan # : 8
Object : west_lobe
Date : 2012/10/25/07:51:25
Scan 8 stored
Scan # : 9
Object : east_lobe
Date : 2012/10/25/07:56:55
Scan 9 stored
Scan # : 10
Object : west_lobe
Date : 2012/10/25/08:02:15
Scan 10 stored
Scan # : 11
Object : east_lobe
Date : 2012/10/25/08:07:45
Scan 11 stored
Scan # : 12
Object : west_lobe
Date : 2012/10/25/08:13:05
Scan 12 stored
Scan # : 13
Object : east_lobe
Date : 2012/10/25/08:18:25
Scan 13 stored
Scan # : 14
Object : west_lobe
Date : 2012/10/25/08:23:45
Scan 14 stored
Scan # : 15
Object : secndry_cal
Date : 2012/10/25/08:29:05
Scan 15 stored
Scan # : 16
Object : secndry_cal
Date : 2012/10/25/08:31:45
Scan 16 stored
Scan # : 17
Object : east_lobe
Date : 2012/10/25/08:34:05
Scan 17 stored
Scan # : 18
Object : west_lobe
Date : 2012/10/25/08:39:45
Scan 18 stored
Scan # : 19
Object : secndry_cal
Date : 2012/10/25/08:45:05
Scan 19 stored
Scan # : 20
Object : east_lobe
Date : 2012/10/25/08:47:45
Scan 20 skipped
Scan # : 21
Object : 1934-638
Date : 2012/10/25/08:47:45
Scan 21 stored
Scan # : 22
Object : 1934-638
Date : 2012/10/25/08:54:05
IF 2 : Frequency = 1891 MHz, bandwidth = 2048MHz, #channels = 33
: Polarizations XX - 9 , XY - 10 , YX - 11 , YY - 12
End of File
Number of rows selected : 8910
Flagged : 5.1%
Antenna off source : 5.1%
ScanType (Point/Paddle): 0%
Scans that don't have valid or selected data will be marked 'skipped'. At the end a summary of the on-line flagging appears, we see about 5% of the data is flagged because antennas were not on source.
The Observation
The task listobs can be used to get a listing of the individual scans comprising the observation, the frequency setup, source list, and antenna locations. Since we've already seen the scans scroll past, we'll use verbose=False to get the summary:
# In CASA
listobs(vis='ngc612.ms.0',verbose=False)
From the source listing we see there are 5 sources, the primary calibrator 1934-638, a secondary calibrator, and two target sources called east_lobe and west_lobe and some data on 0537-441. We also see the data has a 2048 MHz bandwidth, with 1 MHz channels, centered on 2.1 GHz with 4 linear polarizations.
##### Begin Task: listobs #####
listobs(vis="ngc612.ms.0",selectdata=True,spw="",field="",antenna="",
uvrange="",timerange="",correlation="",scan="",intent="",
feed="",array="",observation="",verbose=False,listfile="",
listunfl=False,cachesize=50,overwrite=False)
================================================================================
MeasurementSet Name: /data/ngc612/ngc612.ms.0 MS Version 2
================================================================================
Observer: obs Project: C2728
Observation: ATCA(6 antennas)
Data records: 60465 Total elapsed time = 43730 seconds
Observed from 25-Oct-2012/07:07:39.9 to 25-Oct-2012/19:16:29.9 (UTC)
Compute subscan properties
Fields: 5
ID Code Name RA Decl Epoch SrcId nRows
0 C 1934-638 19:39:25.025995 -63.42.45.63000 J2000 0 6645
1 secndry_cal 00:51:09.502006 -42.26.33.29294 J2000 1 5880
2 east_lobe 01:34:16.760003 -36.30.13.52004 J2000 2 23160
3 west_lobe 01:33:34.630993 -36.29.21.54998 J2000 3 22515
4 C 0537-441 05:38:50.362001 -44.05.08.93995 J2000 4 2265
Spectral Windows: (1 unique spectral windows and 1 unique polarization setups)
SpwID Name #Chans Frame Ch0(MHz) ChanWid(kHz) TotBW(kHz) CtrFreq(MHz) Corrs
0 2049 TOPO 3124.000 1000.000 2048000.0 2100.0000 XX XY YX YY
Antennas: 6 'name'='station'
ID= 0-3: 'CA01'='W98', 'CA02'='W109', 'CA03'='W113', 'CA04'='W140',
ID= 4-5: 'CA05'='W148', 'CA06'='W392'
##### End Task: listobs #####
##########################################
The antenna station names give away that this is a compact configuration, with CA01-CA05 close together and CA06 a long way away (multiply the station numbers by 15 to get distance in meters from station W0). It is actually a 750m array configuration, which means we would normally not include CA06 in the imaging. The galaxy ngc612 is quite large compared to the primary beam at 2 GHz, so two pointings were used to cover the emission with adequate sensitivity. The observation ran for close to 12 hours, this should give good uv coverage with the array in a short E-W configuration.
Examining and Editing the Data
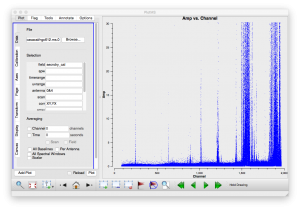
It is always a good idea to examine the data before jumping straight into calibration. Start by flagging data known to be bad from observing logs, then examine the data. At low frequencies there is a lot of RFI, some of it is very strong and persistent so lets get of rid of that first. Use plotms to make a plot of the XY and YX correlations on the secondary calibrator. These correlations should normally have little signal in them, certainly anything stronger than the calibrator flux will be interference. Specify axes of 'channel' and 'amp', zoom in and use the resulting plots to read off the worst channel ranges. To cut down on the amount of data we can use antenna selection "0&4", the longest baseline we're likely to use, if it is bad, the shorter ones will be worse. The following command flags the data in all the bad channel ranges we found, you may decide to excise a few more or fewer channel ranges. At the highest channel number (lowest frequencies) the RFI gets so bad, we've deleted large channel ranges. If you were desperate for spectral information in this range you could do a more careful job. Note that it is not necessary to be very accurate in this step, as we will do further flagging after this first pass.
By default flagdata will save a copy of the existing set of flags before entering any new flags. This is so that you can easily revert any flags you have applied using the flagmanager task. This comes in very handy for those occasions where you accidentally flag more than you intended.
# In CASA
flagdata(vis='ngc612.ms.0',mode='manual',spw='0:243~247;548~551;631~634;895~899;1012~1025;1205~1234;1421~1439;1495~1599;1614~1625;1675~1676;1815~2048')
Next we will do some clipping on the cross correlations. To choose the clip level we checked the amplitudes on a XY and YX plot with all the data, to make sure we're not clipping any good data. We can clip the primary and secondary calibrator at lower levels than the strongly polarized source. After clipping we extend the flagged points using 'growaround=True' and also flag the corresponding parallel hand polarizations. You can also do this editing directly in plotms, as you notice the bad data, but while convenient, this doesn't let you easily redo the same flagging when you want to rerun your calibration script.
# In CASA
flagdata(vis='ngc612.ms.0',mode='clip',clipminmax=[0,6],correlation='ABS,XY,YX')
flagdata(vis='ngc612.ms.0',mode='clip',clipminmax=[0,3],correlation='ABS,XY,YX',field='0')
flagdata(vis='ngc612.ms.0',mode='clip',clipminmax=[0,1.5],correlation='ABS,XY,YX',field='1')
flagdata(vis='ngc612.ms.0',mode='extend',extendpols=True,growaround=True)
As a final step we'll use the 'tfcrop' algorithm to find further bad data. It automatically does a flag extend to the other polarizations. Read the help for flagdata for the details on how these flagging algorithms work.
# In CASA
flagdata(vis='ngc612.ms.0',mode='tfcrop',usewindowstats='sum',correlation='ABS,XY,YX',halfwin=3)
After these steps the data is much improved but after replotting the secondary calibrator we notice one more flaw: there is a single integration with excess flux on the secondary calibrator.
# In CASA
flagdata(vis='ngc612.ms.0',mode='manual',timerange='14:15:30~14:15:40')
After fixing that we inspect the rest of the data and it is looking pretty clean. We could do some more flagging on the ngc612 data, but it is probably not warranted unless we notice problems later in the reduction.
Calibrating the Data
It is now time to begin calibrating the data. The general data reduction strategy is to derive a series of scaling factors or corrections from the calibrators, which are then collectively applied to the science data. For more discussion of the philosophy, strategy, and implementation of calibration of synthesis data within CASA, see Synthesis Calibration in the CASA Reference Manual.
The first thing we want to do is split the wide bandwidth of 2048 MHz up into 8 spectral windows of 256 MHz each so we can solve for the gains and polarisation leakages per spectral window. Make sure you are using CASA 4.7 or later for this step as earlier versions of mstransform had a bug that scrambled the baselines when used in this way.
# In CASA
mstransform(vis='ngc612.ms.0',outputvis='ngc612.ms',datacolumn='data',mode='channel',
regridms=True,spw='0',nspw=8)
Next we set the flux scale for the primary calibrator 1934-638. We have a choice of standards: 'Perley-Butler 2010' works fine at frequencies below 11 GHz; 'Stevens-Reynolds 2016' first appeared in CASA 4.7 and has the same scale below 11 GHz but is more accurate above that frequency, make sure to use this for 15 or 7mm data. We also define a few shortcut names for the primary, secondary and MeasurementSet name to save typing and we've chosen a central antenna as the reference. If you haven't managed to observe 1934-638 in your observation, you may be able to 'borrow' an observation from an observation directly before or after yours if it happened to use the same setup, otherwise your best bet is to check the ATCA Calibrator database for a recent observation of your calibrator and use the flux from that.
# In CASA
#set flux scale for primary calibrator
msname='ngc612.ms'
pri='1934-638'
sec='secndry_cal'
ref='CA04'
setjy(vis=msname,field=pri,scalebychan=True,standard='Perley-Butler 2010')
The next thing we want to do is derive a bandpass solution for our data. Before we can do that though, we need to phase calibrate the data so we can integrate over a long solution interval to do the bandpass solution. Because we're processing polarization data from an Az/El telescope we switch on parallactic angle rotation using parang=True in all the steps from here.
# In CASA
gaincal(vis=msname,caltable='cal.G0',field=pri,refant=ref,gaintype='G',calmode='p',parang=True,solint='60s');
bandpass(vis=msname,caltable='cal.B0',field=pri,spw='',refant=ref,solnorm=True,solint='inf',bandtype='B',gaintable=['cal.G0'],parang=True);
After the bandpass solution we're ready to determine the gains using the secondary calibrator. We also solve for the gains on the primary. We're using a 60s solution interval and apply the bandpass table we've just created.
# In CASA
gaincal(vis=msname,caltable='cal.G1',field=pri+','+sec,refant=ref,spw='*',gaintype='G',calmode='ap',parang=True,solint='60s',gaintable=['cal.B0'])
The remaining calibration to solve for is the polarization leakage or D-terms. For the ATCA we can often use the unpolarized primary calibrator 1934-638 for this, but you can also use the secondary calibrator. In both cases we need to determine the polarization of the secondary so we can correct the gains for source polarization. Here we use polcal with the primary calibrator and apply the previously determined bandpass and gain calibration.
# In CASA
polcal(vis=msname,caltable='cal.D0',field=pri,refant=ref,gaintable=['cal.B0','cal.G1'],poltype='D',solint='inf')
We then use the qufromgain routine to extract polarized source model for the secondary. The qufromgain routine works out the Q and U of the calibrator for each spectral window and returns the mean values. We capture those and use them as a first order correction for the next round of calibration. Since we don't know the flux of the secondary yet, it is assumed to be 1 by gaincal and our source model is relative to this. The qufromgain routine assumes the X feed is offset by 45 degrees for ATCA (except for 7mm observations when it uses 135 degrees, if you have analyzed polarized 7mm data let us know if you think this is incorrect). We also need to select the secondary calibrator with the field id, which is 1. We exclude spectral windows 6 and 7 from the mean because they gave inconsistent results and probably need more RFI flagging. We also excluded antenna CA03 (or index 2) because it has some strange steps in the gains (one of appears to coincide with a change in the focus setting). You are advised to use plotcal to inspect the gain table used in this step. The qufromgain routine looks at the ratio between the X and Y gains, so we specify poln='/' to inspect that.
# In CASA
#plotcal(caltable='cal.G1',xaxis='time',yaxis='amp',poln='/',iteration='antenna,spw')
# In CASA
from recipes.atcapolhelpers import qufromgain
# from casa 6 onwards use
#from casarecipes.atcapolhelpers import qufromgain
qu = qufromgain('cal.G1',fieldids=[1],badspw=[6,7],badant=[2])
smodel=[1,qu[1][0],qu[1][1],0]
As you can see the secondary calibrator is only very weakly polarized (~1%) so this step is not as important here as it might be for a strongly polarized calibator. Now we redo the whole calibration using our best estimates for gains, leakages and secondary polarization. We have to split the gain calibration into two steps so we can specify the source model for the secondary. We use the parameter append=True to get the solutions for the primary and secondary into the same table as that is what we need for the fluxscale step that comes next.
# In CASA
# Redo gaincal
bandpass(vis=msname,caltable='cal.B1',field=pri,spw='',refant=ref,solnorm=True,
solint='inf',bandtype='B',gaintable=['cal.G1','cal.D0'],parang=True);
gaincal(vis=msname,caltable='cal.G2',field=pri,refant=ref,spw='*',
gaintype='G',calmode='ap',parang=True,solint='60s',gaintable=['cal.B1','cal.D0'])
gaincal(vis=msname,caltable='cal.G2',field=sec,refant=ref,spw='*',
gaintype='G',calmode='ap',parang=True,solint='60s',
gaintable=['cal.B1','cal.D0'],smodel=smodel,append=True)
polcal(vis=msname,caltable='cal.D1',field=pri,refant=ref,gaintable=['cal.B1','cal.G2'],
poltype='D',solint='inf')
All that remains now is to correct the flux scale using comparison between the primary and secondary calibrator and then apply all the corrections. We use the gainfield parameter in applycal to indicate which solutions to apply from each calibration table.
# In CASA
# flux calibration
fluxscale(vis=msname,caltable='cal.G2',fluxtable='cal.F0',reference=pri);
# Apply calibration
applycal(vis=msname,gaintable=['cal.B1','cal.D1','cal.F0'],gainfield=[pri,pri,sec],parang=True)
At this point we've got a fully calibrated dataset ready for imaging. The imaging now proceeds almost exactly as in the ATCA tutorial that started with a calibrated dataset [[1]]
Multi-scale Polarization Clean
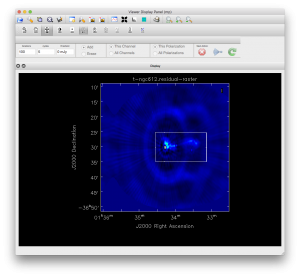
To image this data with 2 pointings we need to use mosaicing, and because the source is quite extended, we will also use the multiscale option. Because the lower part of the band has a lot of interference and the full 1.1-3.1 GHz band is too wide to image properly in one go (even with MFS) we will restrict the imaging for this tutorial to the top 1/3 of the band. (You could try imaging the rest of the band yourself.) To specify this selection we need to use 'spw='0,1,2:0~170', indicating all of spectral windows 0 and 1, and channel 0 to 170 from spectral window 2. At an average frequency of about 2.7 GHz the primary beam is about 17' and the resolution for a 750m array is about 20". Based on this we'll choose a cell size of 5" and and image size of 500 pixels. We will make a test image to see what the field looks like and use the interactive option so we can set the area to clean. In the example we've used a simple box, but you can use polygons or other shapes. Once you have set the clean area you can save it to a file and then enter this as the mask for deeper cleaning later. After some experimentation we've chosen the multiscale parameter to correspond to: a point source, about twice the beam and about the size of the largest 'blob' in the image. Setting the largest scale much smaller tends to leave a negative bowl around the source.
# In CASA
clean(vis='ngc612.ms',imagename='t-ngc612',spw='0,1,2:0~170',imagermode='mosaic',multiscale=[0,8,24],
imsize=500,cell='5.0arcsec',stokes='I',weighting='briggs',robust=0.5,field='*lobe',antenna='!5',interactive=True)
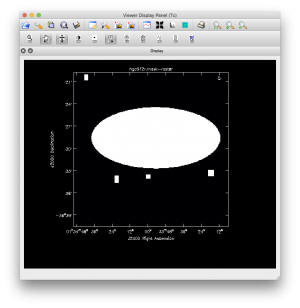
Now that we have a feel for what the source looks like, we can clean a bit deeper and do all the Stokes parameters. A few point sources outside the box appear with a deeper clean, so we should really redo the clean with those sources added into the mask. Figure 3 shows the mask used in the final clean: using the viewer region editor we changed the box to an ellipse and added 4 boxes around point sources. You can get the region file here: Media:ngc612region.txt, after you download it, rename it to ngc612region.crtf. Looking at the final image we see we could probably improve on the clean region, as there is still a negative region visible close to the source.
# In CASA
clean(vis='ngc612.ms',imagename='ngc612h',spw='0,1,2:0~170',imagermode='mosaic',multiscale=[0,8,24],
imsize=500,cell='5.0arcsec',stokes='IQUV',weighting='briggs',robust=0.5,niter=10000,threshold='0.5mJy',
field='*lobe',antenna='!5',mask='ngc612region.crtf' )
Constructing Polarization Intensity and Angle Images
We've made a full polarization image of ngc612. This image is actually a cube with the standard two angular dimensions (right ascension, declination) and a third dimension containing the polarization information. Considering the image cube as a matrix, [math]\displaystyle{ Image[l,m,p] }[/math], the [math]\displaystyle{ l }[/math] and [math]\displaystyle{ m }[/math] axis describe the sky brightness or intensity for the given [math]\displaystyle{ p }[/math] axis. If one opens the viewer and loads the ngc612 continuum image, the default view contains an animator or pane with movie controls. One can step through the polarization axis, displaying the images for the different polarizations.
As created, the image contains four polarizations, one for each of the four Stokes parameters: I, Q, U, and V. Recall that Stokes Q and U describe linear polarization and V describes circular polarization. Specifically, Q describes the amount of linear polarization aligned with a given axis, and U describes the amount of linear polarization at a 45 deg angle to that axis. The V parameter describes the amount of circular polarization, with the sign (positive or negative) describing the sense of the circular polarization (right- or left-hand circularly polarized).
Few celestial sources, with the notable exception of masers, are expected to show circular polarization. Terrestrial and satellite sources, however, are often highly circularly polarized. The V image is therefore often worth forming because any V emission could be indicative of unflagged RFI within the data (or problems with the calibration!). In our image we can see some remnant Stokes V at the position of the strongest source and an error pattern around it that has not cleaned away. This indicates it is probably due to remaining calibration errors.
Because the Q and U images both describe the amount of linear polarization, it is more common to work with a linear polarization intensity image, [math]\displaystyle{ P = \sqrt{Q^2 +U^2} }[/math] and the polarization position angle [math]\displaystyle{ tan 2\chi = U/Q }[/math].
1. Extract the Q and U planes from the full Stokes image cube, forming separate images for each Stokes parameter.
# In CASA
imsubimage(imagename='ngc612h.image',outfile='ngc612h.Q',stokes='Q')
imsubimage(imagename='ngc612h.image',outfile='ngc612h.U',stokes='U')
2. Combine the Q and U images using the parameter mode='poli' of immath to form the linear polarization image. Because P is formed from the sum of the squares of the Q and U values, it is biased by the noise. You can set the sigma value to debias this (it subtracts the square of this value from the Q2+U2 before taking the square-root). Use the imstat task to get the rms in Stokes V, which will usually give a good estimate of the noise. You could just look at the output and enter the rms value as sigma in immath, adding the correct units. Here we show how to do this from a script by capturing the output of imstat into a variable and then extracting the rms value; because the returned value is an array we need to index it with 0 to get the value as a floating point number: s['rms'][0]. Note: due to issues with the image produced this way, the script below uses a different strategy to get the same result.
# In CASA
s=imstat(imagename='ngc612h.image',stokes='V')
sigma=s['rms'][0]
# the next line works, but the image produced has NaN values which causes problems in the viewer, to avoid this we calculate things using mode='evalexpr'
#immath(imagename=['ngc612h.Q','ngc612h.U'],mode='poli',outfile='ngc612h.P',sigma=str(sigma))
P2='IM0^2+IM1^2-'+str(sigma**2)
immath(imagename=['ngc612h.Q','ngc612h.U'],mode='evalexpr',outfile='ngc612h.P',expr='iif('+P2+'>0, sqrt('+P2+'),0.0)')
2. Combine the Q and U images using the parameter mode='pola' of immath to form the polarization position angle image. To avoid displaying the position angle of noise, we can set a threshold intensity of the linear polarization, above which we wish to calculate the polarization angle, using the parameter polithresh. An appropriate level here might be the [math]\displaystyle{ 8\sigma }[/math] level of around 1 mJy/beam:
# In CASA
immath(imagename=['ngc612h.Q','ngc612h.U'],mode='pola',outfile='ngc612h.X',polithresh='1mJy/beam')
3. Now use the viewer to display all the images together:
- load the cube as a raster image and set the levels so you can see the extended emission
- load the linear polarization image as a contour image, set the contours to 1, 3, 10, 30, 100 times 1 mJy/beam
- load the polarization image as a vector image, you can switch the line color to black to make it stand out more
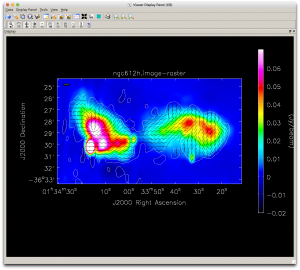
The result should look something like Figure 4.
4. We'll extract some statistics so you can compare them with your own results:
- The total polarized flux in the lobes above the 2 mJy contour.
- The change in average polarization angle across the western lobe.
# In CASA
mystats1=imstat(imagename='ngc612h.P',mask='ngc612h.P>0.002')
mystats2=imstat(imagename='ngc612h.X',region='box [ [ 325pix , 260pix] , [345pix, 275pix ] ]')
mystats3=imstat(imagename='ngc612h.X',region='box [ [ 365pix , 250pix] , [385pix, 265pix ] ]')
print 'Total polarized flux density: %6.3f Jy' % mystats1['flux']
print 'Pol. angles in western lobe: %4.1f and %4.1f degrees' % (mystats2['mean'],mystats3['mean'])
Running the above commands should produce something close to the following output:
Total polarized flux density: 1.183 Jy Pol. angles in western lobe: -60.5 and 23.8 degrees
5. As further practice you could try to image the rest of the band by changing the channel selection and imaging the middle and lower part of the band. You may need to do some additional flagging to get a decent image.
Questions about this tutorial? Please contact the NRAO Helpdesk.
Last checked on CASA Version 5.6.2.